Abstract
Background
New molecular biology-based methods of bacterial identification are expected to help elucidate the relationship between colorectal cancer (CRC) and intestinal microbiota. Although there is increasing evidence revealing the potential role of microbiota in CRC, it remains unclear whether microbial dysbiosis is the cause or the result of CRC onset.
Aim
We investigated the changes of intestinal environments in CRC or adenoma.
Methods
We analyzed 13 groups of microbiota, 8 types of organic acids, and pH in feces obtained from the following 3 groups: individuals with CRC, adenoma, and non-adenoma. Ninety-three patients with CRC and 49 healthy individuals (22 with adenoma and 27 without adenoma) were enrolled.
Results
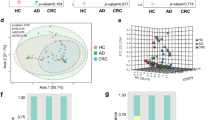
The counts of total bacteria (10.3 ± 0.7 vs. 10.8 ± 0.3 log10 cells/g of feces; p < 0.001), 5 groups of obligate anaerobe, and 2 groups of facultative anaerobes were significantly lower in the CRC group than in the healthy individuals. While the concentrations of short chain fatty acids (SCFAs) were significantly decreased in the CRC group, the pH was increased in the CRC group (7.4 ± 0.8 vs. 6.9 ± 0.6; p < 0.001). Comparison among the CRC, adenoma, and non-adenoma groups revealed that fecal SCFAs and pH in the adenoma group were intermediate to the CRC group and the non-adenoma group. Within the CRC group, no differences in microbiota or organic acids were observed among Dukes stages.
Conclusions
CRC patients showed significant differences in the intestinal environment, including alterations of microbiota, decreased SCFAs, and elevated pH. These changes are not a result of CRC progression but are involved in CRC onset.


Similar content being viewed by others
References
Gill SR, Pop M, Deboy RT, et al. Metagenomic analysis of the human distal gut microbiome. Science. 2006;312:1355–1359.
Hooper LV, Macpherson AJ. Immune adaptations that maintain homeostasis with the intestinal microbiota. Nat Rev Immunol. 2010;10:159–169.
Hooper LV, Gordon JI. Commensal host-bacterial relationships in the gut. Science. 2001;292:1115–1118.
Neish AS. Microbes in gastrointestinal health and disease. Gastroenterology. 2009;136:65–80.
Bjorksten B, Sepp E, Julge K, et al. Allergy development and the intestinal microflora during the first year of life. J Allergy Clin Immunol.. 2001;108:516–520.
Turnbaugh PJ, Ley RE, Mahowald MA, et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006;444:1027–1031.
Rawls JF. Enteric infection and inflammation alter gut microbial ecology. Cell Host Microbe. 2007;22:73–74.
Matsuki T, Watanabe K, Fujimoto J, et al. Use of 16S rRNA gene-targeted group-specific primers for real-time PCR analysis of predominant bacteria in human feces. Appl Environ Microbiol. 2004;70:7220–7228.
Eckburg PB, Bik EM, Bernstein CN, et al. Diversity of the human intestinal microbial flora. Science. 2005;308:1635–1638.
Zoetendal EG, Rajilic-Stojanovic M, de Vos WM. High-throughput diversity and functionality analysis of the gastrointestinal tract microbiota. Gut. 2008;57:1605–1615.
Ohara T, Yoshino K, Kitajima M. Possibility of preventing colorectal carcinogenesis with probiotics. Hepatogastroenterology. 2010;57:1411–1415.
Sobhani I, Tap J, Roudot-Thoraval F, et al. Microbial dysbiosis in colorectal cancer (CRC) patients. PLoS ONE. 2011;6:e16393.
Scanlan PD, Shanahan F, Clune Y, et al. Culture-independent analysis of the gut microbiota in colorectal cancer and polyposis. Environ Microbiol. 2008;10:789–798.
Matsuda K, Tsuji H, Asahara T, et al. Sensitive quantitative detection of commensal bacteria by rRNA-targeted reverse transcription-PCR. Appl Environental Microbiol.. 2007;73:32–39.
Matsuda K, Tsuji H, Asahara T, et al. Establishment of an analytical system for the human fecal microbiota, based on reverse transcription-quantitative PCR targeting of multicopy rRNA molecules. Appl Environental Microbiol.. 2009;75:1961–1969.
Bird AR, Brown IL, Topping DL. Starches, resistant starches, the gut microflora and human health. Curr Issues Intest Microbiol.. 2000;1:25–37.
Wong JM, de Souza R, Kendall CW, et al. Colonic health: fermentation and short chain fatty acids. J Clin Gastroenterol. 2006;40:235–243.
