Abstract
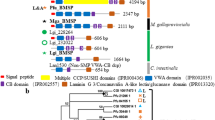
Various novel proteins have been identified from many kinds of mollusk shells. Although such matrix proteins are believed to play important roles in the calcium carbonate crystal formation of shells, no common proteins that interact with calcium carbonate or that are involved in the molecular mechanisms behind shell formation have been identified. Pif consists of two proteins, Pif 80 and Pif 97, which are encoded by a single mRNA. Pif 80 was identified as a key acidic protein that regulates the formation of the nacreous layer in Pinctada fucata, while Pif 97 has von Willebrand factor type A (VWA) and chitin-binding domains. In this study, we identified Pif homologues from Pinctada margaritifera, Pinctada maxima, Pteria penguin, Mytilus galloprovincialis, and in the genome database of Lottia gigantea in order to compare their primary protein sequences. The VWA and chitin-binding domains are conserved in all Pif 97 homologues, whereas the amino acid sequences of the Pif 80 regions differ markedly among the species. Sequence alignment revealed the presence of a novel significantly conserved sequence between the chitin-binding domain and the C-terminus of Pif 97. Further examination of the Pif 80 regions suggested that they share a sequence that is similar to the laminin G domain. These results indicate that all Pif molecules in bivalves and gastropods may be derived from a common ancestral gene. These comparisons may shed light on the correlation between molecular evolution and morphology in mollusk shell microstructure.








Similar content being viewed by others
References
Belcher AM, Wu XH, Christensen RJ, Hansma PK, Stucky GD, Morse DE (1996) Control of crystal phase switching and orientation by soluble mollusc-shell proteins. Nature 381:56–58
Berland S, Marie A, Duplat D, Milet C, Sire JY, Bedouet L (2011) Coupling proteomics and transcriptomics for the identification of novel and variant forms of mollusk shell proteins: a study with P. margaritifera. Chembiochem 12:950–961
Burgess TL, Kelly RB (1987) Constitutive and regulated secretion of proteins. Annu Rev Cell Biol 3:243–293
Elvin CM, Vuocolo T, Pearson RD, East IJ, Riding GA, Eisemann CH, Tellam RL (1996) Characterization of a major peritrophic membrane protein, peritrophin-44, from the larvae of Lucilia cuprina. cDNA and deduced amino acid sequences. J Biol Chem 271:8925–8935
Falini G, Albeck S, Weiner S, Addadi L (1996) Control of aragonite or calcite polymorphism by mollusk shell macromolecules. Science 271:67–69
Fuchigami T, Sasaki T (2005) The shell structure of the recent Patellogastropoda (Mollusca: Gastropoda). Paleontol Res 9:143–168
Gotliv BA, Kessler N, Sumerel JL, Morse DE, Tuross N, Addadi L, Weiner S (2005) Asprich: a novel aspartic acid-rich protein family from the prismatic shell matrix of the bivalve Atrina rigida. Chembiochem 6:304–314
Gregoire C (1957) Topography of the organic components in mother-of pearl. J Biophys Biochem Cytol 3:797–808
Hohenester E, Tisi D, Talts JF, Timpl R (1999) The crystal structure of a laminin G-like module reveals the molecular basis of alpha-dystroglycan binding to laminins, perlecan, and agrin. Mol Cell 4:783–792
Joubert C, Piquemal D, Marie B, Manchon L, Pierrat F, Zanella-Cleon I, Cochennec-Laureau N, Gueguen Y, Montagnani C (2010) Transcriptome and proteome analysis of Pinctada margaritifera calcifying mantle and shell: focus on biomineralization. BMC Genomics 11:613–625
Kawabata S, Nagayama R, Hirata M, Shigenaga T, Agarwala KL, Saito T, Cho J, Nakajima H, Takagi T, Iwanaga S (1996) Tachycitin, a small granular component in horseshoe crab hemocytes, is an antimicrobial protein with chitin-binding activity. J Biochem 120:1253–1260
Kono M, Hayashi N, Samata T (2000) Molecular mechanism of the nacreous layer formation in Pinctada maxima. Biochem Biophys Res Commun 269:213–218
Lee JO, Rieu P, Arnaout MA, Liddington R (1995) Crystal structure of the A domain from the alpha subunit of integrin CR3 (CD11b/CD18). Cell 80:631–638
Mann K, Siedler F, Treccani L, Heinemann F, Fritz M (2007) Perlinhibin, a cysteine-, histidine-, and arginine-rich miniprotein from abalone (Haliotis laevigata) nacre, inhibits in vitro calcium carbonate crystallization. Biophys J 93:1246–1254
