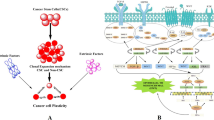
Abstract
The exact mechanisms of spontaneous tumor remission or complete response to treatment are phenomena in oncology that are not completely understood. We use a concept from ecology, the Allee effect, to help explain tumor extinction in a model of tumor growth that incorporates feedback regulation of stem cell dynamics, which occurs in many tumor types where certain signaling molecules, such as Wnts, are upregulated. Due to feedback and the Allee effect, a tumor may become extinct spontaneously or after therapy even when the entire tumor has not been eradicated by the end of therapy. We quantify the Allee effect using an ‘Allee index’ that approximates the area of the basin of attraction for tumor extinction. We show that effectiveness of combination therapy in cancer treatment may occur due to the increased probability that the system will be in the Allee region after combination treatment versus monotherapy. We identify therapies that can attenuate stem cell self-renewal, alter the Allee region and increase its size. We also show that decreased response of tumor cells to growth inhibitors can reduce the size of the Allee region and increase stem cell densities, which may help to explain why this phenomenon is a hallmark of cancer.









Similar content being viewed by others
References
Anastas JN, Moon RT (2013) Wnt signalling pathways as therapeutic targets in cancer. Nat Rev Cancer 13(1):11–26. doi:10.1038/nrc3419
Arnold D, Polking J (1999) Ordinary differential equations using MATLAB, 2nd edn. Prentice Hall, Upper Saddle River
Bachman J, Hillen T (2012) Mathematical optimization of the combination of radiation and differentiation therapies of cancer. Front Mol Cell Oncol. doi:10.3389/fonc.2013.00052
Bayin N, AS M, Placantonakis D (2014) Glioblastoma stem cells: molecular characteristics and therapeutic implications. World J Stem Cells 6(2):230–238
Bienz M, Clevers H (2000) Linking colorectal cancer to wnt signaling. Cell 103(2):311–320
Buchmann A, Alber M, Zartman J (2014) Sizing it up: the mechanical feedback hypothesis of organ growth regulation. Semin Cell Dev Biol 35:73–81
Caja L, Bellomo C, Moustakas A (2015) Transforming growth factor \(\beta \) and bone morphogenetic protein actions in brain tumors. FEBS Lett 589(14):1588–1597
Carracedo A, Ma L, Teruya-Feldstein J, Rojo F, Salmena L, Alimonti A, Egia A, Sasaki AT, Thomas G, Kozma SC, Papa A, Nardella C, Cantley LC, Baselga J, Pandolfi PP (2008) Inhibition of mTORC1 leads to MAPK pathway activation through a PI3K-dependent feedback loop in human cancer. J Clin Invest 118(9):3065–3074. doi:10.1172/JCI34739
Cassidy J, Tabernero J, Twelves C, Brunet R, Butts C, Conroy T, Debraud F, Figer A, Grossmann J, Sawada N, Schöffski P, Sobrero A, Van Cutsem E, Díaz-Rubio E (2004) Xelox (capecitabine plus oxaliplatin): active first-line therapy for patients with metastatic colorectal cancer. J Clin Oncol 22(11):2084–2091. doi:10.1200/JCO.2004.11.069
Challis GB, Stam HJ (1900) The spontaneous regression of cancer. a review of cases from, 1900 to 1987. Acta Oncol 29(5):545–550
Cheah PY, Choo PH, Yao J, Eu KW, Seow-Choen F (2002) A survival-stratification model of human colorectal carcinomas with beta-catenin and p27kip1. Cancer 95(12):2479–2486. doi:10.1002/cncr.10986
Chen Y, Hughes-Fulford M (2001) Human prostate cancer cells lack feedback regulation of low-density lipoprotein receptor and its regulator, srebp2. Int J Cancer 91(1):41–45
Chinnaiyan AM, Prasad U, Shankar S, Hamstra DA, Shanaiah M, Chenevert TL, Ross BD, Rehemtulla A (2000) Combined effect of tumor necrosis factor-related apoptosis-inducing ligand and ionizing radiation in breast cancer therapy. Proc Natl Acad Sci USA 97(4):1754–1759. doi:10.1073/pnas.030545097
