Abstract
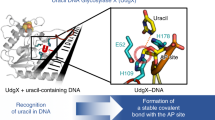
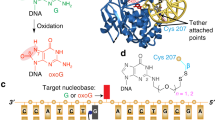
The 1.75-Å crystal structure of the uracil-DNA glycosylase from herpes simplex virus type-1 reveals a new fold, distantly related to dinucleotide-binding proteins. Complexes with a trideoxynucleotide, and with uracil, define the DNA-binding site and allow a detailed understanding of the exquisitely specific recognition of uracil in DNA. The overall structure suggests binding models for elongated single- and double-stranded DNA substrates. Conserved residues close to the uracil-binding site suggest a catalytic mechanism for hydrolytic base excision.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Lindahl, T. Proc. natn. Acad. Sci. U.S.A. 71, 3649–3653 (1974).
Kaboev, O. K., Luchinka, L. A. & Kuziakina, T. I. J. Bact. 164, 421–424 (1985).
Crosby, B., Prakash, L., Davis, H. & Hinkle, D. C. Nucleic Acids Res. 9, 5797–5809 (1981).
Olsen, L. C., Aasland, R., Wittwer, C. U., Krokan, H. E. & Helland, D. E. EMBO J. 9, 3121–3125 (1989).
Upton, C., Stuart, D. T. & McFadden, G. Proc. natn. Acad. Sci. U.S.A. 90, 4518–4522 (1993).
Mullaney, J., Moss, H. W. McL. & McGeogh, D. J. J. gen. Virol. 70, 449–454 (1989).
Caradonna, S., Worrad, D. & Lirette, R. J. Virol. 61, 3040–3047 (1987).
Baer, R. et al. Nature 310, 207–211 (1984).
Davison, A. J. & Scott, J. E. J. gen. Virol. 67, 1759–1816 (1986).
Cone, R., Duncan, J., Hamilton, L. & Friedberg, E. C. Biochemistry 16, 3194–3201 (1977).
Lindahl, T., Ljungquist, S., Siegert, W., Nyberg, B. & Sperens, B. J. biol. Chem. 252, 3286–3294 (1977).
Dianov, G. et al. Nucleic Acids Res. 22, 993–998 (1994).
Lindahl, T. Prog. Nucleic Acid Res. molec. Biol. 22, 135–192 (1979).
Mosbaugh, D. W. Rev. biochem. Tox. 9, 69–130 (1988).
Verri, A., Mazzarello, P., Biamonti, G., Spadari, S. & Focher, F. Nucleic Acids Res. 18, 5775–5780 (1990).
Focher, F., Verri, A., Verzeletti, S., Mazzarello, P. & Spadari, S. Chromosoma. Berl. 102 (Suppl.), S67–S71 (1992).
Slupphaug, G. et al. Nucleic Acids Res. 21, 2579–2584 (1993).
Mauro, D. J., De Reil, J. K., Tallarida, R. J. & Sirover, M. A. Molec. Pharmac. 43, 854–857 (1993).
Hatahet, Z., Kow, Y. W., Purmal, A. A., Cunningham, R. P. & Wallace, S. S. J. biol. Chem. 269, 18814–18820 (1994).
Eftedal, I., Guddal, P. H., Slupphaug, G., Volden, G. & Krokan, H. E. Nucleic Acids Res. 21, 2095–2101 (1993).
Cone, R., Duncan, J., Hamilton, L. & Friedberg, E. C. Biochemistry 16, 3194–3201 (1977).
Lindahl, T., Ljungquist, S., Siegert, W., Nyberg, B. & Sperens, B. J. biol. Chem. 252, 3286–3294 (1977).
Krokan, H. & Wittwer, C. U. Nucleic Acids Res. 9, 2599–2613 (1981).
Pyles, R. B. & Thompson, R. L. J. Virol. 68, 4963–4972 (1994).
Sawa, R. & Pearl, L. H. J. molec. Biol. 234, 910–912 (1993).
Rossman, M. G. et al. in The Enzymes Vol. 11 (ed. Boyer, P. D.) 61–102 (Academic, NewYork, 1975).
Klimasaukas, S., Kumar, S., Roberts, R. J. & Cheng, X. Cell 76, 357–369 (1994).
Bennet, S. E., Jensen, O. N., Barofsky, D. F. & Mosbaugh, D. W. J. biol. Chem. 269, 21870–21879 (1994).
Krokan, H. E. et al. Proc. N.Y. Acad. Sci. (Abstr.) (July 31-Aug. 4, Burlington, VT, 1993).
Varshney, U. & van de Sande, J. H. Biochemistry 30, 4055–4061 (1991).
Leslie, A. G. W. MOSFLM Users Guide (MRC-LMB, Cambridge, 1994).
Pflugrath, J. W. & Messerschmidt, A. Munich Area Detector New EEC System (1992).
Collaborative Computational Project No. 4. Acta crystallogr. D50, 760–763 (1994).
Otwinowski, Z. in Isomorphous Replacement and Anomalous Scattering: Proceedings of the CCP4 Study Weekend 23–38 (SERC Daresbury Laboratory, 1991).
Jones, T. A., Zou, J.-Y., Cowan, S. W. & Kjeldgaard, M. Acta crystallogr. A47, 110–119 (1991).
Kleywegt, G. J. & Jones, T. A. in From First Map to Final Model: Proceedings of the CCP4 Study Weekend (eds Bailey, S. et al.) 59–66 (SERC Daresbury Laboratory, 1994).
Read, R. J. Acta crystallogr. A42, 140–149 (1986).
Brunger, A. T. X-PLOR Version 3.1. A System for X-Ray Crystallography and NMR (Yale Univ. Press, New Haven, CT, 1992).
Laskowski, R. J., MacArthur, M. W., Moss, D. S. & Thornton, J. M. J. Appl. crystallogr. 26, 283–290 (1993).
Kraulis, P. J. J. Appl. Crystallogr. 24, 946–950 (1991).
Flores, T., Moss, D. S. & Thornton, J. M. Protein Engng 7, 31–37 (1994).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Savva, R., McAuley-Hecht, K., Brown, T. et al. The structural basis of specific base-excision repair by uracil–DNA glycosylase. Nature 373, 487–493 (1995). https://doi.org/10.1038/373487a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/373487a0
This article is cited by
-
Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors
Nature Biotechnology (2020)
-
Covalent binding of uracil DNA glycosylase UdgX to abasic DNA upon uracil excision
Nature Chemical Biology (2019)
-
Specificity and Efficiency of the Uracil DNA Glycosylase-Mediated Strand Cleavage Surveyed on Large Sequence Libraries
Scientific Reports (2019)
-
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
Scientific Reports (2017)
-
5-Formylcytosine does not change the global structure of DNA
Nature Structural & Molecular Biology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.