Abstract
Enterovirus 71 (EV71) is a neurotropic virus that causes various clinical manifestations in young children, ranging from asymptomatic to fatal. Different pathotypes of EV71 notably differ in virulence. Several virulence determinants of EV71 have been predicted. However, these reported virulence determinants could not be used to identify the EV71 strains of subgenotype C4, which mainly circulate in China. In this study, VP1 sequences of 37 EV71 strains from severe cases (SC-EV71) and 192 EV71 strains from mild cases (MC-EV71) in mainland China were analyzed to determine the potential virulence determinants in the capsid protein VP1 of EV71. Although most SC-EV71 strains belonged to subgenotype C4a, no specific genetic lineages in C4a were correlated with EV71 virulence. Interestingly, amino acid substitutions at nine positions (H22Q, P27S, N31S/D, E98K, E145G/Q, D164E, T240A/S, V249I, and A289T) were detected by aligning the VP1 sequences of the SC-EV71 and MC-EV71 strains. Moreover, both the constituent ratios of the conservative or mutated residues in the MC-EV71 and SC-EV71 strains and the changes in the VP1 3D structure resulting from these mutations confirmed that the conservative residues (22H, 249V, and 289A) and the mutated residues (27S, 31S/D, 98K, 145G/Q, 164E, and 240A/S) might be potential virulence determinants in VP1 of EV71. Furthermore, these results led to the hypothesis that VP1 acts as a sandwich switch for viral particle stabilization and cellular receptors attachment, and specific mutations in this protein can convert mild cases into severe cases. These findings highlight new opportunities for diagnostic and therapeutic interventions.
Similar content being viewed by others
Introduction
Enterovirus 71 (EV71) is a major causative agent of hand, foot, and mouth disease (HFMD) [1]. HFMD is a type of self-limited disease that presents with mild symptoms, such as fever, sores in the mouth, and a rash or blisters on the hands and feet [2]. However, HFMD caused by EV71 infection is frequently accompanied by a series of severe complications, such as aseptic meningitis, poliomyelitis-like paralysis, brain stem encephalitis, pulmonary edema, and even death [3]. Before 1998, three large outbreaks of HFMD with fatal neurological complications occurred in Bulgaria, Hungary, and Malaysia [4–6]. Additionally, the largest outbreak of HFMD occurred in Taiwan in 1998, resulting in 129,000 mild cases, 405 severe cases with neurological complications and 78 deaths [7]. Since the high morbidity and mortality of severe neurological diseases are caused by EV71 infection, a growing number of researchers have investigated the mechanisms of different pathotypes of EV71. Despite the wide variation in the clinical manifestations of EV71 infection, there was no specific clinical feature for clinicians to predict which patients would develop severe disease leading to death [6]. This phenomenon of patients with EV71 infection having such a wide variety of clinical outcomes may be related to the virulence of the particular viral strain and specific host factors [8, 9]. Some studies have shown that the host immune responses, especially cellular immune reactions, might be an important factor that contributes to the various clinical outcomes [9, 10]. However, the viral factors associated with clinical outcome remain unclear.
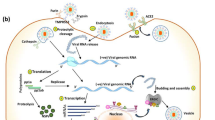
EV71 is a member of the human enterovirus species A in the genus Enterovirus of the family Picornaviridae [11]. It contains a single stranded, positive-sense RNA molecule nearly 7,400 nucleotides in length, which is surrounded by an icosahedral capsid without an envelope [12]. The viral genome is composed of 5′- and 3′-untranslated regions (UTR), which is indispensable for viral replication, and one long open reading frame (ORF) [13]. The ORF can be translated into four structural proteins (VP1–VP4) and seven nonstructural proteins (2A–2C, 3A–3D) [13, 14]. The nonstructural proteins play important roles in processing the viral genome, promoting viral replication, and shutting down the translation of host cell proteins [12]. Except that the capsid protein VP4 is inside the viral particle, VP1, VP2, and VP3 are exposed on the surface of EV71 which recognize the cellular receptors, and are antigenic [12]. In addition, VP1 is the most external and is the main part to constitute the canyon on the surface of picornaviruses, and is potentially involved in the viral pathogenicity [15]. Several researches have indicated that VP1 is significant in immune modulation of EV71 and vaccine development [16–21]. Thus, VP1 plays an important role in the life history of EV71.
To date, some groups have argued that the differences in EV71 strains might contribute to the different severity of the disease [22, 23]. It has been demonstrated that sequences in the 5′-UTR and 3D gene are related with the neurovirulence of both polioviruses and EV71 [24–26]. Other studies also explored the virulence determinants of EV71 strains [27, 28]. Li et al. [28] have identified that amino acid residues G/Q/R at VP1-145, E at VP1-164, K at VP2-68, and certain nucleotides (272G, 488U, and 700A/U) in the 5′-UTR are associated with the virulent phenotype of EV71. Although most EV71 strains which cause severe diseases can be identified by these virulence determinants with high sensitivity, subgenotype C4 EV71 strains cannot be well recognized [28]. Because EV71 strains circulate in mainland China are grouped into subgenotype C4, these previous results cannot be represented the virulence determinants of Chinese EV71 strains. In this study, the VP1 sequences of 37 EV71 strains from severe cases (SC-EV71) and 192 EV71 strains from mild cases (MC-EV71) in mainland China were analyzed. Further, the potential determinants for EV71 virulence in the capsid protein VP1 were explored.
