Abstract
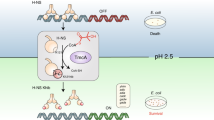
Cysteine residue 69 of theEscherichia coli Ada transcription factor, which accepts a methyl group from methylphosphotriester in methylated DNA, was substituted by each of 19 other amino acids. Only the mutant Ada (C69H), carrying a histidine substitution of Cys69, exhibited a limited degree of transactivating potential for theada promoter inE. coli cells although the mutant protein was completely devoid of methylphosphotriester-DNA methyltransferase activity. Using a multicopy plasmid system for the expression of Ada protein, we have shown that Ada C69H has a transactivating capacity equivalent to that of wild-type Ada protein in the absence of an alkylating agent. This indicates that the zinc-binding capacity of histidine at residue 69 is likely to be sufficient for Ada to recognize and bind to theada promoter. Furthermore, transactivation of theada promoter by Ada C69H was enhanced up to 6-fold by treatment with methylating agents. An additional substitution was made with alanine in Ada C69H, replacing Cys321, the site for acceptance of a methyl group from O6-methylguanine and O4-methylthymine residues in DNA, with alanine. This renders the protein completely inactive as a methyltransferase but this derivative is constitutively active as a transactivator for theada promoter. Therefore, acquisition of a methyl group at Cys321 apparently enhances the transactivating capacity of Ada protein on theada promoter. We propose that the transcription-regulating function of Ada protein is under dual control by methylation of cysteine residues at positions 69 and 321; the former enhances DNA binding, while the latter enhances the transactivating capacity of the protein.
Similar content being viewed by others
References
Akimaru H, Sakumi K, Yoshikai T, Anai M, Sekiguchi M (1990) Positive and negative regulation of transcription by a cleavage product of Ada protein. J Mol Biol 216:261–273
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Cathcart R, Goldthwait DA (1981) Enzymatic excision of 3-methyladenine and 7-methylguanine by a rat liver nuclear fraction. Biochemistry 20:273–280
Chang ACY, Cohen SN (1978) Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the P15A cryptic miniplasmid. J Bacteriol 134:1145–1156
Chueh LL, Nakamura T, Nakatsu Y, Sakumi K, Hayakawa H, Sekiguchi M (1992) Specific amino acid sequences required forO 6-methylguanine-DNA methyltransferase activity: analyses of three residues at or near the methyl acceptor site. Carcinogenesis 13:837–843
Demple B (1986) MutantEscherichia coli Ada proteins simultaneously defective in the repair ofO 6-methylguanine and in gene activation. Nucleic Acids Res 14:5575–5589
Demple B, Sedgwick B, Robins P, Totty N, Waterfield MD, Lindahl T (1985) Active site and complete sequence of the suicidal methyltransferase that counters alkylation mutagenesis. Proc Natl Acad Sci USA 82:2688–2692
Hayakawa H, Koike G, Sekiguchi M (1990) Expression and cloning of complementary DNA for a human enzyme that repairsO 6-methylguanine in DNA. J Mol Biol 213:739–747
Hemminki K (1983) Nucleic acid adducts of chemical carcinogens and mutagens. Arch Toxicol 52:249–285
Ihara K, Kawate H, Chueh LL, Hayakawa H, Sekiguchi M (1994) Requirement of the Pro-Cys-His-Arg sequence forO 6-methylguanine-DNA methyltransferase activity revealed by saturation mutagenesis with negative and positive screening. Mol Gen Genet 243:379–389
Iyama A, Sakumi K, Nakabeppu Y, Sekiguchi M (1994) A unique structural feature of rabbit DNA repair methyltransferase as revealed by cDNA cloning. Carcinogenesis 15:627–633
Lawley PD (1974) Some chemical aspects of dose-response relationships in alkylation mutagenesis. Mutat Res 23:283–295
Lindahl T, Sedgwick B, Sekiguchi M, Nakabeppu Y (1988) Regulation and expression of the adaptive response to alkylating agents. Annu Rev Biochem 57:133–157
McCarthy TV, Lindahl T (1985) Methyl phosphotriesters in alkylated DNA are repaired by the Ada regulatory protein ofE. coli. Nucleic Acids Res 13:2683–2698
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Moore MH, Gulbis JM, Dodson EJ, Demple B, Moody PCE (1994) Crystal structure of a suicidal DNA repair protein: the AdaO 6-methylguanine-DNA methyltransferase fromE. coli. EMBO J 13:1495–1501
Morohoshi F, Hayashi K, Munakata N (1990)Bacillus subtilis ada operon encodes two DNA alkyltransferase. Nucleic Acids Res 18:5473–5480
Morohoshi F, Hayashi K, Munakata N (1991) Molecular analysis ofBacillus subtilis ada mutants deficient in the adaptive response to simple alkylating agents. J Bacteriol 173:7834–7840