Ciurea M, Georgescu A, Purcaru S, Artene S, Emami G, Boldeanu M, Tache D, Dricu A (2014) Cancer stem cells: biological functions and therapeutically targeting. Int J Mol Sci 15(5):8169–8185
Clevers H, Loh KM, Nusse R (2014) Stem cell signaling. An integral program for tissue renewal and regeneration: Wnt signaling and stem cell control. Science 346(6205):1248,012. doi:10.1126/science.1248012
Crosnier C, Stamataki D, Lewis J (2006) Organizing cell renewal in the intestine: stem cells, signals and combinatorial control. Nat Rev Genet 7(5):349–359. doi:10.1038/nrg1840
Dunn GP, Bruce AT, Ikeda H, Old LJ, Schreiber RD (2002) Cancer immunoediting: from immunosurveillance to tumor escape. Nat Immunol 3(11):991–998. doi:10.1038/ni1102-991
Enderling H, Anderson A, Chaplain M, Beheshti A, Hlatky L, Hahnfeld P (2009) Paradoxical dependencies of tumor dormancy and progression on basic cell kinetics. Cancer Res 69(22):8814–8821
Everson TC, Cole WH (1968) Spontaneous regression of cancer. JB Saunders and Co, Philadelphia
Freeman M (2000) Feedback control of intercellular signalling in development. Nature 408(6810):313–319. doi:10.1038/35042500
González-Sancho JM, Aguilera O, García JM, Pendás-Franco N, Peña C, Cal S, García de Herreros A, Bonilla F, Muñoz A (2005) The wnt antagonist DICKKOPF-1 gene is a downstream target of beta-catenin/TCF and is downregulated in human colon cancer. Oncogene 24(6):1098–1103. doi:10.1038/sj.onc.1208303
Greene WH (2003) Econometric analysis, 5th edn. Prentice Hall, Upper Saddle River
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100(1):57–70
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674. doi:10.1016/j.cell.2011.02.013
Hillen T, Enderling H, Hahnfeldt P (2013) The tumor growth paradox and immune system-mediated selection for cancer stem cells. Bull Math Biology 75(1):161–184
Hobohm U (2001) Fever and cancer in perspective. Cancer Immunol Immunother 50(8):391–396
Iwasa Y, Nowak M, Michor F (2006) Evolution of resistance during clonal expansion. Genetics 172:2557–2566
Jessy T (2011) Immunity over inability: the spontaneous regression of cancer. J Nat Sci Biol Med 2(1):43–49. doi:10.4103/0976-9668.82318
Korolev KS, Xavier JB, Gore J (2014) Turning ecology and evolution against cancer. Nat Rev Cancer 14(5):371–380. doi:10.1038/nrc3712
Krausova M, Korinek V (2014) Wnt signaling in adult intestinal stem cells and cancer. Cell Signal 26(3):570–579. doi:10.1016/j.cellsig.2013.11.032
Kwak EL, Clark JW, Chabner B (2007) Targeted agents: the rules of combination. Clin Cancer Res 13(18 Pt 1):5232–5237. doi:10.1158/1078-0432.CCR-07-1385
Lander AD, Gokoffski KK, Wan FYM, Nie Q, Calof AL (2009) Cell lineages and the logic of proliferative control. PLoS Biol 7(1):e15. doi:10.1371/journal.pbio.1000015
Malhotra V, Perry MC (2003) Classical chemotherapy: mechanisms, toxicities and the therapeutic window. Cancer Biol Ther 2(4 Suppl 1):S2–4
Mangsbo SM, Sandin LC, Anger K, Korman AJ, Loskog A, Tötterman TH (2010) Enhanced tumor eradication by combining ctla-4 or pd-1 blockade with cpg therapy. J Immunother 33(3):225–235. doi:10.1097/CJI.0b013e3181c01fcb
Massagué J, Blain SW, Lo RS (2000) Tgfbeta signaling in growth control, cancer, and heritable disorders. Cell 103(2):295–309
Matchett K, Lappin T (2014) Concise reviews: cancer stem cells: from concept to cure. Stem Cells 32(10):2563–2570
Mathijssen RHJ, Sparreboom A, Verweij J (2014) Determining the optimal dose in the development of anticancer agents. Nat Rev Clin Oncol 11(5):272–281. doi:10.1038/nrclinonc.2014.40
Meacham C, Morrison S (2013) Tumour heterogeneity and cancer cell plasticity. Nature 501(7467):328–337