Materials and methods
Ethics statement
This study has obtained ethics approval from the ethics committee at the School of Medicine, Wuhan University, in accordance with the guidelines for the protection of human subjects. Written informed consent was obtained from the parents of all the children involved in our study.
Viruses and VP1 gene sequences
Nine strains were isolated from throat swab specimens collected from HFMD patients in Hubei province and their complete VP1 genes were sequenced in our laboratory. Their VP1 nucleotide sequences had been deposited in the GenBank database under the accession numbers JQ419491–JQ419499 (Table S1). Only one EV71 strain (JQ419499) was isolated from a patient with tremor which was considered as one of neurological complications. Other eight EV71 strains all caused mild HFMD in patients. Complete VP1 gene sequences of 220 EV71 strains (including 36 strains isolated in patients with severe disease) isolated in 1987, 1997–1998, 2000, 2002–2003, and 2007–2011 from various regions of Chinese mainland were downloaded from GenBank (Table S1). Thus, totally 37 SC-EV71 and 192 MC-EV71 strains were analyzed in this study. The SC-EV71 infection causes severe clinical presentations such as meningitis, encephalitis, acute flaccid paralysis (AFP), and tremor. And the MC-EV71 infection causes mild HFMD, the manifestations of which are fever, skin rash, and herpes.
Phylogenetic analysis of the VP1 genes
The phylogenetic tree was constructed by the neighbor-joining method (1,000 bootstrap replicates) with Kimura 2-parameter distance model using the MEGA 5.0 software. The bootstrap value of over 70 % supporting each cluster was shown at the nodes.
Alignment of the amino acid sequences deduced from the VP1 genes
The complete VP1 amino acid sequences of the 37 SC-EV71 and 192 MC-EV71 strains were aligned by ClustalW multiple alignment method using MEGA 5.0 software.
Structural analysis of the VP1 capsid protein
Crystal structures of human enterovirus Protein Data Bank Accession Nos. 4AED and 3VBS were used for presumed SC-EV71 and MC-EV71 strains, respectively [29, 30]. Structural visualization of VP1 protein was done using Pymol software.
Statistical analysis
Statistical analysis was performed with Chi square and Fisher’s exact test by SPSS 15.0 software. Level of significance (α) was set at 0.05.
Results
Phylogenetic analysis based on the VP1 genes of EV71
To explore whether genetic lineages are related to the virulence of Chinese EV71, a phylogenetic tree based on the VP1 genes of 229 EV71 strains was constructed using the neighbor-joining method. The 192 MC-EV71 strains were grouped into two distinct genotypes, A and C (C0, C4a and C4b) (Fig. 1). With the exception of one SC-EV71 strain belonging to subgenotype C3, the SC-EV71 strains were subdivided into clade C4a of the cladogram (Fig. 1). These results suggested that the EV71 strains that belong to cluster C4a, which circulate in mainland China, were capable of causing severe disease. Regrettably, the SC-EV71 strains were scattered in various branches of C4a, and it could be that no special genetic lineages in C4a correlated with EV71 virulence.
Phylogenetic tree based on the complete VP1 gene sequences of the 229 EV71 strains. The phylogenetic tree was generated using the neighbor-joining method (bootstrap analysis with 1,000 replicates) with MEGA version 5.0 software. The percentage bootstrap of supporting each branch no lower than 70 % is shown at the nodes. The scale bar indicates the number of nucleotide substitutions per site. The black triangle indicated EV71 strains from severe cases
Analysis of the amino acid sequence of VP1 between the SC-EV71 and MC-EV71 strains
As one component of the viral capsid, the VP1 protein plays important roles during EV71 infection. To determine the differences in the VP1 protein that may be associated with EV71 virulence, the VP1 protein sequences from 37 SC-EV71 and 192 MC-EV71 strains were aligned (Table 1; Fig. 2). Interestingly, the VP1 sequences of 15 SC-EV71 strains were identical with those of 69 MC-EV71 strains. To simplify further analyses, the VP1 sequences of these 84 EV71 strains were used as the reference sequence, referred to as the consensus (Fig. 2a). To determine the potential virulence determinants in VP1, the consensus was compared with the VP1 sequences of 22 additional SC-EV71 strains (Fig. 2b), and amino acid substitutions at nine positions (H22Q, P27S, N31S/D, E98K, E145G/Q, D164E, T240A/S, V249I, and A289T) were detected in VP1 (Fig. 2b). The conservative residues 22H, 27P, 31N, 98E, 145E, 164D, 240T, 249V, and 289A were present in the consensus, while the mutated residues 22Q, 27S, 31S/D, 98K, 145G/Q, 164E, 240A/S, 249I, and 289T were found in the SC-EV71 strains. Because of the polymorphisms in VP1, we further analyzed whether these amino acid substitutions occurred in the remaining 123 MC-EV71 strains (Fig. 2b). Based on this comparison, the 123 MC-EV71 strains were divided into two categories (Fig. 2b). One category consisted of 23 MC-EV71 strains that did not contain any of these amino acid substitutions, and the VP1 sequences of these 23 EV71 strains were designated into sequence cluster 32 (Table 1). The other category consisted of 100 MC-EV71 strains that contained at least one of these substitutions (Fig. 2b). Given that these amino acid substitutions were present in 22 SC-EV71 and 100 MC-EV71 strains (Fig. 2b), the VP1 sequences of those 22 SC-EV71 and 100 MC-EV71 strains were used in further analyses to determine the potential virulence determinants of EV71 (Fig. 2c). The sequence alignment showed that 13 variations of the VP1 sequence were present in 22 SC-EV71 strains (sequence clusters 1–13, Table 1), whereas 27 variations of the VP1 sequence were present in 100 MC-EV71 strains (sequence clusters 1–9 and 14–31, Table 1). Whether the amino acid substitutions that were detected in the VP1 protein were related to EV71 virulence needed to be further confirmed.