Tjalsma H, Scholler-Guinard M, Lasonder E, et al. Profiling the humoral immune response in colon cancer patients: diagnostic antigens from Streptococcus bovis. Int J Cancer. 2006;119:2127–2135.
Ellmerich S, Djouder N, Scholler M, Klein JP. Production of cytokines by monocytes, epithelial and endothelial cells activated by Streptococcus bovis. Cytokine. 2000;12:26–31.
Hill MJ, Drasar BS, Hawksworth G, et al. Bacteria and aetiology of cancer of large bowel. Lancet. 1971;1:95–100.
Moore WE, Moore LH. Intestinal floras of populations that have a high risk of colon cancer. Appl Environ Microbiol. 1995;61:3202–3207.
Scheppach W. Effects of short chain fatty acids on gut morphology and function. Gut. 1994;35:S35–S38.
Augenlicht LH, Mariadason JM, Wilson A, et al. Short chain fatty acids and colon cancer. J Nutr. 2002;132:3804S–3808S.
Fearon ER, Vogelstein B. A genetic model for colorectal tumorigenesis. Cell. 1990;61:759–767.
Johnson IT, Lund EK. Review article: nutrition, obesity and colorectal cancer. Aliment Pharmacol Ther. 2007;26:161–181.
Matsubara N. Epigenetic regulation and colorectal cancer. Dis Colon Rectum. 2012;55:96–104.
Liu Z, Qin H, Yang Z, et al. Randomised clinical trial: the effects of perioperative probiotic treatment on barrier function and post-operative infectious complications in colorectal cancer surgery—a double-blind study. Aliment Pharmacol Ther. 2011;33:50–63.
Gianotti L, Morelli L, Galbiati F, et al. A randomized double-blind trial on perioperative administration of probiotics in colorectal cancer patients. World J Gastroenterol. 2010;16:167–175.
Worthley DL, Le Leu RK, Whitehall VL, et al. A human, double-blind, placebo-controlled, crossover trial of prebiotic, probiotic, and synbiotic supplementation: effects on luminal, inflammatory, epigenetic, and epithelial biomarkers of colorectal cancer. Am J Clin Nutr. 2009;90:578–586.
Quintero E. Chemical or immunological tests for the detection of fecal occult blood in colorectal cancer screening? Gastroenterol Hepatol. 2009;32:565–576.
Mueller S, Saunier K, Hanisch C, et al. Differences in fecal microbiota in different European study populations in relation to age, gender, and country: a cross-sectional study. Appl Environ Microbiol. 2006;72:1027–1033.
Zhao L, Xu W, Ibrahim SA, et al. Effects of age and region on fecal microflora in elderly subjects living in Bama, Guangxi. China. Curr Microbiol.. 2011;62:64–70.
Cummings JH, Pomare EW, Branch WJ, et al. Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut. 1987;28:1221–1227.
Kikuchi H, Yamada T. Correlation between water-holding capacity of different types of cellulose in vitro and gastrointestinal retention time in vivo of rats. J Sci Food Agric. 1992;60:139–146.
Matsuki T. Development of quantitative PCR detection method with 16S rRNA gene-targeted genus- and species-specific primers for the analysis of human intestinal microflora and its application. Nippon Saikingaku Zasshi. 2007;62:255–261.
Matsuki T, Watanabe K, Tanaka R, et al. Distribution of bifidobacterial species in human intestinal microflora examined with 16S rRNA-gene-targeted species-specific primers. Appl Environ Microbiol. 1999;65:4506–4512.
Matsuda K, Tsuji H, Asahara T, et al. Sensitive quantification of Clostridium difficile by reverse transcription—quantitative PCR (RT-qPCR) targeting rRNA molecules. Appl Environ Microbiol. 2012;78:5111–5118.
Kikuchi E, Miyamoto Y, Narushima S, et al. Design of species-specific primers to identify 13 species of Clostridium harbored in human intestinal tracts. Microbiol Immunol. 2002;46:353–358.
Acknowledgments
We thank all the subjects who participated in this study. We also thank the nursing staff of the 10E Medical Examination Ward and 8E Surgery Ward in St. Luke’s International Hospital for their cooperation in the prompt collection and storage of fecal samples. We thank Norikatsu Yuki for technical help with the analysis of fecal samples. This study was supported in part by a grant from St. Luke’s Life Science Institute.
Conflicts of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ohigashi, S., Sudo, K., Kobayashi, D. et al. Changes of the Intestinal Microbiota, Short Chain Fatty Acids, and Fecal pH in Patients with Colorectal Cancer. Dig Dis Sci 58, 1717–1726 (2013). https://doi.org/10.1007/s10620-012-2526-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-012-2526-4