Abstract
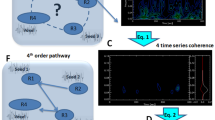
Since the middle of the 1990s, studies of resting-state fMRI/BOLD data have explored the correlation patterns of activity across the whole brain, which is referred to as functional connectivity (FC). Among the many methods that have been developed to interpret FC, a recently proposed model-based approach describes the propagation of fluctuating BOLD activity within the recurrently connected brain network by inferring the effective connectivity (EC). In this model, EC quantifies the strengths of directional interactions between brain regions, viewed from the proxy of BOLD activity. In addition, the tuning procedure for the model provides estimates for the local variability (input variances) to explain how the observed FC is generated. Generalizing, the network dynamics can be studied in the context of an input–output mapping—determined by EC—for the second-order statistics of fluctuating nodal activities. The present paper focuses on the following detection paradigm: observing output covariances, how discriminative is the (estimated) network model with respect to various input covariance patterns? An application with the model fitted to experimental fMRI data—movie viewing versus resting state—illustrates that changes in local variability and changes in brain coordination go hand in hand.



Similar content being viewed by others
References
Battaglia D, Witt A, Wolf F, Geisel T (2012) Dynamic effective connectivity of inter-areal brain circuits. PLoS Comput Biol 8:e1002438. https://doi.org/10.1371/journal.pcbi.1002438
Belliveau JW, Cohen MS, Weisskoff RM, Buchbinder BR, Rosen BR (1991) Functional studies of the human brain using high-speed magnetic resonance imaging. J Neuroimaging 1:36–41
Biswal B, Yetkin FZ, Haughton VM, Hyde JS (1995) Functional connectivity in the motor cortex of resting human brain using echo-planar MRI. Magn Reson Med 34:537–541
Bolt T, Prince EB, Nomi JS, Messinger D, Llabre MM, Uddin LQ (2017) Combining region- and network-level brain-behavior relationships in a structural equation model. Neuroimage 165:158–169. https://doi.org/10.1016/j.neuroimage.2017.10.007
Boynton G, Engel S, Glover G, Heeger D (1996) Linear systems analysis of functional magnetic resonance imaging in human V1. J Neurosci 16:4207–4221
Brookes MJ, Woolrich M, Luckhoo H, Price D, Hale JR, Stephenson MC, Barnes GR, Smith SM, Morris PG (2011) Investigating the electrophysiological basis of resting state networks using magnetoencephalography. Proc Natl Acad Sci USA 108:783–788. https://doi.org/10.1073/pnas.1112685108
Bullmore E, Sporns O (2009) Complex brain networks: graph theoretical analysis of structural and functional systems. Nat Rev Neurosci 10:186–198
Cabral J, Hugues E, Sporns O, Deco G (2011) Role of local network oscillations in resting-state functional connectivity. Neuroimage 57:130–139. https://doi.org/10.1016/j.neuroimage.2011.04.010
Chang C, Glover GH (2010) Time-frequency dynamics of resting-state brain connectivity measured with fMRI. Neuroimage 50:81–98. https://doi.org/10.1016/j.neuroimage.2009.12.011
Chang LJ, Gianaros PJ, Manuck SB, Krishnan A, Wager TD (2015) A sensitive and specific neural signature for picture-induced negative affect. PLoS Biol 13:e1002180. https://doi.org/10.1371/journal.pbio.1002180
Choi S, Amari S, Cichocki A (2000) Natural gradient learning for spatio-temporal decorrelation: recurrent network. IEICE Trans Fundamentals 83:2715–2722
Ciuciu P, Abry P, He BJ (2014) Interplay between functional connectivity and scale-free dynamics in intrinsic fMRI networks. Neuroimage 95:248–263. https://doi.org/10.1016/j.neuroimage.2014.03.047
Conturo TE, Lori NF, Cull TS, Akbudak E, Snyder AZ, Shimony JS, McKinstry RC, Burton H, Raichle ME (1999) Tracking neuronal fiber pathways in the living human brain. Proc Natl Acad Sci USA 96:422–427
Cordes D, Haughton V, Arfanakis K, Wendt G, Turski P, Moritz C, Quigley M, Meyerand M (2000) Mapping functionally related regions of brain with functional connectivity MR imaging. Am J Neuroradiol 21:1636–1644
Deco G, Jirsa VK, McIntosh AR (2011) Emerging concepts for the dynamical organization of resting-state activity in the brain. Nat Rev Neurosci 12:43–56. https://doi.org/10.1038/nrn2961
Deco G, Ponce-Alvarez A, Mantini D, Romani G, Hagmann P, Corbetta M (2013) Resting-state functional connectivity emerges from structurally and dynamically shaped slow linear fluctuations. J Neurosci 33:11239–11252. https://doi.org/10.1038/nrn2961
