Abstract
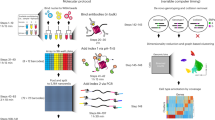
Recent technological advances have enabled massively parallel chromatin profiling with scATAC-seq (single-cell assay for transposase accessible chromatin by sequencing). Here we present ATAC with select antigen profiling by sequencing (ASAP-seq), a tool to simultaneously profile accessible chromatin and protein levels. Our approach pairs sparse scATAC-seq data with robust detection of hundreds of cell surface and intracellular protein markers and optional capture of mitochondrial DNA for clonal tracking, capturing three distinct modalities in single cells. ASAP-seq uses a bridging approach that repurposes antibody:oligonucleotide conjugates designed for existing technologies that pair protein measurements with single-cell RNA sequencing. Together with DOGMA-seq, an adaptation of CITE-seq (cellular indexing of transcriptomes and epitopes by sequencing) for measuring gene activity across the central dogma of gene regulation, we demonstrate the utility of systematic multi-omic profiling by revealing coordinated and distinct changes in chromatin, RNA and surface proteins during native hematopoietic differentiation and peripheral blood mononuclear cell stimulation and as a combinatorial decoder and reporter of multiplexed perturbations in primary T cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Data are available at the Gene Expression Omnibus under accession number GSE156478.
Code availability
Custom code to reproduce all analyses and figures is available at https://github.com/caleblareau/asap_reproducibility.
References
Nam, A. S., Chaligne, R. & Landau, D. A. Integrating genetic and non-genetic determinants of cancer evolution by single-cell multi-omics. Nat. Rev. Genet. 22, 3–18 (2021).
Zhu, C., Preissl, S. & Ren, B. Single-cell multimodal omics: the power of many. Nat. Methods 17, 11–14 (2020).
Schier, A. F. Single-cell biology: beyond the sum of its parts. Nat. Methods 17, 17–20 (2020).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Mimitou, E. P. et al. Multiplexed detection of proteins, transcriptomes, clonotypes and CRISPR perturbations in single cells. Nat. Methods 16, 409–412 (2019).
Peterson, V. M. et al. Multiplexed quantification of proteins and transcripts in single cells. Nat. Biotechnol. 35, 936–939 (2017).
Ma, S. et al. Chromatin potential identified by shared single-cell profiling of RNA and chromatin. Cell 183, 1103–1116 (2020).
Lareau, C. A. et al. Droplet-based combinatorial indexing for massive-scale single-cell chromatin accessibility. Nat. Biotechnol. 37, 916–924 (2019).
Satpathy, A. T. et al. Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral T cell exhaustion. Nat. Biotechnol. 37, 925–936 (2019).
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 361, 1380–1385 (2018).
Chen, S., Lake, B. B. & Zhang, K. High-throughput sequencing of the transcriptome and chromatin accessibility in the same cell. Nat. Biotechnol. 37, 1452–1457 (2019).
Zhu, C. et al. An ultra high-throughput method for single-cell joint analysis of open chromatin and transcriptome. Nat. Struct. Mol. Biol. 26, 1063–1070 (2019).
Lareau, C. A. et al. Massively parallel single-cell mitochondrial DNA genotyping and chromatin profiling. Nat. Biotechnol. 39, 451–461 (2021).
Walker, M. A. et al. Purifying selection against pathogenic mitochondrial DNA in human T cells. N. Engl. J. Med. 383, 1556–1563 (2020).
Granja, J. M. et al. Single-cell multiomic analysis identifies regulatory programs in mixed-phenotype acute leukemia. Nat. Biotechnol. 37, 1458–1465 (2019).
Stoeckius, M. et al. Cell hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics. Genome Biol. 19, 224 (2018).
Gaublomme, J. T. et al. Nuclei multiplexing with barcoded antibodies for single-nucleus genomics. Nat. Commun. 10, 2907 (2019).
McGinnis, C. S. et al. MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices. Nat. Methods 16, 619–626 (2019).