Mertins S (2014) Cancer stem cells: a systems biology view of their role in prognosis and therapy. Anticancer Drugs 25(4):353–367
Najdi R, Holcombe RF, Waterman ML (2011) Wnt signaling and colon carcinogenesis: beyond apc. J Carcinog 10:5. doi:10.4103/1477-3163.78111
Nakamura T, Matsumoto K, Kiritoshi A, Tano Y, Nakamura T (1997) Induction of hepatocyte growth factor in fibroblasts by tumor-derived factors affects invasive growth of tumor cells: in vitro analysis of tumor-stromal interactions. Cancer Res 57(15):3305–3313
O’Connor M, Xiang D, Shigdar S, Macdonald J, Li Y, Wang T, Pu C, Wang Z, Qiao L, Duan W (2014) Cancer stem cells: a contentious hypothesis now moving forward. Cancer Lett 344(2):180–187
Perko L (2001) Differential equations and dynamical systems, vol 7, (3rd edn). Springer, New York. http://www.loc.gov/catdir/enhancements/fy0826/00058305-d.html
Pinto D, Clevers H (2005) Wnt, stem cells and cancer in the intestine. Biol Cell 97(3):185–196. doi:10.1042/BC20040094
Poleszczuk J, Hahnfeldt P, Enderling H (2015) Evolution and phenotype selection of cancer stem cells. PLoS Comput Biol 11(3):e1004,025
Radtke F, Clevers H (2005) Self-renewal and cancer of the gut: two sides of a coin. Science 307(5717):1904–1909. doi:10.1126/science.1104815
Reya T, Clevers H (2005) Wnt signalling in stem cells and cancer. Nature 434(7035):843–850. doi:10.1038/nature03319
Rhodes A, Hillen T (2016) Mathematical modelling of the role of survivin on dedifferentiation and radioresistance in cancer (submitted)
Scott J, Hjelmeland A, Chinnaiyan P, Anderson A (2014) Microenvironmental variables must influence intrinsic phenotype parameters of cancer stem cells to affect tumourigenicity. PLoS Comput Biol 10(1):e1003,433
Sottoriva A, Verhoeff J, Borovski T, McWeeney S, Naurnov L, Medema J, Sloot P, Vermuelen L (2010) Cancer stem cell tumor model reveals invasive morphology and increased phenotypical heterogeneity. Cancer Res 70(1):46–56
Sottoriva A, Vermuelen L, Tavare S (2011) Modeling evolutionary dynamics of epigenetic mutations in hierarchically organized tumors. PLoS Comput Biol 7(5):e1001,132
Stephens PA, Sutherland WJ, Freckleton RP (1999) What is the Allee effect? Oikos 87(1):185–190
Uno T, Takeda K, Kojima Y, Yoshizawa H, Akiba H, Mittler RS, Gejyo F, Okumura K, Yagita H, Smyth MJ (2006) Eradication of established tumors in mice by a combination antibody-based therapy. Nat Med 12(6):693–698. doi:10.1038/nm1405
Youssefpour H, Li X, Lander AD, Lowengrub JS (2012) Multispecies model of cell lineages and feedback control in solid tumors. J Theor Biol 304:39–59. doi:10.1016/j.jtbi.2012.02.030
Zhou BBS, Zhang H, Damelin M, Geles KG, Grindley JC, Dirks PB (2009) Tumour-initiating cells: challenges and opportunities for anticancer drug discovery. Nat Rev Drug Discov 8(10):806–823. doi:10.1038/nrd2137
Acknowledgments
AK gratefully acknowledges support from Predoctoral Training Grant T32HD060555 from the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NIH/NICHD). JL would like to thank the National Institute of Health, National Cancer Institute, for funding through the Grants P50GM76515 for a Center of Excellence in Systems Biology at the University of California, Irvine, and P30CA062203 for the Chao Comprehensive Cancer Center at the University of California, Irvine. JL also gratefully acknowledges support from the National Science Foundation, Division of Mathematical Sciences. The research of TH is supported by NSERC. Part of this work was carried out during a research visit of TH to the Mathematical Biosciences Institute (MBI) in Ohio, USA. The authors are grateful to the anonymous referees for their constructive comments and suggestions.