Deco G, Kringelbach ML (2014) Great expectations: using whole-brain computational connectomics for understanding neuropsychiatric disorders. Neuron 84:892–905. https://doi.org/10.1016/j.neuron.2014.08.034
Deco G, Tononi G, Boly M, Kringelbach ML (2015) Rethinking segregation and integration: contributions of whole-brain modelling. Nat Rev Neurosci 16:430–439. https://doi.org/10.1038/nrn3963
Deco G, Van Hartevelt T, Fernandes H, Stevner A, Kringelbach M (2017) The most relevant human brain regions for functional connectivity: evidence for a dynamical workspace of binding nodes from whole-brain computational modelling. Neuroimage 146:197–210
Engel AK, Gerloff C, Hilgetag CC, Nolte G (2013) Intrinsic coupling modes: multiscale interactions in ongoing brain activity. Neuron 80:867–886. https://doi.org/10.1016/j.neuron.2013.09.038
Fox MD, Raichle ME (2007) Spontaneous fluctuations in brain activity observed with functional magnetic resonance imaging. Nat Rev Neurosci 8:700–711. https://doi.org/10.1038/nrn2201
Freestone DR, Karoly PJ, Nešić D, Aram P, Cook MJ, Grayden DB (2014) Estimation of effective connectivity via data-driven neural modeling. Front Neurosci 28:383. https://doi.org/10.3389/fnins.2014.00383
Fries P (2015) Rhythms for cognition: communication through coherence. Neuron 88:220–235
Friston KJ, Mechelli A, Turner R, Price CJ (2000) Nonlinear responses in fMRI: the Balloon model, Volterra kernels, and other hemodynamics. Neuroimage 12:466–477. https://doi.org/10.1006/nimg.2000.0630
Friston KJ, Harrison L, Penny W (2003) Dynamic causal modelling. Neuroimage 19:1273–1302
Friston KJ (2011) Functional and effective connectivity: a review. Brain Connect 1:8
Friston KJ, Kahan J, Biswal B, Razi A (2014) A DCM for resting state fMRI. Neuroimage 94:396–407
Gilson M, Moreno-Bote R, Ponce-Alvarez A, Ritter P, Deco G (2016) Estimation of directed effective connectivity from fMRI functional connectivity hints at asymmetries of cortical connectome. PLoS Comput Biol 12:e1004762
Gilson M, Deco G, Friston K, Hagmann P, Mantini D, Betti V, Romani GL, Corbetta M (2017) Effective connectivity inferred from fMRI transition dynamics during movie viewing points to a balanced reconfiguration of cortical interactions. Neuroimage. https://doi.org/10.1016/j.neuroimage.2017.09.061
Goebel R, Roebroeck A, Kim DS, Formisano E (2003) Investigating directed cortical interactions in time-resolved fMRI data using vector autoregressive modeling and granger causality mapping. Magn Reson Imaging 21:1251–1261
Hagmann P, Cammoun L, Gigandet X, Meuli R, Honey CJ, Wedeen VJ, Sporns O (2008) Mapping the structural core of human cerebral cortex. PLoS Biol 6:e159. https://doi.org/10.1371/journal.pbio.0060159
Hall EL, Robson SE, Morris PG, Brookes MJ (2014) The relationship between MEG and fMRI. Neuroimage 102(Pt 1):80–91. https://doi.org/10.1016/j.neuroimage.2013.11.005
He BJ (2011) Scale-free properties of the functional magnetic resonance imaging signal during rest and task. J Neurosci 31:13786–13795
Heeger DJ, Ress D (2002) What does fMRI tell us about neuronal activity? Nat Rev Neurosci 3:142–151. https://doi.org/10.1038/nrn730
Honey CJ, Thivierge JP, Sporns O (2010) Can structure predict function in the human brain? Neuroimage 52:766–776. https://doi.org/10.1016/j.neuroimage.2010.01.071
Hutchison RM, Womelsdorf T, Allen EA, Bandettini PA, Calhoun VD, Corbetta M, Della Penna S, Duyn JH, Glover GH, Gonzalez-Castillo J, Handwerker DA, Keilholz S, Kiviniemi V, Leopold DA, de Pasquale F, Sporns O, Walter M, Chang C (2013) Dynamic functional connectivity: promise, issues, and interpretations. Neuroimage 80:360–378. https://doi.org/10.1016/j.neuroimage.2013.05.079
Iturria-Medina Y, Sotero RC, Canales-Rodríguez EJ, Alemán-Gómez Y, Melie-García L (2008) Studying the human brain anatomical network via diffusion-weighted MRI and graph theory. Neuroimage 40:1064–1076. https://doi.org/10.1016/j.neuroimage.2007.10.060
Linden DEJ, Turner DL (2016) Real-time functional magnetic resonance imaging neurofeedback in motor neurorehabilitation. Curr Opin Neurol 29:412–418. https://doi.org/10.1097/WCO.0000000000000340
Lütkepohl H (2005) New introduction to multiple time series analysis. Springer Science & Business Media, New York
Malsburg C (1981) The correlation theory of brain function. Tech. rep, Max Planck Institute for Biophysical Chemistry in Göttingen
Messé A, Rudrauf D, Benali H, Marrelec G (2014) Relating structure and function in the human brain: relative contributions of anatomy, stationary dynamics, and non-stationarities. PLoS Comput Biol 10:e1003530. https://doi.org/10.1371/journal.pcbi.1003530