Gehring, J., Hwee Park, J., Chen, S., Thomson, M. & Pachter, L. Highly multiplexed single-cell RNA-seq by DNA oligonucleotide tagging of cellular proteins. Nat. Biotechnol. 38, 35–38 (2020).
Srivatsan, S. R. et al. Massively multiplex chemical transcriptomics at single-cell resolution. Science 367, 45–51 (2020).
Shin, D., Lee, W., Lee, J. H. & Bang, D. Multiplexed single-cell RNA-seq via transient barcoding for simultaneous expression profiling of various drug perturbations. Sci. Adv. 5, eaav2249 (2019).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Kim, H. J., Lin, Y., Geddes, T. A., Yang, J. Y. H. & Yang, P. CiteFuse enables multi-modal analysis of CITE-seq data. Bioinformatics 36, 4137–4143 (2020).
Hao, Y., Hao, S., Andersen-Nissen, E. & Mauck, W. M. Integrated analysis of multimodal single-cell data. Preprint at bioRxiv https://doi.org/10.1101/2020.10.12.335331 (2020).
Ludwig, L. S. et al. Lineage tracing in humans enabled by mitochondrial mutations and single-cell genomics. Cell 176, 1325–1339 (2019).
Lareau, C. A., Ludwig, L. S. & Sankaran, V. G. Longitudinal assessment of clonal mosaicism in human hematopoiesis via mitochondrial mutation tracking. Blood Adv. 3, 4161–4165 (2019).
Ju, Y. S. et al. Somatic mutations reveal asymmetric cellular dynamics in the early human embryo. Nature 543, 714–718 (2017).
Buenrostro, J. D. et al. Integrated single-cell analysis maps the continuous regulatory landscape of human hematopoietic differentiation. Cell 173, 1535–1548 (2018).
Velten, L. et al. Human haematopoietic stem cell lineage commitment is a continuous process. Nat. Cell Biol. 19, 271–281 (2017).
Hartmann, F. J. & Bendall, S. C. Immune monitoring using mass cytometry and related high-dimensional imaging approaches. Nat. Rev. Rheumatol. 16, 87–99 (2019).
Sancho, D., Gómez, M. & Sánchez-Madrid, F. CD69 is an immunoregulatory molecule induced following activation. Trends Immunol. 26, 136–140 (2005).
Caruso, A. et al. Flow cytometric analysis of activation markers on stimulated T cells and their correlation with cell proliferation. Cytometry 27, 71–76 (1997).
Dong, C. et al. ICOS co-stimulatory receptor is essential for T-cell activation and function. Nature 409, 97–101 (2001).
San José, E., Borroto, A., Niedergang, F., Alcover, A. & Alarcón, B. Triggering the TCR complex causes the downregulation of nonengaged receptors by a signal transduction-dependent mechanism. Immunity 12, 161–170 (2000).
Calderon, D. et al. Landscape of stimulation-responsive chromatin across diverse human immune cells. Nat. Genet. 51, 1494–1505 (2019).
Gate, R. E. et al. Genetic determinants of co-accessible chromatin regions in activated T cells across humans. Nat. Genet. 50, 1140–1150 (2018).
Swanson, E., Lord, C., Reading, J. & Heubeck, A. T. Integrated single cell analysis of chromatin accessibility and cell surface markers. Preprint at bioRxiv https://doi.org/10.1101/2020.09.04.283887 (2020).
Corces, M. R. et al. Lineage-specific and single-cell chromatin accessibility charts human hematopoiesis and leukemia evolution. Nat. Genet. 48, 1193–1203 (2016).
Liu, L., Takeda, K. & Akkoyunlu, M. Disease stage-specific pathogenicity of CD138 (syndecan 1)-expressing T cells in systemic lupus erythematosus. Front. Immunol. 11, 1569 (2020).
