Abstract
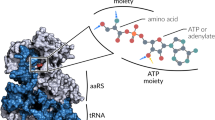
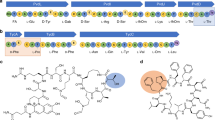
THE aminoacyl-transfer RNA synthetases (aaRS) catalyse the attachment of an amino acid to its cognate transfer RNA molecule in a highly specific two-step reaction. These proteins differ widely in size and oligomeric state, and have limited sequence homology. Out of the 18 known aaRS, only 9 (ref. 1), referred to as class I synthetases (GlnRS, TyrRS, MetRS, GluRS, ArgRS, ValRS, IleRS, LeuRS, TrpRS), display two short common consensus sequences ('HIGH' and 'KMSKS') which indicate, as observed in three crystal structures2–4, the presence of a structural domain (the Rossman fold) that binds ATP. We report here the sequence of Escherichia coll ProRS, a dimer of relative molecular mass 127,402, which is homologous to both ThrRS and SerRS. These three latter aaRS share three new sequence motifs with AspRS, AsnRS, LysRS, HisRS and the β subunit of PheRS. These three motifs (motifs 1, 2 and 3), in a search through the entire data bank, proved to be specific for this set of aaRS (referred to as class II). Class II may also contain AlaRS and GlyRS, because these sequences have a typical motif 3. Surprisingly, this partition of aaRS in two classes is found to be strongly correlated on the functional level with the acylation occurring either on the 2′ OH (class I) or 3′ OH (class II) of the ribose of the last nucleotide of tRNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Burbaum, J., Starzyk, R. M. & Schimmel, P. Proteins 7, 99–111 (1990).
Brick, P., Bhat, T. N. & Blow, D. M. J. molec. Biol. 208, 83–98 (1988).
Zelwer, C., Risler, J.-L. & Brunie, S. J. molec. Biol. 155, 63–81 (1982).
Rould, M. A., Perona, J. J., Söll, D. & Steitz, T. A. Science 246, 1135–1142 (1989).
Bohman, K. & Isaksson, L. A. Molec. gen. Genet. 177, 603–605 (1980).
Dale, R. M. K., McClure, B. A. & Houchins, J. P. Plasmid 13, 31–40 (1985).
Tabor, S. & Richardson, C. C. Proc. natn. Acad. Sci. U.S.A. 84, 4767–4771 (1987).
Brendel, V. & Trifonov, E. V. Nucleic Acids Res. 10, 4411–4427 (1984).
Springer, M., Graffe, M., Dondon, J. & Grunberg-Manago, M. EMBO J. 8, 2417–2427 (1989).
Moine, H. thesis., Univ. Louis Pasteur, Strasbourg (1990).
Molina, A. J., Peterson, R. & Yang, D.C.H. J. biol. Chem. 264, 16608–16612 (1989).
Gampel, A. & Tzagoloff, A. Proc. natn. Acad Sci. U.S.A. 86, 6023–6027 (1989).
Anselme, J. & Härtlein, M. Gene 84, 481–485 (1989).
Leveque, F., Plateau, P., Dessen, P. & Blanquet, S. Nucleic. Acids Res. 18, 305–312 (1990).
Wek, R. C., Jackson, B. M. & Hinnenbusch, A. G. Proc. natn. Acad. Sci. U.S.A. 86, 4579–4583 (1989).
Gribskov, M., MacLachlan, A. D. & Eisenberg, D. Proc. natn. Acad. Sci. U.S.A. 84, 4355–4358 (1987).
Jasin, M., Regan, L. & Schimmel, P. R. Nature 306, 441–447 (1983).
Prevost, G., Eriani, G., Kern, D., Dirheimer, G. & Gangloff, J. Eur. J. Biochem. 180, 351–358 (1989).
Eriani, G. thesis., Univ. Louis Pasteur, Strasbourg (1990).
Hecht, S. M. in Transfer-RNA Structure, Properties and Recognition (eds Schimmel, P. R., Söll, D. & Abelson, J. N.) 345–360 (Cold Spring Harbor Laboratory, New York, 1979).
Weiner, A. M. & Maizels, N. Proc. natn. Acad. Sci. U.S.A. 84, 7383–7387 (1987).
Fraser, T. H. & Rich, A. Proc. natn. Acad. Sci. U.S.A. 72, 3044–3048 (1975).
von der Haar, F. & Cramer, F. Biochemistry 15, 4131–4136 (1976).
Fersht, A. R. & Kaethner, M. M. Biochemistry 15, 3342–3346 (1976).
Brune, M., Schumann, R. & Wittinghofer, F. Nucleic Acids Res. 13, 7139–7147 (1985).
Devereux, J., Haeberli, P. & Smithies, O. Nucleic Acids Res. 12, 387–395 (1984).
Eriani, G., Dirheimer, G. & Gangloff, J. Nucleic Acids Res. 17, 5725–5736 (1989).
Berger, S. L., Wallace, D. M., Puskas, R. S. & Eschenfeldt, W. H. Biochemistry 22, 2365–2373 (1983).
Argos, P. J. molec. Biol. 193, 385–396 (1987).
Mirande, M. & Waller, J. P. J. biol. Chem. 263, 18443–18451 (1988).
Nilssen, T. W. et al. Proc. natn. Acad. Sci. U.S.A. 85, 3604–3607 (1988).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Eriani, G., Delarue, M., Poch, O. et al. Partition of tRNA synthetases into two classes based on mutually exclusive sets of sequence motifs. Nature 347, 203–206 (1990). https://doi.org/10.1038/347203a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/347203a0
This article is cited by
-
Computational Insight into Substrate-Induced Conformational Changes in Methionyl-tRNA Synthetase of Mycobacterium Tuberculosis
The Protein Journal (2023)
-
Symmetrical distributions of aminoacyl-tRNA synthetases during the evolution of the genetic code
Theory in Biosciences (2023)
-
Establishing a substrate-assisted mechanism for the pre-transfer editing in SerRS and IleRS: a QM/QM investigation
Structural Chemistry (2023)
-
Role of Mutations of Mitochondrial Aminoacyl-tRNA Synthetases Genes on Epileptogenesis
Molecular Neurobiology (2023)
-
Metabolic engineering of Escherichia coli BW25113 for the production of 5-Aminolevulinic Acid based on CRISPR/Cas9 mediated gene knockout and metabolic pathway modification
Journal of Biological Engineering (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.