Mehta-Pandejee G, Robinson PA, Henderson JA, Aquino KM, Sarkar S (2017) Inference of direct and multistep effective connectivities from functional connectivity of the brain and of relationships to cortical geometry. J Neurosci Methods 283:42–54
Mitra A, Snyder AZ, Hacker CD, Raichle ME (2014) Lag structure in resting-state fMRI. J Neurophysiol 111:2374–2391. https://doi.org/10.1152/jn.00804.2013
Mitra A, Snyder AZ, Tagliazucchi E, Laufs H, Raichle ME (2015) Propagated infra-slow intrinsic brain activity reorganizes across wake and slow wave sleep. Elife 4:e10781. https://doi.org/10.7554/eLife. 10781
Pallares V, Insabato A, Sanjuan A, Kühn S, Mantini D, Deco G, Gilson M (2017) Subject- and behavior-specific signatures extracted from fMRI data using whole-brain effective connectivity. biorxiv https://doi.org/10.1101/201624
Palmigiano A, Geisel T, Wolf F, Battaglia D (2017) Flexible information routing by transient synchrony. Nat Neurosci 20:1014–1022. https://doi.org/10.1038/nn.4569
Park HJ, Friston K (2013) Structural and functional brain networks: from connections to cognition. Science 342:1238411. https://doi.org/10.1126/science.1238411
Raichle ME, MacLeod AM, Snyder AZ, Powers WJ, Gusnard DA, Shulman GL (2001) A default mode of brain function. Proc Natl Acad Sci USA 98:676–682. https://doi.org/10.1073/pnas.98.2.676
Richardson M (2012) Large scale brain models of epilepsy: dynamics meets connectomics. J Neurol Neurosurg Psychiatry 83:1238–1248
Sala-Llonch R, Peña-Gómez C, Arenaza-Urquijo EM, Vidal-Piñeiro D, Bargalló N, Junqué C, Bartrés-Faz D (2012) Brain connectivity during resting state and subsequent working memory task predicts behavioural performance. Cortex 48(9):1187–1196. https://doi.org/10.1016/j.cortex.2011.07.006
Sanz Leon P, Knock SA, Woodman MM, Domide L, Mersmann J, McIntosh AR, Jirsa V (2013) The virtual brain: a simulator of primate brain network dynamics. Front Neuroinform 7:10. https://doi.org/10.3389/fninf.2013.00010
Schirner M, Rothmeier S, Jirsa VK, McIntosh AR, Ritter P (2015) An automated pipeline for constructing personalized virtual brains from multimodal neuroimaging data. Neuroimage 117:343–357. https://doi.org/10.1016/j.neuroimage.2015.03.055
Shen H (2014) Neuroscience: tuning the brain. Nature 507:290–292. https://doi.org/10.1038/507290a
Sporns O (2013) The human connectome: origins and challenges. Neuroimage 80:53–61. https://doi.org/10.1016/j.neuroimage.2013.03.023
Stephan KE, Mathys C (2014) Computational approaches to psychiatry. Curr Opin Neurobiol 25:85–92. https://doi.org/10.1016/j.conb.2013.12.007
Tzourio-Mazoyer N, Landeau B, Papathanassiou D, Crivello F, Etard O, Delcroix N, Mazoyer B, Joliot M (2002) Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage 15:273–289. https://doi.org/10.1006/nimg.2001.0978
Uhlhaas P, Singer W (2006) Neural synchrony in brain disorders: relevance for cognitive dysfunctions and pathophysiology. Neuron 52:155–168
Valdes-Sosa PA, Roebroeck A, Daunizeau J, Friston K (2011) Effective connectivity: influence, causality and biophysical modeling. Neuroimage 58:339–361. https://doi.org/10.1016/j.neuroimage.2011.03.058
Zamora-López G, Chen Y, Deco G, Kringelbach ML, Zhou C (2016) Functional complexity emerging from anatomical constraints in the brain: the significance of network modularity and rich-clubs. Sci Rep 6:38424. https://doi.org/10.1038/srep38424
Zamora-López G, Zhou C, Kurths J (2011) Exploring brain function from anatomical connectivity. Front Neurosci 5:83. https://doi.org/10.3389/fnins.2011.00083
Acknowledgements
The author thanks Moritz Deger and Martin Nawrot for organizing the 12th International Neural Coding Workshop, NC2016. The author is also grateful to Pierre Yger, Ruben Moreno-Bote, Vicente Pallarez, Andrea Insabato, Gustavo Deco and Morten Kringelbach for constructive discussions.
Author information
Authors and Affiliations
Corresponding author
Additional information
MG acknowledges funding from the Marie Sklodowska-Curie Action (Grant H2020-MSCA-656547) and the Human Brain Project (ramp-up phase, Grant FP7-FET-ICT-604102).
This article belongs to a Special Issue on Neural Coding.
Rights and permissions
About this article
Cite this article
Gilson, M. Analysis of fMRI data using noise-diffusion network models: a new covariance-coding perspective. Biol Cybern 112, 153–161 (2018). https://doi.org/10.1007/s00422-017-0741-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00422-017-0741-y