Tan, H. et al. Integrative proteomics and phosphoproteomics profiling reveals dynamic signaling networks and bioenergetics pathways underlying T cell activation. Immunity 46, 488–503 (2017).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Hogan, P. G. Transcriptional regulation by calcium, calcineurin, and NFAT. Genes Dev. 17, 2205–2232 (2003).
Macian, F. NFAT proteins: key regulators of T-cell development and function. Nat. Rev. Immunol. 5, 472–484 (2005).
Basak, S., Shih, V. F.-S. & Hoffmann, A. Generation and activation of multiple dimeric transcription factors within the NF-κB signaling system. Mol. Cell. Biol. 28, 3139–3150 (2008).
Rubin, A. J. et al. Coupled single-cell CRISPR screening and epigenomic profiling reveals causal gene regulatory networks. Cell 176, 361–376 (2019).
Korkmaz, G. et al. Functional genetic screens for enhancer elements in the human genome using CRISPR–Cas9. Nat. Biotechnol. 34, 192–198 (2016).
Gasperini, M. et al. A genome-wide framework for mapping gene regulation via cellular genetic screens. Cell 176, 1516 (2019).
Simeonov, D. R. et al. Discovery of stimulation-responsive immune enhancers with CRISPR activation. Nature 549, 111–115 (2017).
Fulco, C. P. et al. Activity-by-contact model of enhancer-promoter regulation from thousands of CRISPR perturbations. Nat. Genet. 51, 1664–1669 (2019).
Perkins, D. et al. Regulation of CTLA-4 expression during T cell activation. J. Immunol. 156, 4154–4159 (1996).
Scholzen, T. & Gerdes, J. The Ki-67 protein: from the known and the unknown. J. Cell. Physiol. 182, 311–322 (2000).
Valk, E., Rudd, C. E. & Schneider, H. CTLA-4 trafficking and surface expression. Trends Immunol. 29, 272–279 (2008).
Kaech, S. M. et al. Selective expression of the interleukin 7 receptor identifies effector CD8 T cells that give rise to long-lived memory cells. Nat. Immunol. 4, 1191–1198 (2003).
Soares, A. P. et al. Longitudinal changes in CD4+ T-cell memory responses induced by BCG vaccination of newborns. J. Infect. Dis. 207, 1084–1094 (2013).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Bartosovic, M., Kabbe, M. & Castelo-Branco, G. Single-cell profiling of histone modifications in the mouse brain. Preprint at bioRxiv https://doi.org/10.1101/2020.09.02.279703 (2020).
Wu, S. J. et al. Single-cell analysis of chromatin silencing programs in developmental and tumor progression. Preprint at bioRxiv https://doi.org/10.1101/2020.09.04.282418 (2020).
Pierce, S. E., Granja, J. M. & Greenleaf, W. J. High-throughput single-cell chromatin accessibility CRISPR screens enable unbiased identification of regulatory networks in cancer. Preprint at bioRxiv https://doi.org/10.1101/2020.11.02.364265 (2020).
Swanson, E. et al. TEA-seq: a trimodal assay for integrated single cell measurement of transcription, epitopes, and chromatin accessibility. Preprint at bioRxiv https://doi.org/10.1101/2020.09.04.283887 (2020).
Replogle, J. M. et al. Combinatorial single-cell CRISPR screens by direct guide RNA capture and targeted sequencing. Nat. Biotechnol. 38, 954–961 (2020).
Wroblewska, A. et al. Protein barcodes enable high-dimensional single-cell CRISPR screens. Cell 175, 1141–1155 (2018).
Dixit, A. et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866 (2016).
Jaitin, D. A. et al. Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-seq. Cell 167, 1883–1896 (2016).
Gerlach, J. P. et al. Combined quantification of intracellular (phospho-)proteins and transcriptomics from fixed single cells. Sci. Rep. 9, 1469 (2019).
Chung, H. et al. Simultaneous single cell measurements of intranuclear proteins and gene expression. Preprint at bioRxiv https://doi.org/10.1101/2021.01.18.427139 (2021).