Scheme for the analysis of the virulence determinants in the VP1 capsid protein of EV71. a The VP1 deduced amino acid sequences of 37 SC-EV71 and 192 MC-EV71 strains were aligned. Fifteen SC-EV71 and 69 MC-EV71 strains had the same VP1 amino acid sequences, and this sequence was designated the consensus sequence. The non-consensus sequences included the VP1 protein sequences of the remaining 22 SC-EV71 and 123 MC-EV71 strains. b Amino acid substitutions at nine positions were found by comparing the consensus sequence with the VP1 sequences of the remaining 22 SC-EV71 strains. Furthermore, according to whether these amino acid substitutions occurred in the 123 non-consensus MC-EV71 strains, the 123 MC-EV71 strains were divided into two categories. In total, 23 MC-EV71 strains did not contain these substitutions, whereas 100 MC-EV71 strains contained at least one of these substitutions. c The VP1 sequences of the 22 SC-EV71 and 100 MC-EV71 strains were used in further analyses to find out potential virulence determinants. SC-EV71: EV71 strains from severe cases. MC-EV71: EV71 strains from mild cases
Combinations among the potential virulent residues in VP1 of EV71
It has been reported that several amino acid mutations in the viral genome were related to EV71 virulence [22, 27, 28]. In the present study, the constituent ratios of these amino acid substitutions (H22Q, P27S, N31S/D, E98K, E145G/Q, D164E, T240A/S, V249I, and A289T) at a single position in the 22 SC-EV71 and 100 MC-EV71 strains were calculated to identify whether these substitutions were associated with EV71 virulence (Table 2). Because the number of single amino acid substitutions in the SC-EV71 strains was fewer, it was difficult to calculate the p value of the constituent ratios of single substitutions between the SC-EV71 and MC-EV71 strains. Because of the high mutational rate of viral RNA replication during EV71 evolution and the complex pathogenesis of EV71, the virulence of EV71 was usually associated with multiple amino acid residues rather than a single substitution. Thus, the virulent combinations of the mutated residues were explored (Fig. 3).
Amino acid combinations among the potential virulence residues in group I and group II. Group I contained three conservative residues 22H, 249V, and 289A which were classified into seven sets 22H, 249V, 289A, 22H249V, 22H289A, 249V289A, and 22H249V289A. Group II contained the mutated residues 27S, 31S/D, 98K, 145G/Q, 164E, and 240A/S which were grouped into 63 sets. Then, the seven sets of group I residues were combined with each set of group II residues to find out the potential virulence combinations in VP1. The “/” indicated that the virulent residues sets contained at lest one of the residues in group I and group II. Six sets of group II residues significantly differed between the MC-EV71 and SC-EV71 (p < 0.05) were showed in bold. SC-EV71: EV71 strains from severe cases. MC-EV71: EV71 strains from mild cases
The constituent ratios of the mutated residues 22Q, 249I, and 289T in MC-EV71 were higher than in SC-EV71 (Table 2), which suggested that the constituent ratios of the conservative residues 22H, 249V, and 289A in SC-EV71 were higher than in MC-EV71. The three conservative residues 22H, 249V, and 289A were classified in group I. Moreover, the constituent ratios of the mutated residues 27S, 31S/D, 98K, 145G/Q, 164E, and 240A/S in the SC-EV71 strains were higher than in the MC-EV71 strains (Table 2). These mutated residues were classified in group II. In group I, the constituent ratio of the residues 22H249V289A significantly differed between the MC-EV71 and SC-EV71 strains (Χ 2 = 4.243, p = 0.039, Table 1). Therefore, the conservative residues 22H249V289A might be the potential virulence determinants in the VP1 protein of EV71. In group II, the mutated residues were more likely to be present in the SC-EV71 strains, which indicated that these mutated residues in VP1 might be related to EV71 virulence.
