Abstract
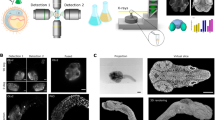
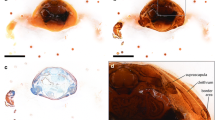
We describe a new methodology for rapid 2D and 3D computer analysis and visualisation of gene expression and gene product pattern in the context of anatomy and tissue architecture. It is based on episcopic imaging of embryos and tissue samples, as they are physically sectioned, thereby producing inherently aligned digital image series and volume data sets, which immediately permit the generation of 3D computer representations. The technique uses resin as embedding medium, eosin for unspecific tissue staining, and colour reactions (β-galactosidase/Xgal or BCIP/NBT) for specific labelling of gene activity and mRNA pattern. We tested the potential of the method for producing high-resolution volume data sets of adult human and porcine tissue samples and of specifically and unspecifically stained mouse, chick, quail, frog, and zebrafish embryos. The quality of the episcopic images resembles the quality of digital images of true histological sections with respect to resolution and contrast. Specifically labelled structures can be extracted using simple thresholding algorithms. Thus, the method is capable of quickly and precisely detecting molecular signals simultaneously with anatomical details and tissue architecture. It has no tissue restrictions and can be applied for analysis of human tissue samples as well as for analysis of all developmental stages of embryos of a wide variety of biomedically relevant species.






Similar content being viewed by others
References
Blasberg RG, Tjuvajev JG (2003) Molecular-genetic imaging: current and future perspectives. J Clin Invest 111(11):1620–1629
Costa LdaF, Barbosa MS, Manoel ET, Streicher J, Müller GB (2004) Mathematical characterization of three-dimensional gene expression patterns. Bioinformatics 20(11):1653–1662
Costa LdaF, Beletti ME, Müller GB, Rasskin-Gutman D, Sternik G, Ibanes M, Belmonte JCI (2005) Field approach to three-dimensional gene expression pattern characterization. Appl Phys Lett 86(14):143901
Denk W, Horstmann H (2004) Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLoS Biol 2(11):e329
Ewald AJ, McBride H, Reddington M, Fraser SE, Kerschmann R (2002) Surface imaging microscopy, an automated method for visualizing whole embryo samples in three dimensions at high resolution. Dev Dyn 225(3):369–375
Hamburger V, Hamilton HL (1992) A series of normal stages in the development of the chick embryo. 1951. Dev Dyn 195(4):231–272
Herschman HR (2003) Molecular imaging: looking at problems, seeing solutions. Science 302(5645):605–608
Huisken J, Swoger J, Del Bene F, Wittbrodt J, Stelzer EH (2004) Optical sectioning deep inside live embryos by selective plane illumination microscopy. Science 305(5686):1007–1009
Jacobs RE, Fraser SE (1994) Magnetic resonance microscopy of embryonic cell lineages and movements. Science 263(5147):681–684
Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF (1995) Stages of embryonic development of the zebrafish. Dev Dyn 203(3):253–310
Kolker SJ, Tajchman U, Weeks DL (2000) Confocal imaging of early heart development in Xenopus laevis. Dev Biol 218(1):64–73
Louie AY, Huber MM, Ahrens ET, Rothbacher U, Moats R, Jacobs RE, Fraser SE, Meade TJ (2000) In vivo visualization of gene expression using magnetic resonance imaging. Nat Biotechnol 18(3):321–325
Massoud TF, Gambhir SS (2003) Molecular imaging in living subjects: seeing fundamental biological processes in a new light. Genes Dev 17(5):545–580
Matsui T, Raya A, Kawakami Y, Callol-Massot C, Capdevila J, Rodriguez-Esteban C, Izpisua Belmonte JC (2005) Noncanonical Wnt signaling regulates midline convergence of organ primordia during zebrafish development. Genes Dev 19(1):164–175
Ng JK, Kawakami Y, Buscher D, Raya A, Itoh T, Koth CM, Rodriguez Esteban C, Rodriguez-Leon J, Garrity DM, Fishman MC, Izpisua Belmonte JC (2002) The limb identity gene Tbx5 promotes limb initiation by interacting with Wnt2b and Fgf10. Development 129(22):5161–5170
Schneider JE, Bamforth SD, Farthing CR, Clarke K, Neubauer S, Bhattacharya S (2003) High-resolution imaging of normal anatomy, and neural and adrenal malformations in mouse embryos using magnetic resonance microscopy. J Anat 202(2):239–247
Sharpe J, Ahlgren U, Perry P, Hill B, Ross A, Hecksher-Sørensen J, Baldock R, Davidson D (2002) Optical projection tomography as a tool for 3D microscopy and gene expression studies. Science 296(5567):541–545
Streicher J, Weninger WJ, Müller GB (1997) External marker-based automatic congruencing: a new method of 3D reconstruction from serial sections. Anat Rec 248(4):583–602
Streicher J, Donat MA, Strauss B, Sporle R, Schughart K, Müller GB (2000) Computer-based three-dimensional visualization of developmental gene expression. Nat Genet 25(2):147–152
Weissleder R, Ntziachristos V (2003) Shedding light onto live molecular targets. Nat Med 9(1):123–128
Weninger WJ, Mohun T (2002) Phenotyping transgenic embryos: a rapid 3-D screening method based on episcopic fluorescence image capturing. Nat Genet 30(1):59–65
Weninger WJ, Meng S, Streicher J, Müller GB (1998) A new episcopic method for rapid 3-D reconstruction: applications in anatomy and embryology. Anat Embryol (Berl) 197(5):341–348
Acknowledgements
The authors are grateful to M. Knapp, E. Gröller, K. Dorfmeister, G.M. Gruber, B. Maurer, and M.T. Nödl for technical support. This work was supported by the HFSP grant, RGP 0039/2002-C203.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Weninger, W.J., Geyer, S.H., Mohun, T.J. et al. High-resolution episcopic microscopy: a rapid technique for high detailed 3D analysis of gene activity in the context of tissue architecture and morphology. Anat Embryol 211, 213–221 (2006). https://doi.org/10.1007/s00429-005-0073-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00429-005-0073-x