Abstract
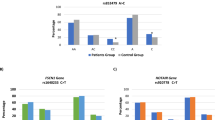
Deleterious mutations in several genes that are involved in repair of damage to DNA have been associated with an increased risk of breast cancer. Recent studies have shown sequence variants in two such genes, RAD50 and NBS1, which can be predisposed to breast cancer. The aim of this study is to elucidate the contribution of RAD50 and NBS1 germline mutations to the etiology of non-BRCA1/2 hereditary breast cancer in China. We conducted a mutational analysis of RAD50 and NBS1 in genomic DNA from 384 Chinese women with early-onset breast cancer and/or affected relatives. All the coding exons and adjacent intronic splice junction rejoins of RAD50 and NBS1 were screened using PCR-DHPLC and DNA sequencing analysis. Among all cases, no obviously deleterious mutations were observed in RAD50; one synonymous change c.102G>A at codon 34 and one single nucleotide polymorphism IVS9 + 19C>T were identified in NBS1. Furthermore, there was no remarkable difference in the allele frequency of NBS1 c.553G>C (E185Q) between cases (172/384) and controls (182/420). Our results exclude the possible role of RAD50 and NBS1 in familial breast cancer predisposition in Chinese women, and there is no evidence for the recommendation of RAD50 and NBS1 for genetic testing in China.
Similar content being viewed by others
References
Rahman N, Stratton MR (1998) The genetics of breast cancer susceptibility. Annu Rev Genet 32:95–121
Walsh T, King MC (2007) Ten genes for inherited breast cancer. Cancer Cell 11:103–105
Zhu XD, Kuster B, Mann M, Petrini JH, de Lange T (2000) Cell-cycle-regulated association of RAD50/MRE11/NBS1 with TRF2 and human telomeres. Nat Genet 25:347–352
Stracker TH, Theunissen JW, Morales M, Petrini JH (2004) The Mre11 complex and the metabolism of chromosome breaks: the importance of communicating and holding things together. DNA Repair 3:845–854
Bartkova J, Tommiska J, Oplustilova L, Aaltonen K, Tamminen A, Heikkinen T, Mistrik M, Aittomaki K, Blomqvist C, Heikkila P, Lukas J, Nevanlinna H, Bartek J (2008) Aberrations of the MRE11-RAD50-NBS1 DNA damage sensor complex in human breast cancer: MRE11 as a candidate familial cancer-predisposing gene. Mol Oncol 2:296–316
Falck J, Coates J, Jackson SP (2005) Conserved modes of recruitment of ATM, ATR and DNA-PKcs to sites of DNA damage. Nature 434:605–611
Heikkinen K, Rapakko K, Karppinen SM, Erkko H, Knuutila S, Lundan T, Mannermaa A, Borresen-Dale AL, Borg A, Barkardottir RB, Petrini J, Winqvist R (2006) RAD50 and NBS1 are breast cancer susceptibility genes associated with genomic instability. Carcinogenesis 27:1593–1599
Bogdanova N, Feshchenko S, Schurmann P, Waltes R, Wieland B, Hillemanns P, Rogov YI, Dammann O, Bremer M, Karstens JH, Sohn C, Varon R, Dork T (2008) Nijmegen breakage syndrome mutations and risk of breast cancer. Int J Cancer 122:802–806
Wang X, Szabo C, Qian C, Amadio PG, Thibodeau SN, Cerhan JR, Petersen GM, Liu W, Couch FJ (2008) Mutational analysis of thirty-two double-strand DNA break repair genes in breast and pancreatic cancers. Cancer Res 68:971–975
Cao AY, Hu Z, Yin WJ, Jin W, Shao ZM (2010) Some common mutations of RAD50 and NBS1 in western populations do not contribute significantly to Chinese non-BRCA1/2 hereditary breast cancer. Breast Cancer Res Treat 121:247–249
Tommiska J, Seal S, Renwick A, Barfoot R, Baskcomb L, Jayatilake H, Bartkova J, Tallila J, Kaare M, Tamminen A, Heikkila P, Evans DG, Eccles D, Aittomaki K, Blomqvist C, Bartek J, Stratton MR, Nevanlinna H, Rahman N (2006) Evaluation of RAD50 in familial breast cancer predisposition. Int J Cancer 118:2911–2916
Buslov KG, Iyevleva AG, Chekmariova EV, Suspitsin EN, Togo AV, Kuligina E, Sokolenko AP, Matsko DE, Turkevich EA, Lazareva YR, Chagunava OL, Bit-Sava EM, Semiglazov VF, Devilee P, Cornelisse C, Hanson KP, Imyanitov EN (2005) NBS1 657del5 mutation may contribute only to a limited fraction of breast cancer cases in Russia. Int J Cancer 114:585–589