Residues in group I and group II were both likely to be present in the SC-EV71 strains; therefore, we tried to determine the potential virulent combinations of these residues. The conservative residues in group I were designated into seven sets: 22H, 249V, 289A, 22H249V, 22H289A, 249V289A, and 22H249V289A. The mutated residues in group II were designated into 63 sets. The constituent ratios of six sets of group II were significantly different between the SC-EV71 and MC-EV71 strains (Table 3). Furthermore, to synthetically explore the multiple virulence determinants in VP1 of EV71, we wanted to know whether these virulent residues in group I and group II synergistically affected EV71 virulence. Therefore, the seven sets of group I residues were combined with each set of the group II residues. In total, 441 combinations were found between the sets of group I and group II. Furthermore, 108 combinations significantly differed between the SC-EV71 and MC-EV71 strains (Tables 4, 5, 6, 7, 8, 9, 10), indicating that they may be related to the clinical manifestations caused by EV71 infection. These results also confirmed that amino acid substitutions at these nine positions were correlated with EV71 virulence. For example, the combination 22H289A + 98K/240A/S meant that the conservative residues 22H and 289A were both present in VP1, while at least one of the mutated residues 98K or 240A/S were present in VP1. This combination was more likely to appear in the SC-EV71 strains than in the MC-EV71 strains (Χ 2 = 10.060, p = 0.002) (Table 8). If this combination was used as the criterion to identify the SC-EV71 strains, the sensitivity was 31.82 % (Table 8). Collectively, these observed combinations provided powerful insight into strategies to control and prevent the deterioration of mild HFMD during the early stages of EV71 infection, and these combinations would act together as the diagnostic criteria for severe diseases caused by EV71 strains in the future.
The virulence determinants in VP1 based on 3D structure analysis
Like other picornaviruses, the capsid protein VP1 of EV71 is the main constituent of the canyon-like depression and the hydrophobic pocket [28]. The BC, DE, EF, and HI loops of VP1 are located at the surface of the canyon and play important roles in the binding of EV71 to the cellular receptors. Since the identified amino acid residues in VP1 might be the potential determinants of EV71 virulence, we tried to map the location of these residues in the VP1 structure and predict the influences made these amino acid substitutions on viral capsid conformation.
Based on 3D structure of VP1, residues 22, 27, 31, and 289 were located in the termini of VP1, and the residues 98, 145, 164, and 240 were located in the BC, DE, EF, and HI loops, respectively, on the surface of VP1 (Fig. 4a). Notably, the crystal structures of two different EV71 strains are available [29, 30]. Plevka et al. [29] exhibited the structure of one EV71 strain with residues 22Q, 98E, 145Q, 164E, 240S, and 289T in VP1 (Protein Data Bank Accession No. 4AED). In contrast, Wang et al. [30] reported the structure of EV71 strain with residues 22H, 98K, 145E, 164D, 240T, and 289A in VP1 (Protein Data Bank Accession No. 3VBS). Thus, we predicted how these residues influence EV71 virulence by comparing the two structures of VP1. When the A at VP1-289 was substituted with a T, a hydrogen bond formed between 289T in VP1 and 94G in VP3, causing the rearrangement of the viral capsid proteins (Fig. 4b). This rearrangement would result in the decreased stability of the viral particles. When the E at VP1-98 was substituted with a K, the positive charge near the fivefold axes of EV71 increased and promoted the interaction of EV71 and the negatively charged region of the cellular membrane (Fig. 4c). The mutated residues 145G/Q, 164E, and 240A/S located in the DE, EF, and HI loops, respectively, of VP1 would enhance the binding of EV71 to the target cells (Fig. 4d–f). These results further supported the finding that the amino acid substitutions detected in VP1 were potential virulence determinants of EV71.
Analyzing the changes in the 3D structure of VP1 caused by the potential virulent residues. a Molecular surface representation of the VP1 capsid protein is shown in gray. The potential virulence determinants are shown in magenta. The N terminus, C terminus, and core region of the VP1 protein contained 3, 1, and 4 residues, respectively. VP1-164 is located on the other side of the molecular surface. b Comparison of the interactions between the G residue at VP3-94 and the A or T residues at VP1-289. Conformations of the amino acid substitutions at four positions c E98K, d E145Q, e D164E, and f T240S all located at the bottom of the core hydrophobic region of VP1 in both the native state and the mutated state. The orientation was similar to that of the VP1 cartoon in a. The mutated residues 98K, 145Q, 164E, and 240S in the VP1 protein that increase virulence are shown in gray, while the corresponding conservative native residues 98E, 145E, 164E, and 240T are shown in blue
Discussion
Since 2007, EV71 infection has caused severe diseases with high incidence rates and mortality in the Western Pacific Region, especially in mainland China [31–33]. In this study, we demonstrated that amino acid residues at nine positions (22H, 27S, 31S/D, 98K, 145G/Q, 164E, 240A/S, 249V, and 289A) might be potential virulence determinants in the VP1 proteins of EV71 strains circulating in mainland China. Furthermore, we speculated that VP1 acts as a sandwich switch to affect viral particle stabilization and the binding of EV71 to cellular receptors, and these virulent residues in VP1 may help determine which infection will result in severe cases and mild cases in mainland China.