Katzenelenbogen, Y. et al. Coupled scRNA-seq and intracellular protein activity reveal an immunosuppressive role of TREM2 in cancer. Cell 182, 872–885 (2020).
Thurner, B. et al. Generation of large numbers of fully mature and stable dendritic cells from leukapheresis products for clinical application. J. Immunol. Methods 223, 1–15 (1999).
Joung, J. et al. Genome-scale CRISPR–Cas9 knockout and transcriptional activation screening. Nat. Protoc. 12, 828–863 (2017).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Lareau, C. A., Ma, S., Duarte, F. M. & Buenrostro, J. D. Inference and effects of barcode multiplets in droplet-based single-cell assays. Nat. Commun. 11, 866 (2020).
Melsted, P., Ntranos, V. & Pachter, L. The barcode, UMI, set format and BUStools. Bioinformatics 35, 4472–4473 (2019).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Lander, E. S. & Waterman, M. S. Genomic mapping by fingerprinting random clones: a mathematical analysis. Genomics 2, 231–239 (1988).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Stuart, T., Srivastava, A., Lareau, C. & Satija, R. Multimodal single-cell chromatin analysis with Signac. Preprint at bioRxiv https://doi.org/10.1101/2020.11.09.373613 (2020).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Soneson, C. & Robinson, M. D. Bias, robustness and scalability in single-cell differential expression analysis. Nat. Methods 15, 255–261 (2018).
Mulè, M. P., Martins, A. J. & Tsang, J. S. Normalizing and denoising protein expression data from droplet-based single cell profiling. Preprint at bioRxiv https://doi.org/10.1101/2020.02.24.963603 (2020).
Kang, H. M. et al. Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat. Biotechnol. 36, 89–94 (2018).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Acknowledgements
We acknowledge support from the Broad Institute and the Whitehead Institute Flow Cytometry Core facilities. This research was supported by National Institutes of Health grants nos. R01 DK103794 (V.G.S.) and R01 HL146500 (V.G.S.); National Institutes of Health/National Human Genome Research Institute grants nos. R21 HG-009748 (P.S.) and RM1 HG0110014 (P.S.); Grants-in-Aid by the Japan Society for Promotion of Science for Specially Promoted Research no. 16H06295 (S.S.); the Japan Agency for Medical Research and Development for Leading Advanced Projects for Medical Innovation (S.S.); a gift from the Lodish Family to Boston Children’s Hospital (V.G.S.); the New York Stem Cell Foundation (NYSCF) (V.G.S.); the Howard Hughes Medical Institute and Klarman Cell Observatory (A.R.); and the Chan Zuckerberg Initiative/Silicon Valley Community Foundation Human Cell Atlas grant no. HCA3-0000000309 (P.S.). V.G.S. is an NYSCF Robertson Investigator. C.A.L. is supported by a Stanford Science Fellowship. L.S.L is supported by an Emmy Noether fellowship by the German Research Foundation (LU 2336/2-1).
Author information
Authors and Affiliations
Contributions
E.P.M. and P.S. conceived and designed the methods with input from L.S.L., K.Y.C. and C.A.L. E.P.M., K.Y.C., L.S.L. and P.S. designed experiments with input from C.A.L., R.S., V.G.S. and A.R. E.P.M., K.Y.C., E.P., W.L., P.I.T., T.K., M.H. and L.S.L. performed experiments. C.A.L. led data analysis with substantial contributions from E.P.M., K.Y.C., Y.H. and Y.T. A.L.Z.-F., T.-S.H., B.Y. and K.L.N. provided insights and developed key reagents and protocols for protein detection. J.B.W. provided insights and discussions for experimental planning. R.S., S.S., L.S.L., V.G.S., A.R. and P.S. each supervised various aspects of the work. E.P.M., C.A.L., K.Y.C., L.S.L. and P.S. drafted the manuscript with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
C.A.L., L.S.L., V.G.S. and A.R. are listed as co-inventors on a patent related to mtscATAC-seq (U.S. provisional patent application 62/683,502). In the past 3 years, R.S. has worked as a consultant for Bristol-Myers Squibb, Regeneron and Kallyope and served as a scientific advisory board member for Immunai and Resolve BioSciences. A.R. is a founder and equity holder of Celsius Therapeutics, an equity holder in Immunitas Therapeutics and, until August 31, 2020, was a scientific advisory board member of Syros Pharmaceuticals, Neogene Therapeutics, Asimov and Thermo Fisher Scientific. From August 1, 2020, A.R. is an employee of Genentech. P.S. is listed as co-inventor on a patent related to this work (U.S. provisional patent application 62/515-180).