Marie B, Le Roy N, Zanella-Cleon I, Becchi M, Marin F (2011) Molecular evolution of mollusc shell proteins: insights from proteomic analysis of the edible mussel Mytilus. J Mol Evol 72:531–546
Marin F, Corstjens P, de Gaulejac B, de Vrind-de JE, Westbroek P (2000) Mucins and molluscan calcification. Molecular characterization of mucoperlin, a novel mucin-like protein from the nacreous shell layer of the fan mussel Pinna nobilis (Bivalvia, Pteriomorphia). J Biol Chem 275:20667–20675
Marshall RC, Gillespie JM (1982) The tryptophan-rich keratin protein fraction of claws of the lizard Varanus gouldii. Comp Biochem Physiol B 71:623–628
Marxen JC, Nimtz M, Becker W, Mann K (2003) The major soluble 19.6 kDa protein of the organic shell matrix of the freshwater snail Biomphalaria glabrata is an N-glycosylated dermatopontin. Biochim Biophys Acta 1650:92–98
Matsushiro A, Miyashita T, Miyamoto H, Morimoto K, Tonomura B, Tanaka A, Sato K (2003) Presence of protein complex is prerequisite for aragonite crystallization in the nacreous layer. Mar Biotechnol (N Y) 5:37–44
Michenfelder M, Fu G, Lawrence C, Weaver JC, Wustman BA, Taranto L, Evans JS, Morse DE (2003) Characterization of two molluscan crystal-modulating biomineralization proteins and identification of putative mineral binding domains. Biopolymers 70:522–533
Miyamoto H, Miyashita T, Okushima M, Nakano S, Morita T, Matsushiro A (1996) A carbonic anhydrase from the nacreous layer in oyster pearls. Proc Natl Acad Sci USA 93:9657–9660
Miyamoto H, Yano M, Miyashita T (2003) Similarities in the structure of nacrein, the shell-matrix protein, in a bivalve and a gastropod. J Moll Stud 69:87–89
Miyashita T, Takagi R, Okushima M, Nakano S, Miyamoto H, Nishikawa E, Matsushiro A (2000) Complementary DNA cloning and characterization of pearlin, a new class of matrix protein in the nacreous layer of oyster pearls. Mar Biotechnol (N Y) 2:409–418
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–25
Samata T, Hayashi N, Kono M, Hasegawa K, Horita C, Akera S (1999) A new matrix protein family related to the nacreous layer formation of Pinctada fucata. FEBS Lett 462:225–229
Sarashina I, Yamaguchi H, Haga T, Iijima M, Chiba S, Endo K (2006) Molecular evolution and functionally important structures of molluscan dermatopontin: implications for the origins of molluscan shell matrix proteins. J Mol Evol 62:307–318
Shen Z, Jacobs-Lorena M (1999) Evolution of chitin-binding proteins in invertebrates. J Mol Evol 48:341–347
Shen X, Belcher AM, Hansma PK, Stucky GD, Morse DE (1997) Molecular cloning and characterization of lustrin A, a matrix protein from shell and pearl nacre of Haliotis rufescens. J Biol Chem 272:32472–32481
Smirnov SP, McDearmon EL, Li S, Ervasti JM, Tryggvason K, Yurchenco PD (2002) Contributions of the LG modules and furin processing to laminin-2 functions. J Biol Chem 277:18928–18937
Sudo S, Fujikawa T, Nagakura T, Ohkubo T, Sakaguchi K, Tanaka M, Nakashima K, Takahashi T (1997) Structures of mollusc shell framework proteins. Nature 387:563–564
Suetake T, Tsuda S, Kawabata S, Miura K, Iwanaga S, Hikichi K, Nitta K, Kawano K (2000) Chitin-binding proteins in invertebrates and plants comprise a common chitin-binding structural motif. J Biol Chem 275:17929–17932
Suzuki M, Saruwatari K, Kogure T, Yamamoto Y, Nishimura T, Kato T, Nagasawa H (2009) An acidic matrix protein, Pif, is a key macromolecule for nacre formation. Science 325:1388–1390
Suzuki M, Kameda J, Sasaki T, Saruwatari K, Nagasawa H, Kogure T (2010) Characterization of the multilayered shell of a limpet, Lottia kogamogai (Mollusca: Patellogastropoda), using SEM-EBSD and FIB-TEM techniques. J Struct Biol 171:223–230
Suzuki M, Iwashima A, Tsutsui N, Ohira T, Kogure T, Nagasawa H (2011) Identification and characterisation of a calcium carbonate-binding protein, blue mussel shell protein (BMSP), from the nacreous layer. Chembiochem 12:2478–2487
Takahashi K, Yamamoto H, Onoda A, Doi M, Inaba T, Chiba M, Kobayashi A, Taguchi T, Okamura T, Ueyama N (2004) Highly oriented aragonite nanocrystal–biopolymer composites in an aragonite brick of the nacreous layer of Pinctada fucata. Chemm Comm 8:996–997