Author information
Authors and Affiliations
Corresponding authors
Appendices
Appendix 1: Approximation of the Separatrix for System (9) using the Stable Manifold Theorem.
We use the Stable Manifold Theorem (SMT) to prove that the separatrix described in Theorem 1 near the equilibrium point \(\mathbf {P_2}(S_2,A_2)\) of system (9) exists and to approximate it. We follow the technique presented in (Perko 2001). We recall that \(\mathbf {P_2}\) occurs at the unique intersection of the curves \(\{p(S,a) = 0.5\}\) and \(\{F(S,a) = 0\}\). We state the SMT here for reference.
Theorem 2
(The Stable Manifold Theorem). Let E be an open subset of \({\mathbb {R}}^n\) containing the origin, let \(\mathbf {f} \in C^1(E)\), and let \(\phi _t\) be the flow of the nonlinear system \(\dot{\mathbf {x}} = \mathbf {f(x)}\). Suppose that \(\mathbf {f(0) = 0}\) and that \(D\mathbf {f(0)}\) has k eigenvalues with negative real part and \(n-k\) eigenvalues with positive real part. Then there exists a k-dimensional differentiable manifold M tangent to the stable subspace \(E^m\) of the linear system \(\dot{\mathbf {x}} = \mathbf {Ax}\) at \(\mathbf {0}\) where \(\varvec{A} = D \mathbf {f(0)}\), such that for all \(t \ge 0\), \(\phi _t(M) \subset M\) for all \(\mathbf {x_0} \in M\) and
In our case, we make an affine change of coordinates to system (9) which sends \(\mathbf {P_2} \rightarrow {\mathbf {0}}\) and use the constructive proof of the SMT (see Perko 2001, p. 108) to find the separatrix M.
1.1 Affine Change of Coordinates
To apply the SMT, we need to first make the affine change of coordinates: \(\mathbf {c}: (S,a) \rightarrow (S,a) - (S_2,A_2)\). We let \((S^*,a^*) = \mathbf {c}(S,a)\). Then, applying \(\mathbf {c}\) to (9), and noting that \((S,a) = (S^*,a^*) + (S_2,A_2)\), and \(\frac{\partial }{\partial t}(S,a) = \frac{\partial }{\partial t}((S^*,a^*)+(S_2,A_2)) = \frac{\partial }{\partial t}(S^*,a^*)\), we obtain
The Jacobian for (16) is
where
In this coordinate system, \((S^*,a^*) = (0,0)\) is an equilibrium point and \(P^*(0,0)\) corresponds to \(\mathbf {P_2}(S_2,A_2)\). To use the SMT, we need to first determine \(\varvec{A} = D\mathbf {f}(\mathbf {0}) = \varvec{J}^*(0,0)\). We have
We first note, as in the original \(\varvec{J}(S_1,S_2)\), that since \(p^*_{S^*} <0\) and \((f^*_2)_{a^*}(0,0)>0\), \(2p^*_{S^*} (0,0) k S_2(f_2)_{a^*} (0,0) <0\) and since \(p^*_{a^*}>0\) and \((f_2^*)_{S^*}>0\), \( 2p^*_{a^*} (0,0) k S_2 (f_2)_{S^*} (0,0) >0\). Therefore,
and hence \(\varvec{J}^*(0,0)\) has one positive and one negative eigenvalue, and \(P^*\) is a saddlepoint. We also recall that \(S_2\) and \(A_2\) satisfy
Using (19), we simplify (17) to calculate the elements of \({\varvec{A}}\),