Additional information
Peer review information Nature Biotechnology thanks Dan Xie, Golnaz Vahedi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Additional technical and computational validation of ASAP-seq workflows.
a. PBMCs and compensation beads were stained with fluorophore-conjugated antibodies and subjected to the ASAP-seq workflow with samples withdrawn at the indicated steps and assessed for fluorophore intensity by flow cytometry. CD19 (staining B cells), CD11c (dendritic cells) and CD4 (lymphocytes and monocytes) signal on fixed cells is hardly affected by permeabilization alone, but after the 37 °C incubation for 1 h to mimic the Tn5 transposition reaction, some signal reduction is observed. b. Barcoding scheme of TSA tags using the bridge oligo for TotalSeqTM-A (BOA). TSA tags do not contain UMIs, so to allow molecule counting, UBIs (N9V) are incorporated via the bridge oligo. c. Species mixing experiment as in Fig. 1c, using the Post-SPRI approach for tag recovery. Points are colored based on species classification using ATAC fragments. d. ATAC library complexity and TSS enrichment for fragments from each species under the two protein-tag library approaches. e. Comparison of protein tag complexity between libraries prepared using the pre- and post-SPRI approach. f. Comparison of ATAC library complexity between mtscATAC-seq and ASAP-seq. Boxplots: center line, median; box limits, first and third quartiles; whiskers, 1.5× interquartile range. g. Two-dimensional embedding of the PBMC hashing data using t-SNE. The four major clusters (black) correspond to the four hashing antibodies used to stain the PBMCs. 13,772 cells were recovered and 1,396 doublets (red) were detected. h. UMAP embedding resolving PBMC cell types based on chromatin accessibility for cells processed by mtscATAC-seq and ASAP-seq. Data for the two different samples were processed together using cell ranger-atac aggr before dimensionality reduction. i,j. Selected protein markers (i) and corresponding gene score activities (j) superimposed on the ATAC-clustered PBMCs (for the ASAP-seq sample) as in (h).
Extended Data Fig. 2 Additional validation and comparison of modular ASAP-seq workflows.
a. Barcoding scheme of TSB tags using the bridge oligo for TotalSeqB (BOB). TSB tags contain UMIs (encompassing the antibody barcode), negating the requirement for a UBI on the bridge oligo. b. Pairwise comparison of centered log-ratio (CLR) normalized TSA and TSB counts under OMNI lysis conditions (n = 5,236 cells). Counts were collapsed for unique molecules using UBIs (TSA panel) or UMIs (TSB panel). c. Comparison of CLR normalised TSB counts under the two lysis conditions. Statistical comparisons are two-sided Wilcoxon rank sum test with Bonferroni adjusted p-values (ns = not significant; * padj = 0.0002; ** padj = 2.2×10–16). d. UMAP embedding and cluster annotation of the LLL (n = 5,236) and OMNI (n = 4,748) processed cells. Data for the two different samples were processed together using cell ranger-atac aggr before dimensionality reduction. e. TSA and TSB CLR counts projected on the LLL embeddings. f. TSA and TSB CLR counts projected on the OMNI embeddings.
Extended Data Fig. 3 Supporting information for ASAP-seq bone marrow analyses.