It had been demonstrated that the VP1 capsid protein is related to EV71 virulence in the host [17, 34–36]. The VP1 protein was selected as a potential source of virulence determinants of EV71 for the following reasons: (i) The amino acid residues 66–132 of VP1 contain the major dimerization domain and are needed for the formation of the entire capsid to enhance the pathogenicity and stability of EV71 [37]. (ii) The VP1 coding region contains an adaptor sensor that is necessary for cellular receptor attachment and the initiation of viral uncoating [30]. For example, amino acid residues VP1-172 and VP1-145 play important roles in EV71 binding to the receptors SCARB2 and PSGL-1, respectively [35, 38]. Additionally, the amino acids 40–100 of VP1 play important roles in binding and interacting with human Annexin 2 protein to enhance viral infectivity [39]. (iii) Some amino acid residues in the VP1 protein apparently interact with viral RNA and assist with its entry into the cytoplasm. (iv) The VP1 protein contains important neutralizing epitope regions. Studies have shown that the N-terminal half of the VP1 protein is a strong target for high-titered human neutralizing antibodies [17]. (v) VP1 plays an important role in protecting some animals against EV71 infection. VP1-containing milk produced by transgenic mice protects the suckling mice against EV71 infection by acting as a potential oral vaccine [40]. Additionally, a novel antiviral peptide derived from the VP1 capsid protein of EV71 can inhibit viral infection [41]. Lactoferrin can bind to the VP1 protein and inhibits EV71 infection [18]. (vi) It has been reported that a few mutations in the VP1 gene can lead to drug resistance and virulence change [20, 42].
Previous studies have indicated that the mutation of some amino acid residues in the VP1 protein can affect EV71 virulence [28, 43–45], and amino acid residues at positions 22, 31, 145, 164, 240, 249, and 289 in VP1 have been predicted to be related to EV71 virulence [28, 46]. However, few studies have systematically explored the virulence determinants of the EV71 strains circulating in mainland China based on the clinical symptoms of EV71 infection. In our study, amino acid substitutions at nine positions (H22Q, P27S, N31S/D, E98K, E145G/Q, D164E, T240A/S, V249I, and A289T) in VP1 were associated with the virulence phenotypes of the EV71 strains in mainland China. Residues at seven positions (22, 31, 145, 164, 240, 249, and 289) in VP1 were consistent with the previous presumptions. However, in contrast with the previous research, our study extensively and systematically explored the potential virulence determinants in the VP1 protein from Chinese EV71 strains. Moreover, we demonstrated that residues at the seven positions in VP1 were potential virulence determinants of Chinese EV71 strains by comparing the constituent ratios of these residues in the SC-EV71 and MC-EV71 strains and the resulting changes in the 3D structure of VP1 caused by these residues. Notably, novel amino acid substitutions at two positions, VP1-27 and VP1-98, were found to be related to the virulence of EV71 by comparing the constituent ratios of the residue substitutions in the SC-EV71 and MC-EV71 strains and the changes in the 3D structure of the protein caused by these substitutions. It has been reported that the viral RNA may interact with residue 30Q in VP1 during uncoating [30]. So, residue 27S which located nearby position 30Q in VP1 might play an important role in viral RNA entering into target cells during EV71 uncoating by conformational change. And the mutation from E to K at VP1-98 would increase the positive charge of the fivefold axes of EV71, thus promoting the adsorption of EV71 into the target cells. These new findings would be significant and helpful for designing the viral vaccine and antiviral drugs in the future.
In this study, a hypothesis was proposed to elucidate the virulence phenotypes of EV71. Based on these results, VP1 acts as a sandwich switch to affect the virulence of the EV71 strains in mainland China (Fig. 5). The mutated residues 98K, 145G/Q, 164E, and 240A/S were located in the external loops BC, DE, EF, and HI of VP1, respectively. These mutated residues, which were located in the core region of VP1, might increase the binding of EV71 to the receptors such as PSGL-1, SCARB2, and Annexin 2, among others. The conserved residues 22H and 289A were located in the N and C termini of VP1, respectively. The conservative residues in the VP1 termini might participate in the stabilization of the viral particle during EV71 adsorption into the target cells. The hypothesis indicated that the residues in the termini of VP1 maintained conservative and the mutations in the residues in the VP1 core region to hydrophobic and positive charged residues would increase EV71 virulence. Clinically, the discovery of these potential virulence determinants in VP1 of EV71 may be very important in the prevention and treatment of the diseases caused by EV71 infection. Moreover, these findings may help determine which patients infected with EV71 will develop severe diseases from mild symptoms, and specific therapy can be administered in the early stages of EV71 infection to prevent patients for suffering life-threatening complications. This study provided new insight into the pathogenicity of EV71.