Substituting (20) into (18), we have the following expression for \(\varvec{A}=\varvec{J}^*(0,0)\),
1.2 Preliminary Calculations for the SMT
Following (Perko 2001) and taking \(\mathbf {x}= (S^*,a^*)\), we can rewrite the system (16) as
where \(\varvec{A}=\varvec{J}^*(0,0)\) and \(\mathbf {F(x)} = \mathbf {f^*(x)} - \mathbf {Ax}\). We next need to find an invertible matrix \(\varvec{C}\) such that
where \(L_1\) and \(L_2\) are the negative and positive eigenvalues, respectively, of \(\varvec{A}=(A_{ij})\). We first calculate the trace, T, and determinant, D, of A,
We note, from the calculations above, that \(D<0\). We proceed to calculate \(0 = \det (A - L I)\) to obtain the quadratic equation
The quadratic formula gives us:
Since \(D<0\), we find that \(L_{1,2}\) are both real and have opposite sign; hence, \(L_1 < 0 <L_2\). It can be verified that \(\mathbf {v}_1 = [(L_1 - A_{22}), A_{21}]^T\) and \(\mathbf {v}_2 = [ (L_2 - A_{22}), A_{21}]^T\) are eigenvectors corresponding (respectively) to \(L_1\) and \(L_2\). Here the superscript T denotes the transpose. Therefore, we have
We make another change of variables, taking \(\mathbf {y}=\varvec{C}^{-1}(\mathbf {x})\), and writing (22) as
where \(\varvec{B}\) is from (23) and \(\mathbf {G(y)} = \mathbf {C^{-1}F(Cy)}\).
1.3 Applying the SMT
By the SMT (taking \(\mathbf {a}=(a_1,a_2)\)),
is the solution to (24), where
We solve for \(\mathbf {u}\) using the method of successive approximation. We let \(\mathbf {u}^{(0)} (t,\mathbf {a}) = \mathbf {0}\) and
To solve for \(j=1\), we note that \(\mathbf {G(0)}= \mathbf {{C}^{-1} F(C\cdot 0)} = \mathbf {{C}^{-1}F(\mathbf {0}) }= \mathbf {0}\) since \(f_1(0,0) = f_2(0,0) = 0\). Therefore,
For the next approximation, we first calculate \(\varvec{U}(t-s) \mathbf {G}(\mathbf {u}^{(1)}(s,\mathbf {a})) = \varvec{U}(t-s) \varvec{C}^{-1} \mathbf {F(Cw)} = \varvec{H}_1 \mathbf {F(Cw)}\), where \(\varvec{H}_1 = \varvec{U}(t-s)\varvec{C}^{-1}\) and \(\mathbf {w}= (e^{L_1 s} a_1, 0)^T \). Simplifying \(\varvec{H}_1\) gives us:
Then,
Hence,
We note that our stable manifold will be of the form \(y_2= \psi ^{(2)}_2(y_1)\), where \(\psi ^{(2)}_2(a_1) = u^{(2)}_2(0,a_1,0)\). Since \(\mathbf {U}(t)\mathbf {a}\) and (28) only contribute trivially to \(u^{(2)}_2\), we will not perform further calculations on them.
Next, we calculate \(\varvec{V}(t-s) \mathbf {G}(\mathbf {u}^{(1)}(s,\mathbf {a})) = \varvec{V}(t-s) \varvec{C}^{-1} \mathbf {F}(\mathbf {Cw}) = \varvec{H}_2 \mathbf {F}(\mathbf {Cw})\). As before, we first calculate \(\varvec{H}_2\):
We thus have,
Taking the integral of the right-hand term on the domain \([t,\infty )\) gives us \(\frac{e^{L_1 t}}{L_2-L_1}(0, g(\cdot ))^T\). We find that \(g(\cdot ) = L_1^2 - TL_1 + D = 0\). Therefore, this term does not contribute to the stable manifold.