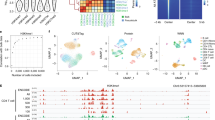
a. Annotation of reduced dimension space with the Doublet Enrichment score from ArchR. Arrow indicates the monocytic progenitor population. b. Histogram of scores from panel (a). c. Feature plots for six additional antibody tags in the reduced dimension space. d. Correlation heatmap between 25 most variable TF activities and surface markers. e. Percent of cells in each ArchR cluster (y axis) mapping to the indicated Seurat cluster (x axis) after label transfer using the protein tags only f. Substitution rate (observed over expected) of mgatk-identified heteroplasmic mutations (y axis) in each class of mononucleotide and trinucleotide change resolved by the heavy (H) and light (L) strands of the mitochondrial genome. g. Projection of 13711 G > A in single cells; threshold for + was 5% heteroplasmy. h. Distribution of observed mtDNA mutations in cells among major cell lineages. i. Association of antibody tag abundance to cell clones determined by mtDNA genotypes, highlighting the erythroid marker CD71. j. Developmental trajectory of erythroid differentiation using semi-supervised pseudotime analysis. k. Expression of select cell surface markers along the erythroid developmental trajectory highlighted in (j). Rows are min-max normalized. l. Expression of chromatin activity scores along the monocytic developmental trajectory for genes encoding proteins shown in Fig. 3h. m. Expression of chromatin activity scores along the erythroid developmental trajectory for genes encoding proteins shown in (k).
Extended Data Fig. 4 Supporting information for combined ASAP-seq and CITE-seq readouts.
a. Antibody tag complexity per condition and technology. Median tag complexity is 1.7-2x higher in CITE-seq compared to ASAP-seq and 1.3-1.6x higher in stimulation compared to control sample. Boxplots: center line, median; box limits, first and third quartiles; whiskers, 1.5× interquartile range. The lower panels show the per-cluster mean tag abundance for the 50 most variable antibodies and corresponding Pearson correlations. b,c. Cellular distribution of protein tags measured by ASAP-seq (left) and CITE-seq (right) for control (top) and stimulated conditions (bottom) for, (b) CD278 (ICOS) and (c) CD71 (TFRC). d. Protein tag measurement importance in predicting cell cluster and stimulation from two different Random Forest models. Negative controls (rat epitopes) are shown in red. e-g. ASAP-seq and CITE-seq data co-embedding utilizing protein abundances. Cells are highlighted by (e) chromatin/RNA cluster identity, (f) stimulation condition and (g) technology assayed. h-j. UMAPs of chromatin accessibility, mRNA expression, and surface protein levels for (h) CD28, (i) CD4, and (j) CD52. k. Summary of changes in chromatin accessibility, gene expression and surface protein abundance for 103 expressed genes in B cells following T cell stimulation. l,m. UMAPs of chromatin accessibility, mRNA expression, and surface protein levels for genes with differential expression in B cells, including (l) CD184 (CXCR4) and (m) CD25 (IL2RA).
Extended Data Fig. 5 Supporting information for DOGMA-seq.
a-e. QC metrics of indicated modalities captured by DOGMA-seq applied on the stimulated PBMC sample. (a) TSS score, (b) ATAC fragment complexity, (c) % mtDNA content, (d) number of genes/cell and (e) protein tag complexity in the two different cell preparations compare similarly to the control PBMC sample in Fig. 5b-f. f. Percent of UMIs detected in the GEX library that map to mtRNA is higher in the digitonin-treated cells. g-h. Percent of UMIs mapping to exons is higher in the digitonin-treated (DIG) compared to LLL-treated cells (g), but similar when mitochondrial transcripts are excluded (h). i. CD138 tag counts projected on the three modality WNN stimulation clusters. j. Gene activity scores, transcript and protein tag counts projected for the indicated markers on the control and stimulated 3WNN clusters. k. Heatmaps showing percent overlap between clusters detected by 3WNN compared to 2WNN variations applied on the control PBMC dataset. l. Mean coverage along the mtDNA genome in control and stimulated PBMCs. m. Substitution rate (observed over expected) of mgatk-identified heteroplasmic mutations (y axis) in each class of mononucleotide and trinucleotide change resolved by the heavy (H) and light (L) strands of the mitochondrial genome for all cells in the PBMC-LLL condition. n. Observed (red) and permuted (gray) log2 heteroplasmy changes across the 106 identified variants. Statistical test: two-sided Kolmogorov–Smirnov Test. o. 3WNN UMAP embedding of control and stimulated PBMC samples under LLL and DIG processing. Dashed box indicates activated T cell clusters. p. Comparison of peak to gene linkage for genes detected in both protein and RNA modalities. Each dot is a peak to gene link with the z score representing the magnitude of the association. Boxplots: center line, median; box limits, first and third quartiles; whiskers, 1.5× interquartile range.