A sandwich-switch hypothesis developed from the 3D structure of the VP1 protein. The conservative residues 22H and 289A were present in the N- and C-termini of VP1, respectively. The amino acid substitutions E98K, E145G/Q, D164E, and T240A/S were present in the external loops BC (green), DE (rose), EF (red), and HI (light blue) of the VP1 protein, respectively. When no significant amino acid substitutions occurred in the two termini of VP1, the mutations of the amino acid residues 98E, 145E, 164D, and 240T (all in mandarin blue) into residues 98K, 145G/Q, 164E, and 240A/S (all in yellow), respectively, would increase the binding of EV71 to the target cellular receptors, such as PSGL-1, SCARB2, and Annexin 2, among others (receptor X), which would result in the progression of mild HFMD into severe disease. PSGL-1 P-selectin glycoprotein ligand-1, SCARB2 scavenger receptor B2, sER smooth endoplasmic reticulum, rER rough endoplasmic reticulum
In fact, the mechanism for the different severity of EV71 pathogenic is complex. Perhaps virus and host factors co-contribute to the diversity of the clinical presentations for EV71 infection [8, 31]. To date, three studies have indicated that host immune response, especially cellular immunity, is a major factor in the pathogenesis of EV71-associated neurological disease [1, 9, 10]. Chang et al. [9] presumed that virus would escape the killing or clearance by weak cellular immunity, thus resulting in viral diffusion, persistent systemic inflammatory response, and subsequent severe diseases. Moreover, the glucose-6-phosphate dehydrogenase (G6PD) deficiency has been identified as a probably predisposing factor for EV71 infection and may make the aggravation of clinical outcome, the mechanism of which may be the oxidative stress [47]. Host factors are complex and play an important role in clinical outcome induced by EV71 infection. Otherwise, it cannot be ignored that the viral factors such as viral genomic differences, virulence, and genotype may be also involved in the severity of clinical outcome [28, 33, 46]. In this study, most SC-EV71 strains were belonging to subgenotype C4a. Previously, reports have also declared that no severe or fatal cases caused by C4b EV71 strains, while the large outbreaks of HFMD with severe neurological complications are mainly induced by C4a EV71 strains [8, 48]. The reason for this phenomenon may be conjectured that the viral rapid evolution promotes the speedy spread of EV71 in the human population and causes some novel biologic characteristic of EV71, such as immune escape or high virulence [12]. However, our result showed that no specific lineages in subgenotype C4a were associated with severe or mild cases. This indicated that EV71 strains isolated from patients with different clinical symptoms derived from the common ancestor and evolved to different lineages by gradualness mutation [8]. And it also suggested that in subgenotype C4a, slight genetic mutations in VP1 gene would not change the viral lineages but might affect the viral virulence. Collectively, both virus and host factors contribute to the phenotype of EV71-associated disease.
Because there is no suitable animal model for EV71 infection in humans, it is hard to do research on the pathogenicity of EV71 [39]. EV71 infection is lethal to suckling mice, while mice more than a few weeks old are no susceptible to EV71 infection [39, 49]. Moreover, the virulent phenotype of EV71 infection in mice is different from that in humans [50]. Two studies have predicted that EV71 strains with VP1-145E are associated with milder disease in humans [28, 51]. However, Huang et al. [42] demonstrated that residues 149M of VP2 and 145E of VP1 cooperatively increased the infectivity and lethality of EV71 infection in mice. Since there are no suitable animal models for identifying the determinants of EV71 virulence, this study on analyzing the potential virulence determinants of