Talts JF, Timpl R (1999) Mutation of a basic sequence in the laminin alpha2LG3 module leads to a lack of proteolytic processing and has different effects on beta1 integrin-mediated cell adhesion and alpha-dystroglycan binding. FEBS Lett 458:319–323
Talts JF, Mann K, Yamada Y, Timpl R (1998) Structural analysis and proteolytic processing of recombinant G domain of mouse laminin alpha2 chain. FEBS Lett 426:71–76
Temkin I (2010) Molecular phylogeny of pearl oysters and their relatives (Mollusca, Bivalvia, Pterioidea). BMC Evol Biol 10:342–369
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–80
Treccani L, Mann K, Heinemann F, Fritz M (2006) Perlwapin, an abalone nacre protein with three four-disulfide core (whey acidic protein) domains, inhibits the growth of calcium carbonate crystals. Biophys J 91:2601–2608
Tryggvason K (1993) The laminin family. Curr Opin Cell Biol 5:877–882
Tsukamoto D, Sarashina I, Endo K (2004) Structure and expression of an unusually acidic matrix protein of pearl oyster shells. Biochem Biophys Res Commun 320:1175–1180
Tuckwell D (1999) Evolution of von Willebrand factor A (VWA) domains. Biochem Soc Trans 27:835–840
Wada K (1961) Crystal growth of molluscan shells. Bull Nat Pearl Res Lab 7:703–785
Weiner S, Hood L (1975) Soluble protein of the organic matrix of mollusk shells: a potential template for shell formation. Science 190:987–9
Weiss IM, Kaufmann S, Mann K, Fritz M (2000) Purification and characterization of perlucin and perlustrin, two new proteins from the shell of the mollusc Haliotis laevigata. Biochem Biophys Res Commun 267:17–21
Whittaker CA, Hynes RO (2002) Distribution and evolution of von Willebrand/integrin A domains: widely dispersed domains with roles in cell adhesion and elsewhere. Mol Biol Cell 13:3369–3387
Yano M, Nagai K, Morimoto K, Miyamoto H (2007) A novel nacre protein N19 in the pearl oyster Pinctada fucata. Biochem Biophys Res Commun 362:158–163
Yu H, Talts JF (2003) Beta1 integrin and alpha-dystroglycan binding sites are localized to different laminin-G-domain-like (LG) modules within the laminin alpha5 chain G domain. Biochem J 371:289–299
Yurchenco PD, Sung U, Ward MD, Yamada Y, O'Rear JJ (1993) Recombinant laminin G domain mediates myoblast adhesion and heparin binding. J Biol Chem 268:8356–8365
Acknowledgments
We are grateful to Tasaki Shinju Co. Ltd in Hyogo Prefecture, Japan, for providing us with the shells and live oysters of P. margaritifera, P. maxima, and P. penguin. We are grateful to Dr. James Weaver, Dr. Steve Weiner, and Dr. Lia Addadi for providing the shells of L. gigantea. This work was supported by Grants-in-Aid for Scientific Research (17GS0311, 22248037, and 22228006) from the Japan Society for the Promotion of Science (JSPS). M.S. was supported by a Research Fellowship of JSPS for young scientists.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. 1a
Sequence strategy for determination of the complete cDNA sequence for each Pif homologues. (a) pmPif. (b) pmxPif. (c) ppPif. The sequence of each primer number is shown in supplementary Table 1. (JPEG 39 kb)
Fig. 1b
(JPEG 36 kb)
Fig. 1c
(JPEG 35 kb)
Fig. 2a
The complete cDNA and deduced amino acid sequence of pmPif. (b) The complete cDNA and deduced amino acid sequence of pmxPif. (c) The complete cDNA and deduced amino acid sequence of ppPif. Orange box is a signal peptide. Blue box indicates the VWA domain. Yellow box is the Peritrophin A-type chitin-binding domain. Purple box is BXBB (B means a basic amino acid residue) that is a Kex2-like protease cleavage site. Green box is the sequence that has the similarity of aragonite-binding protein (Pif 80). The underlines indicate the polyadenylation signal sequences. (JPEG 241 kb)
Fig. 2b
(JPEG 228 kb)
Fig. 2c
(JPEG 476 kb)
Table 1
(DOCX 22 kb)
Rights and permissions
About this article
Cite this article
Suzuki, M., Iwashima, A., Kimura, M. et al. The Molecular Evolution of the Pif Family Proteins in Various Species of Mollusks. Mar Biotechnol 15, 145–158 (2013). https://doi.org/10.1007/s10126-012-9471-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-012-9471-2