Extended Data Fig. 6 Supporting information for ASAP-seq based decoding of perturbations in primary T cells.
a. Schematic for CRISPR perturbation experiment in primary human T cells. CD4 + T cells from healthy donors were stimulated for 72 hours, followed by a resting period of four days to enable expansion. On Day 7, cells were electroporated with Cas9 RNPs and then rested for an additional 8 days before secondary stimulation. b. Heatmap of cell demultiplexing with hashing antibodies, indicating normalized abundance of each hashtag. c. Assessment of the effect of CRISPR perturbations on three indicated protein surface markers. d. UMAP embedding overlaid with expression of the eight indicated surface protein markers. e. Allele-specific CRISPR editing outcomes for ZAP70 gRNA1 (left) and ZAP70 gRNA2 (right). The wildtype allele is indicated by **. f. Volcano plots showing TF motifs with significantly changed chromatin accessibility profiles between NTC cells and the indicated gRNAs (FDR < = 0.05, chromVAR accessibility change > = 0.25). g. Correlation of chromVAR median accessibility changes or FDR (bottom right panel) between the indicated gRNAs. h. Genomic tracks of TNFRSF18 and HAVCR2 loci with corresponding CLR-normalized protein abundance ridge plots. CLR-normalized protein abundance from the PBMC stimulation experiment is indicated by the corresponding boxplots. Differentially accessible regions are highlighted in blue.
Extended Data Fig. 7 Supporting information for intracellular ASAP-seq workflow.
a,b. Selected protein markers (a) and corresponding gene activity scores (b) superimposed on the ATAC-clustered PBMCs from the intracellular staining experiment (see Fig. 3a). c. Heatmap of cell demultiplexing with hashing antibodies, indicating normalized abundance of each hashtag for 24 different perturbation conditions. d. Violin plots showing distribution of CLR normalized protein counts for indicated proteins and their associated gRNA. e. Genomic tracks of IFNG and GZMB loci, indicating pseudo-bulk ATAC signal tracks across six Louvain clusters with corresponding log-normalized gene activity score violin plots shown to the right. Differentially accessible regions are highlighted in blue.
Supplementary information
Supplementary Information
ASAP-seq and DOGMA-seq step-by-step protocols.
Supplementary Tables 1–5
Lists of antibodies, oligos, cluster annotations and stimulation-induced modality changes.
Rights and permissions
About this article
Cite this article
Mimitou, E.P., Lareau, C.A., Chen, K.Y. et al. Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nat Biotechnol 39, 1246–1258 (2021). https://doi.org/10.1038/s41587-021-00927-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-00927-2
This article is cited by
-
Single Cell Atlas: a single-cell multi-omics human cell encyclopedia
Genome Biology (2024)
-
DANCE: a deep learning library and benchmark platform for single-cell analysis
Genome Biology (2024)
-
Research progress and application of single-cell sequencing in head and neck malignant tumors
Cancer Gene Therapy (2024)
-
Simultaneous single-cell analysis of 5mC and 5hmC with SIMPLE-seq
Nature Biotechnology (2024)
-
Deep generative modeling of transcriptional dynamics for RNA velocity analysis in single cells
Nature Methods (2024)