Chinese EV71 strains was performed only at the molecular level. We found that the VP1 amino acid sequences of 15 SC-EV71 strains were identical to 69 MC-EV71 strains, which indicated that the virulence determinants of some SC-EV71 strains might be located in other regions of the EV71 genome. Li et al. [28] identified virulence determinants at one amino acid position in protease 2A and at three nucleotide positions of the 5′-UTR of EV71. Accordingly, further study should include the characterization the whole genome of the EV71 strains, and the development of suitable animal models to identify the potential virulence determinants will be needed.
References
P. McMinn, I. Stratov, L. Nagarajan, S. Davis, Clin. Infect. Dis. 32(2), 236–242 (2001)
L.M. Sun, H.Y. Zheng, H.Z. Zheng, X. Guo, J.F. He, D.W. Guan, M. Kang, Z. Liu, C.W. Ke, J.S. Li, L. Liu, R.N. Guo, H. Yoshida, J.Y. Lin, Jpn. J. Infect. Dis. 64(1), 13–18 (2011)
P. Chen, Z. Song, Y. Qi, X. Feng, N. Xu, Y. Sun, X. Wu, X. Yao, Q. Mao, X. Li, W. Dong, X. Wan, N. Huang, X. Shen, Z. Liang, W. Li, J. Biol. Chem. 287(9), 6406–6420 (2013)
L.M. Shindarov, M.P. Chumakov, M.K. Voroshilova, S. Bojinov, S.M. Vasilenko, I. Iordanov, I.D. Kirov, E. Kamenov, E.V. Leshchinskaya, G. Mitov, I.A. Robinson, S. Sivchev, S. Staikov, J. Hyg. Epidemiol. Microbiol. Immunol. 23(3), 284–295 (1979)
G. Nagy, S. Takatsy, E. Kukan, I. Mihaly, I. Domok, Arch. Virol. 71(3), 217–227 (1982)
L.G. Chan, U.D. Parashar, M.S. Lye, F.G. Ong, S.R. Zaki, J.P. Alexander, K.K. Ho, L.L. Han, M.A. Pallansch, A.B. Suleiman, M. Jegathesan, L.J. Anderson, Clin. Infect. Dis. 31(3), 678–683 (2000)
M. Ho, E.R. Chen, K.H. Hsu, S.J. Twu, K.T. Chen, S.F. Tsai, J.R. Wang, S.R. Shih, N. Engl. J. Med. 341(13), 929–935 (1999)
Y. Zhang, X. Tan, A. Cui, N. Mao, S. Xu, Z. Zhu, J. Zhou, J. Shi, Y. Zhao, X. Wang, X. Huang, S. Zhu, Y. Zhang, W. Tang, H. Ling, W. Xu, PLoS One 8(2), e56341 (2013)
L.Y. Chang, C.A. Hsiung, C.Y. Lu, T.Y. Lin, F.Y. Huang, Y.H. Lai, Y.P. Chiang, B.L. Chiang, C.Y. Lee, L.M. Huang, Pediatr. Res. 60(4), 466–471 (2006)
K.D. Yang, M.Y. Yang, C.C. Li, S.F. Lin, M.C. Chong, C.L. Wang, R.F. Chen, T.Y. Lin, J. Infect. Dis. 183(6), 850–856 (2001)
C.C. Yip, S.K. Lau, B. Zhou, M.X. Zhang, H.W. Tsoi, K.H. Chan, X.C. Chen, P.C. Woo, K.Y. Yuen, Arch. Virol. 155(9), 1413–1424 (2010)
X. Chen, Q. Zhang, J. Li, W. Cao, J.X. Zhang, L. Zhang, W. Zhang, Z.J. Shao, Y. Yan, Virology 398(2), 251–261 (2010)
S.W. Huang, D. Kiang, D.J. Smith, J.R. Wang, Exp. Biol. Med. (Maywood) 236(8), 899–908 (2011)
Y.F. Chan, I.C. Sam, S. AbuBakar, Infect. Genet. Evol. 10(3), 404–412 (2010)
N.H. De Jesus, Virol. J. 4, 70 (2007)
H.F. Chen, M.H. Chang, B.L. Chiang, S.T. Jeng, Vaccine 24(15), 2944–2951 (2006)
C.S. Tan, M.J. Cardosa, Arch. Virol. 152(6), 1069–1073 (2007)
T.Y. Weng, L.C. Chen, H.W. Shyu, S.H. Chen, J.R. Wang, C.K. Yu, H.Y. Lei, T.M. Yeh, Antiviral Res. 67(1), 31–37 (2005)
C.N. Wu, Y.C. Lin, C. Fann, N.S. Liao, S.R. Shih, M.S. Ho, Vaccine 20(5–6), 895–904 (2001)
S.R. Shih, M.C. Tsai, S.N. Tseng, K.F. Won, K.S. Shia, W.T. Li, J.H. Chern, G.W. Chen, C.C. Lee, Y.C. Lee, K.C. Peng, Y.S. Chao, Antimicrob. Agents Chemother. 48(9), 3523–3529 (2004)
G.H. Chang, Y.J. Luo, X.Y. Wu, B.Y. Si, L. Lin, Q.Y. Zhu, Virol. J. 7, 106 (2010)
A. Hagiwara, T. Yoneyama, S. Takami, I. Hashimoto, Arch. Virol. 79(3–4), 273–283 (1984)
J. Melnick, Rev. Infect. Dis. 6, 387–390 (1984)
C. Christodoulou, F. Colbere-Garapin, A. Macadam, L.F. Taffs, S. Marsden, P. Minor, F. Horaud, J. Virol. 64(10), 4922–4929 (1990)
A. McGoldrick, A.J. Macadam, G. Dunn, A. Rowe, J. Burlison, P.D. Minor, J. Meredith, D.J. Evans, J.W. Almond, J. Virol. 69(12), 7601–7605 (1995)
M. Arita, H. Shimizu, N. Nagata, Y. Ami, Y. Suzaki, T. Sata, T. Iwasaki, T. Miyamura, J. Gen. Virol. 86(Pt 5), 1391–1401 (2005)
G.H. Chang, L. Lin, Y.J. Luo, L.J. Cai, X.Y. Wu, H.M. Xu, Q.Y. Zhu, Virus Res. 151(1), 66–73 (2010)
R. Li, Q. Zou, L. Chen, H. Zhang, Y. Wang, PLoS One 6(10), e26237 (2011)