Uhrhammer N, Delort L, Bignon YJ (2009) Rad50 c.687delT does not contribute significantly to familial breast cancer in a French population. Cancer Epidemiol Biomarkers Prev 18:684–685
Mosor M, Ziolkowska-Suchanek I, Roznowski K, Baranowska M, Januszkiewicz-Lewandowska D, Nowak J (2010) RAD50 gene mutations are not likely a risk factor for breast cancer in Poland. Breast Cancer Res Treat 123:607–609
Steffen J, Nowakowska D, Niwinska A, Czapczak D, Kluska A, Piatkowska M, Wisniewska A, Paszko Z (2006) Germline mutations 657del5 of the NBS1 gene contribute significantly to the incidence of breast cancer in Central Poland. Int J Cancer 119:472–475
Gorski B, Debniak T, Masojc B, Mierzejewski M, Medrek K, Cybulski C, Jakubowska A, Kurzawski G, Chosia M, Scott R, Lubinski J (2003) Germline 657del5 mutation in the NBS1 gene in breast cancer patients. Int J Cancer 106:379–381
Gorski B, Cybulski C, Huzarski T, Byrski T, Gronwald J, Jakubowska A, Stawicka M, Gozdecka-Grodecka S, Szwiec M, Urbanski K, Mitus J, Marczyk E, Dziuba J, Wandzel P, Surdyka D, Haus O, Janiszewska H, Debniak T, Toloczko-Grabarek A, Medrek K, Masojc B, Mierzejewski M, Kowalska E, Narod SA, Lubinski J (2005) Breast cancer predisposing alleles in Poland. Breast Cancer Res Treat 92:19–24
Carlomagno F, Chang-Claude J, Dunning AM, Ponder BA (1999) Determination of the frequency of the common 657Del5 Nijmegen breakage syndrome mutation in the German population: no association with risk of breast cancer. Genes Chromosomes Cancer 25:393–395
Roznowski K, Januszkiewicz-Lewandowska D, Mosor M, Pernak M, Litwiniuk M, Nowak J (2008) I171V germline mutation in the NBS1 gene significantly increases risk of breast cancer. Breast Cancer Res Treat 110:343–348
Bogdanova N, Schurmann P, Waltes R, Feshchenko S, Zalutsky IV, Bremer M, Dork T (2008) NBS1 variant I171V and breast cancer risk. Breast Cancer Res Treat 112:75–79
Desjardins S, Beauparlant JC, Labrie Y, Ouellette G, Durocher F (2009) Variations in the NBN/NBS1 gene and the risk of breast cancer in non-BRCA1/2 French Canadian families with high risk of breast cancer. BMC Cancer 9:181
Steffen J, Varon R, Mosor M, Maneva G, Maurer M, Stumm M, Nowakowska D, Rubach M, Kosakowska E, Ruka W, Nowecki Z, Rutkowski P, Demkow T, Sadowska M, Bidzinski M, Gawrychowski K, Sperling K (2004) Increased cancer risk of heterozygotes with NBS1 germline mutations in Poland. Int J Cancer 111:67–71
Kobayashi J, Antoccia A, Tauchi H, Matsuura S, Komatsu K (2004) NBS1 and its functional role in the DNA damage response. DNA Repair 3:855–861
Lu J, Wei Q, Bondy ML, Li D, Brewster A, Shete S, Yu TK, Sahin A, Meric-Bernstam F, Hunt KK, Singletary SE, Ross MI, Wang LE (2006) Polymorphisms and haplotypes of the NBS1 gene are associated with risk of sporadic breast cancer in non-Hispanic white women < or = 55 years. Carcinogenesis 27:2209–2216
Smith TR, Levine EA, Freimanis RI, Akman SA, Allen GO, Hoang KN, Liu-Mares W, Hu JJ (2008) Polygenic model of DNA repair genetic polymorphisms in human breast cancer risk. Carcinogenesis 29:2132–2138
Kuschel B, Auranen A, McBride S, Novik KL, Antoniou A, Lipscombe JM, Day NE, Easton DF, Ponder BA, Pharoah PD, Dunning A (2002) Variants in DNA double-strand break repair genes and breast cancer susceptibility. Hum Mol Genet 11:1399–1407
Forsti A, Angelini S, Festa F, Sanyal S, Zhang Z, Grzybowska E, Pamula J, Pekala W, Zientek H, Hemminki K, Kumar R (2004) Single nucleotide polymorphisms in breast cancer. Oncol Rep 11:917–922
Silva SN, Tomar M, Paulo C, Gomes BC, Azevedo AP, Teixeira V, Pina JE, Rueff J, Gaspar JF (2010) Breast cancer risk and common single nucleotide polymorphisms in homologous recombination DNA repair pathway genes XRCC2, XRCC3, NBS1 and RAD51. Cancer Epidemiol 34:85–92
Zhang L, Zhang Z, Yan W (2005) Single nucleotide polymorphisms for DNA repair genes in breast cancer patients. Clin Chim Acta 359:150–155