The SMT allows us to calculate the second approximation to the separatrix, \(\hbox {M}^* = u_2^{(2)}(0,a_1,0)\), as
where \(\mathbf {Cw} = e^{L_1 s} a_1 (L_1 - A_{22}, A_{21})^T\), and by (16),
We now solve \(- \int _0^{\infty } I_1 ds=\int _0^\infty e^{-L_2s}f^*_1 (\mathbf {Cw}) ds \) and \(\int _0^\infty I_2ds=\int _0^\infty e^{-L_2s} f^*_2 (\mathbf {Cw}) ds \). Substituting (31) into \(I_1\), we obtain
We split \(I_1\) into three parts:
We can directly integrate \(I_{12}\) and \(I_{13}\) to obtain
We now work to simplify \(I_{11}\) by taking \(u = e^{L_1 s}\). The change of variables gives us
where we find that \(u_1 = 1\) and \(u_2 = \lim _{s\rightarrow \infty } e^{L_1 s} = 0\) since \(L_1 < 0\). We take \(c_1 = a_1 A_{21}\) and \(c_2 = a_1(L_1 - A_{22})\).
We would like to do a partial fraction decomposition for the integrand term in (35) not containing \(u^{-T/L_1}\), \(I_{11}^*\). Noting that both the numerator and denominator are of degree 2, we first perform long division to obtain a fraction p / q where deg \(p< \) deg q. Fully multiplying the terms in the fraction, and setting \(c_3 = A_2c_2 + S_2c_1\) and \(c_4 = A_2 \xi _1 + S_2 \xi _2 + A_2S_2 \xi _1 \xi _2 +1\) gives us
Setting \(c_5 = c_2 / \xi _1 + c_1 / \xi _2\) and \(c_6 = A_2 / \xi _2 + S_2/ \xi _1 + 1/(\xi _1 \xi _2)\), the second term of (36) becomes
where
Therefore, the integrand in (35), \(u^{-T/L_1}I_{11}^*\), can be written as
We note that the first term can be integrated,
where the first equality comes about from the observation that \(-T/L_1 = -1 - L_2/L_1\). To summarize, if we set
we can rewrite (34) as
To solve \(\int _0^1 I^{**}_{11} du\), we will need to use a hypergeometric function and the beta function. Indeed, we have the formula
where \(B(a,b) = \int _0^a t^{a-1}(1-t)^{b-1} dt\) and \(_2F_1(a_1,a_2;b_1;x) = \sum _{k=0}^\infty \frac{(a_1)_k(a_2)_k}{(b_1)_k} \frac{x^k}{k!}\). In our case, we split \(I^{**}_{11}\) naturally as a sum of two terms, and for the first integral, we have \(b-1 = -T/L_1\), hence \(b = 1-T/L_1 = -L_2/L_1\), \(0=c-b-1\), hence \(c = b+1 = 2-T/L_1\), \(a=1\), and \(x = (-\xi _1 C_1)/(\xi _1 A_2 +1)\), where we have pulled \((\xi _1 A_2 +1)^{-1}\) from the denominator. We want to first find an explicit representation for \(B(b,c-b)\),
Using the hypergeometric function and (37), our final formula for \(-\int _0^\infty I_1 ds = \int _0^\infty -e^{L_2 s} f_1^* (\mathbf {Cw})ds\) is
where
and
We now begin work to solve \(\int _0^\infty I_2 ds = \int _0^\infty e^{-L_2 s} f_2(\mathbf {Cw}) ds\). Using the same substitution as earlier, i.e., \(u = e^{L_1 s}\), and again taking \(c_1 = a_1 A_{21}\) and \(c_2 = a_1 (L_1 - A_{22})\), we obtain
where the last equality comes from long division in the first integral and full integration of the second. The constants are as follows:
We concentrate now on the first integral in (39). The first three terms multiplied by \(u^{-T/L_1}\) can be integrated in a straightforward manner. The last one can be integrated using a hypergeometric function as described earlier. We thus obtain
where \(_2F_1^3 = {}_2 F_1\left( 1, -L_2/L_1; 1-L_2/L_1; \frac{-\lambda c_1}{1+ \lambda A_2}\right) \). Therefore, using (30), (38) and (40) we obtain an explicit solution for \(M^*\),
where the constants and hypergeometric functions were specified earlier.