P. Plevka, R. Perera, J. Cardosa, R.J. Kuhn, M.G. Rossmann, Science 336(6086), 1274 (2012)
X. Wang, W. Peng, J. Ren, Z. Hu, J. Xu, Z. Lou, X. Li, W. Yin, X. Shen, C. Porta, T.S. Walter, G. Evans, D. Axford, R. Owen, D.J. Rowlands, J. Wang, D.I. Stuart, E.E. Fry, Z. Rao, Nat. Struct. Mol. Biol. 19(4), 424–429 (2012)
T.Y. Lin, S.J. Twu, M.S. Ho, L.Y. Chang, C.Y. Lee, Emerg. Infect. Dis. 9(3), 291–293 (2003)
C.C. Liu, H.W. Tseng, S.M. Wang, J.R. Wang, I.J. Su, J. Clin. Virol. 17(1), 23–30 (2000)
H. Shimizu, A. Utama, N. Onnimala, C. Li, Z. Li-Bi, M. Yu-Jie, Y. Pongsuwanna, T. Miyamura, Pediatr. Int. 46(2), 231–235 (2004)
C. Li, H. Wang, S.R. Shih, T.C. Chen, M.L. Li, Curr. Med. Chem. 14(8), 847–856 (2007)
P. Chen, Z. Song, Y. Qi, X. Feng, N. Xu, Y. Sun, X. Wu, X. Yao, Q. Mao, X. Li, W. Dong, X. Wan, N. Huang, X. Shen, Z. Liang, W. Li, J. Biol. Chem. 287(9), 6406–6420 (2012)
C.C. Liu, A.H. Chou, S.P. Lien, H.Y. Lin, S.J. Liu, J.Y. Chang, M.S. Guo, Y.H. Chow, W.S. Yang, K.H. Chang, C. Sia, P. Chong, Vaccine 29(26), 4362–4372 (2011)
S.K. Lal, P. Kumar, W.M. Yeo, A. Kar-Roy, V.T. Chow, J. Med. Virol. 78(5), 582–590 (2006)
Y. Nishimura, H. Lee, S. Hafenstein, C. Kataoka, T. Wakita, J.M. Bergelson, H. Shimizu, PLoS Pathog. 9(7), e1003511 (2013)
S.L. Yang, Y.T. Chou, C.N. Wu, M.S. Ho, J. Virol. 85(22), 11809–11820 (2011)
H.L. Chen, J.Y. Huang, T.W. Chu, T.C. Tsai, C.M. Hung, C.C. Lin, F.C. Liu, L.C. Wang, Y.J. Chen, M.F. Lin, C.M. Chen, Vaccine 26(23), 2882–2889 (2008)
B. Premanand, T.K. Kiener, T. Meng, Y.R. Tan, Q. Jia, V.T. Chow, J. Kwang, Antiviral Res. 95(3), 311–315 (2012)
S.W. Huang, Y.F. Wang, C.K. Yu, I.J. Su, J.R. Wang, Virology 422(1), 132–143 (2012)
D. Guan, S. van der Sanden, H. Zeng, W. Li, H. Zheng, C. Ma, J. Su, Z. Liu, X. Guo, X. Zhang, L. Liu, M. Koopmans, C. Ke, PLoS One 7(9), e44386 (2012)
J. Li, X. Huo, Y. Dai, Z. Yang, Y. Lei, Y. Jiang, G. Li, J. Zhan, F. Zhan, Virus Res. 169(1), 195–202 (2012)
J. Zhu, Z. Luo, J. Wang, Z. Xu, H. Chen, D. Fan, N. Gao, G. Ping, Z. Zhou, Y. Zhang, J. An, PLoS One 8(2), e56318 (2013)
J.S. Wu, N. Zhao, H. Pan, C.M. Wang, B. Wu, H.M. Zhang, H.X. He, D. Liu, S. Amer, S.L. Liu, J. Virol. Methods 193(2), 713–728 (2013)
J.B. Ou, C.M. Zhang, S.M. Fu, X. Huang, L.H. Huang, Zhongguo Dang Dai Er Ke Za Zhi 15(9), 751–755 (2013)
Y. Zhang, Z. Zhu, W. Yang, J. Ren, X. Tan, Y. Wang, N. Mao, S. Xu, S. Zhu, A. Cui, Y. Zhang, D. Yan, Q. Li, X. Dong, J. Zhang, Y. Zhao, J. Wan, Z. Feng, J. Sun, S. Wang, D. Li, W. Xu, Virol. J. 7, 94 (2010)
Y.F. Wang, C.T. Chou, H.Y. Lei, C.C. Liu, S.M. Wang, J.J. Yan, I.J. Su, J.R. Wang, T.M. Yeh, S.H. Chen, C.K. Yu, J. Virol. 78(15), 7916–7924 (2004)
K. Fujii, N. Nagata, Y. Sato, K.C. Ong, K.T. Wong, S. Yamayoshi, M. Shimanuki, H. Shitara, C. Taya, S. Koike, Proc. Natl. Acad. Sci. U.S.A. 110(36), 14753–14758 (2013)
S.C. Chang, W.C. Li, G.W. Chen, K.C. Tsao, C.G. Huang, Y.C. Huang, C.H. Chiu, C.Y. Kuo, K.N. Tsai, S.R. Shih, T.Y. Lin, J. Med. Virol. 84(6), 931–939 (2012)
Acknowledgments
We are grateful to all the staffs engaged in investigating the epidemiology and pathogenicity of EV71 for their enormous contributions. We especially thank the researchers for their achievement in the viral structure. This work was supported by the National Natural Sciences Foundation of China (Nos. 81171577, 81371790, 81371422, and 81171127), the National Basic Research Program of China (2010CB529803), the National Science and Technology Major Projects of New Drugs (2012ZX09103301-028), and the Fundamental Research Funds for the Central Universities of China (Nos. 201130102020001, 201130102020002, and 2012301020207).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, Y., Fu, C., Wu, S. et al. A novel finding for enterovirus virulence from the capsid protein VP1 of EV71 circulating in mainland China. Virus Genes 48, 260–272 (2014). https://doi.org/10.1007/s11262-014-1035-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-014-1035-2