Gorski B, Jakubowska A, Huzarski T, Byrski T, Gronwald J, Grzybowska E, Mackiewicz A, Stawicka M, Bebenek M, Sorokin D, Fiszer-Maliszewska L, Haus O, Janiszewska H, Niepsuj S, Gozdz S, Zaremba L, Posmyk M, Pluzanska M, Kilar E, Czudowska D, Wasko B, Miturski R, Kowalczyk JR, Urbanski K, Szwiec M, Koc J, Debniak B, Rozmiarek A, Debniak T, Cybulski C, Kowalska E, Toloczko-Grabarek A, Zajaczek S, Menkiszak J, Medrek K, Masojc B, Mierzejewski M, Narod SA, Lubinski J (2004) A high proportion of founder BRCA1 mutations in Polish breast cancer families. Int J Cancer 110:683–686
Li WF, Hu Z, Rao NY, Song CG, Zhang B, Cao MZ, Su FX, Wang YS, He PQ, Di GH, Shen KW, Wu J, Lu JS, Luo JM, Liu XY, Zhou J, Wang L, Zhao L, Liu YB, Yuan WT, Yang L, Shen ZZ, Huang W, Shao ZM (2008) The prevalence of BRCA1 and BRCA2 germline mutations in high-risk breast cancer patients of Chinese Han nationality: two recurrent mutations were identified. Breast Cancer Res Treat 110:99–109
Meijers-Heijboer H, van den Ouweland A, Klijn J, Wasielewski M, de Snoo A, Oldenburg R, Hollestelle A, Houben M, Crepin E, van Veghel-Plandsoen M, Elstrodt F, van Duijn C, Bartels C, Meijers C, Schutte M, McGuffog L, Thompson D, Easton D, Sodha N, Seal S, Barfoot R, Mangion J, Chang-Claude J, Eccles D, Eeles R, Evans DG, Houlston R, Murday V, Narod S, Peretz T, Peto J, Phelan C, Zhang HX, Szabo C, Devilee P, Goldgar D, Futreal PA, Nathanson KL, Weber B, Rahman N, Stratton MR (2002) Low-penetrance susceptibility to breast cancer due to CHEK2(*)1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nat Genet 31:55–59
Vahteristo P, Bartkova J, Eerola H, Syrjakoski K, Ojala S, Kilpivaara O, Tamminen A, Kononen J, Aittomaki K, Heikkila P, Holli K, Blomqvist C, Bartek J, Kallioniemi OP, Nevanlinna H (2002) A CHEK2 genetic variant contributing to a substantial fraction of familial breast cancer. Am J Hum Genet 71:432–438
Offit K, Pierce H, Kirchhoff T, Kolachana P, Rapaport B, Gregersen P, Johnson S, Yossepowitch O, Huang H, Satagopan J, Robson M, Scheuer L, Nafa K, Ellis N (2003) Frequency of CHEK2*1100delC in New York breast cancer cases and controls. BMC Med Genet 4:1
Osorio A, Rodriguez-Lopez R, Diez O, de la Hoya M, Ignacio MJ, Vega A, Esteban-Cardenosa E, Alonso C, Caldes T, Benitez J (2004) The breast cancer low-penetrance allele 1100delC in the CHEK2 gene is not present in Spanish familial breast cancer population. Int J Cancer 108:54–56
Jekimovs CR, Chen X, Arnold J, Gatei M, Richard DJ, Spurdle AB, Khanna KK, Chenevix-Trench G (2005) Low frequency of CHEK2 1100delC allele in Australian multiple-case breast cancer families: functional analysis in heterozygous individuals. Br J Cancer 92:784–790
Song CG, Hu Z, Yuan WT, Di GH, Shen ZZ, Huang W, Shao ZM (2006) CHEK2 c.1100delC may not contribute to genetic background of hereditary breast cancer from Shanghai of China. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 23:443–445
Collaborative Group on Hormonal Factors in Breast Cancer (2001) Familial breast cancer: collaborative reanalysis of individual data from 52 epidemiological studies including 58, 209 women with breast cancer and 101, 986 women without the disease. Lancet 358:1389–1399
Berry DA (2001) Role of population-based studies in assessing genetic cancer risk. J Natl Cancer Inst 93:1188–1189
Xiao W, Oefner PJ (2001) Denaturing high-performance liquid chromatography: a review. Hum Mutat 17:439–474
Single nucleotide polymorphisms [http://www.ncbi.nlm.nih.gov/SNP/]
Acknowledgments
We thank all the family members for their willingness to cooperate with our study. This research was supported in part by the grants from the National Basic Research Program of China (2006CB910501), National Natural Science Foundation of China (30371580, 30572109); Shanghai Science and Technology Committee (03J14019, 06DJ14004, 06DZ19504).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Min He, Gen-Hong Di and A-Yong Cao contributed equally for this study.
Rights and permissions
About this article
Cite this article
He, M., Di, GH., Cao, AY. et al. RAD50 and NBS1 are not likely to be susceptibility genes in Chinese non-BRCA1/2 hereditary breast cancer. Breast Cancer Res Treat 133, 111–116 (2012). https://doi.org/10.1007/s10549-011-1700-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-011-1700-2