Myers LC, Terranova MP, Nash HM, Markus MA, Verdine GL (1992) Zinc binding by the methylation signaling domain of theEscherichia coli Ada protein. Biochemistry 31:4541–4547
Myers LC, Terranova MP, Ferentz AE, Wagner G, Verdine GL (1993) Repair of DNA methylphosphotriesters through a metallocativated cysteine nucleophile. Science 261:1164–1167
Myers LC, Jackow F, Verdine GL (1995) Metal dependence of transcriptional switching inEscherichia coli Ada. J Biol Chem 270:6664–6670
Nakabeppu Y, Sekiguchi M (1986) Regulatory mechanisms for induction of synthesis of repair enzymes in response to alkylating agents: Ada protein acts as a transcriptional regulator. Proc Natl Acad Sci USA 83:6297–6301
Nakabeppu Y, Kondo H, Kawabata S, Iwanaga S, Sekiguchi M (1985a) Purification and structure of the intact Ada regulatory protein ofEscherichia coli K12,O 6-methylguanine-DNA methyltransferase. J Biol Chem 260:7281–7288
Nakabeppu Y, Mine Y, Sekiguchi M (1985b) Regulation of expression of the clonedada gene inEscherichia coli. Mutat Res 146:155–167
Nakabeppu Y, Oda S, Sekiguchi M (1993) Proliferative activation of quiescent Rat-1A cells by ΔFosB. Mol Cell Biol 13:4157–4166
Nakamura T, Tokumoto Y, Sakumi K, Koike G, Nakabeppu Y, Sekiguchi M (1988) Expression of theada gene ofEscherichia coli in response to alkylating agents. Identification of transcriptional regulatory elements. J Mol Biol 202:483–494
Ohkubo T, Sakashita H, Sakuma T, Kainosho M, Sekiguchi M, Morikawa K (1994) Methylation dependent functional switch mechanism newly found in theEscherichia coli Ada protein. J Am Chem Soc 116:6035–6036
Sakashita H, Sakuma T, Ohkubo T, Kainosho M, Sakumi K, Sekiguchi M, Morikawa K (1993) Folding topology and DNA binding of the N-terminal fragment of Ada protein. FEBS Lett 323:252–256
Sakumi K, Sekiguchi M (1989) Regulation of expression of theada gene controlling the adaptive response. Interactions with theada promoter of the Ada protein and RNA polymerase. J Mol Biol 205:373–385
Samson L, Cairns J (1977) A new pathway for DNA repair inEscherichia coli. Nature 267:281–283
Sedgwick B (1989) In vitro proteolytic cleavage of theEscherichia coli Ada protein by theompT gene product. J Bacteriol 171:2249–2251
Sedgwick B, Robins P, Totty N, Lindahl T (1988) Functional domains and methyl acceptor sites of theEscherichia coli Ada protein. J Biol Chem 263:4430–4433
Sekiguchi M, Nakabeppu Y (1987) Adaptive response: induced synthesis of DNA repair enzymes by alkylating agents. Trends Genet 3:51–54
Shevell DE, Walker GC (1991) A region of the Ada DNA-repair protein required for the activation ofada transcription is not necessary for activation ofalkA. Proc Natl Acad Sci USA 88:9001–9005
Shevell DE, LeMotte PK, Walker GC (1988) Alteration of the carboxyl-terminal domain of Ada protein influences its inducibility, specificity, and strength as a transcriptional activator. J Bacteriol 170:5263–5271
Shevell DE, Friedman BM, Walker GC (1990) Resistance to alkylation damage inEscherichia coli: role of the Ada protein in induction of the adaptive response. Mutat Res 233:53–72
Takano K, Nakabeppu Y, Sekiguchi M (1988) Functional sites of the Ada regulatory protein ofEscherichia coli. Analysis by amino acid substitutions. J Mol Biol 201:261–271
Takano K, Nakamura T, Sekiguchi M (1991) Roles of two types ofO 6-methylguanine-DNA methyltransferases in DNA repair. Mutat Res 254:37–44
Teo I, Sedgwick B, Demple B, Li B, Lindahl T (1984) Induction of resistance to alkylating agents inE. coli: theada + gene product serves both as a regulatory protein and as an enzyme for repair of mutagenic damage. EMBO J 3:2151–2157
Teo I, Sedgwick B, Kilpatrick MW, McCarthy TV, Lindahl T (1986) The intracellular signal for induction of resistance to alkylating agents inE. coli. Cell 45:315–324
Vallee BL, Auld DS (1990) Zinc coordination, function, and structure of zinc enzymes and other proteins. Biochemistry 29:5647–5659
Weinfeld M, Drake AF, Saunders JK, Paterson MC (1985) Stereospecific removal of methyl phosphotriesters from DNA by anEscherichia coli ada + extract. Nucleic Acids Res 13:7067–7077
Yoshikai T, Nakabeppu Y, Sekiguchi M (1988) Proteolytic cleavage of Ada protein that carries methyltransferase and transcriptional regulator activities. J Biol Chem 263:19174–19180
Author information
Authors and Affiliations
Additional information
Communicated by R. Devoret
Rights and permissions
About this article
Cite this article
Taketomi, A., Nakabeppu, Y., Ihara, K. et al. Requirement for two conserved cysteine residues in the Ada protein ofEscherichia coli for transactivation of theada promoter. Molec. Gen. Genet. 250, 523–532 (1996). https://doi.org/10.1007/BF02174440
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02174440