1.4 Linear and Quadratic Approximation of \(M^*\)
We take a linear and quadratic portion of \(M^*\) in order to more easily ascertain the effects of the system parameters. Using the expansion for the hypergeometric function, and noting that the untransformed stable manifold will intersect (0, 0), we remove all nonlinear terms and rewrite \(M^*\) as \(y_2 = m y_1\), where m is the slope of the line \(y_2\) to obtain
where \(c_7\) is the constant
where \(c_4^{**} = c_4^*/y_1\) and \(c_6^{**} = c_6^*/y_1\). Next, since \((S^*, a^*) = \mathbf {x} = \mathbf {Cy}\), we can obtain
Keeping all quadratic terms, we obtain
where
The system can be solved for \(x = (S^*,a^*)\) using the quadratic formula.
In Fig. 1, we plot \(M^*\), \(M^*_l\) and \(M^*_q\) for specific parameter values in the original coordinate system (S, a). To plot \(M^*\) and \(M^*_q\) in the original coordinate system (S, a), we input a discreet set of values for \(y_1\) and use Eqs. (41) or (44), respectively, to find \(y_2\). We can then find \((S^*,a^*) = \mathbf {C^{-1}y}\) and hence \((S,a) = (S^*+S_2,a^*+A_2)\). To plot \(M^*_l\) in the original coordinate system (S, a), we directly find \((S^*,a^*)\) using Eq. (42) and translate to (S, a). To simplify notation, when we refer to \(M^*\), \(M^*_l\) and \(M^*_q\) in the main text, we are referring to these functions after coordinate transformation to (S, a).
Appendix 2: Calculation of the Coefficient of Determination, \(R^2\)
Here, we briefly describe the calculation of the coefficient of determination, or \(R^2\), as a measure of the goodness of fit of a regression model to the data. One can find more in depth development and analysis of \(R^2\) in Greene (2003).
The coefficient of determination is calculated as follows. Define the dependent variables as \(\{y_i\}_{i=1}^n\) (in our case, the \(y_i\) correspond to the values of \(A_I\) for a given parameter input); taking \(\bar{y} = \sum _{i} y_i / n\), we define the total variation in y as
which is just the sum of squared deviations. For each \(y_i\), we associate a \(b_i\), which is the value of the regression equation at independent variable \(x_i\). We similarly define the residual sum of squares as
The unadjusted \(R^2\) is calculated as
Note that if the data perfectly fit the model (i.e., \(y_i = b_i\) \(\forall i\)), then \(R^2=1\), indicating that a model with a perfect fit to the data has \(R^2=1\), and decreases to 0 as the goodness of fit is reduced. The adjusted \(R^2\), \(\bar{R^2}\), corrects to an increase in \(R^2\) that can occur due to incorporation of additional degrees of freedom into a model and thus should be used in lieu of \(R^2\) when comparing goodness of fit between models with different degrees of freedom. It is defined as
where n is the total number of observations, d is the number of regression coefficients, \(n-d\) is the degrees of freedom of \(SS_{res}\), and \(n-1\) is the degrees of freedom of \(SS_{t}\). In the main text, the adjusted \(R^2\) is presented without the bar notation.
Rights and permissions
About this article
Cite this article
Konstorum, A., Hillen, T. & Lowengrub, J. Feedback Regulation in a Cancer Stem Cell Model can Cause an Allee Effect. Bull Math Biol 78, 754–785 (2016). https://doi.org/10.1007/s11538-016-0161-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-016-0161-5