Abstract
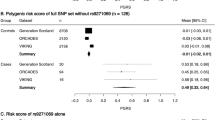
A previous study using cumulative genetic risk estimations in multiple sclerosis (MS) successfully tracked the aggregation of susceptibility variants in multi-case and single-case families. It used a limited description of susceptibility loci available at the time (17 loci). Even though the full roster of MS risk genes remains unavailable, we estimated the genetic burden in MS families and assess its disease predictive power using up to 64 single-nucleotide polymorphism (SNP) markers according to the most recent literature. A total of 708 controls, 3251 MS patients and their relatives, as well as 117 twin pairs were genotyped. We validated the increased aggregation of genetic burden in multi-case compared with single-case families (P=4.14e−03) and confirm that these data offer little opportunity to accurately predict MS, even within sibships (area under receiver operating characteristic (AUROC)=0.59 (0.55, 0.53)). Our results also suggest that the primary progressive and relapsing-type forms of MS share a common genetic architecture (P=0.368; difference being limited to that corresponding to ±2 typical MS-associated SNPs). We have confirmed the properties of individual genetic risk score in MS. Comparing with previous reference point for MS genetics (17 SNPs), we underlined the corrective consequences of the integration of the new findings from GWAS and meta-analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Oksenberg JR, Baranzini SE, Sawcer S, Hauser SL . The genetics of multiple sclerosis: SNPs to pathways to pathogenesis. Nat Rev Genet 2008; 9: 516–526.
Bush WS, Sawcer SJ, de Jager PL, Oksenberg JR, McCauley JL, Pericak-Vance MA et al. Evidence for polygenic susceptibility to multiple sclerosis—the shape of things to come. Am J Hum Genet 2010; 86: 621–625.
Gourraud PA, Harbo HF, Hauser SL, Baranzini SE . The genetics of multiple sclerosis: an up-to-date review. Immunol Rev 2012; 248: 87–103.
Sawcer S, Hellenthal G, Pirinen M, Spencer CC, Patsopoulos NA, Moutsianas L et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 2011; 476: 214–219.
Burton PR, Clayton DG, Cardon LR, Craddock N, Deloukas P, Duncanson A et al. Association scan of 14,500 nonsynonymous SNPs in four diseases identifies autoimmunity variants. Nat Genet 2007; 39: 1329–1337.
Hafler DA, Compston A, Sawcer S, Lander ES, Daly MJ, De Jager PL et al. Risk alleles for multiple sclerosis identified by a genomewide study. N Engl J Med 2007; 357: 851–862.
Comabella M, Craig DW, Camina-Tato M, Morcillo C, Lopez C, Navarro A et al. Identification of a novel risk locus for multiple sclerosis at 13q31.3 by a pooled genome-wide scan of 500,000 single nucleotide polymorphisms. PLoS One 2008; 3: e3490.
ANZGENE-consortium. Genome-wide association study identifies new multiple sclerosis susceptibility loci on chromosomes 12 and 20. Nat Genet 2009; 41: 824–828.
Baranzini SE, Wang J, Gibson RA, Galwey N, Naegelin Y, Barkhof F et al. Genome-wide association analysis of susceptibility and clinical phenotype in multiple sclerosis. Hum Mol Genet 2009; 18: 767–778.
De Jager PL, Jia X, Wang J, de Bakker PI, Ottoboni L, Aggarwal NT et al. Meta-analysis of genome scans and replication identify CD6, IRF8 and TNFRSF1A as new multiple sclerosis susceptibility loci. Nat Genet 2009; 41: 776–782.
Jakkula E, Leppa V, Sulonen AM, Varilo T, Kallio S, Kemppinen A et al. Genome-wide association study in a high-risk isolate for multiple sclerosis reveals associated variants in STAT3 gene. Am J Hum Genet 2010; 86: 285–291.
Nischwitz S, Cepok S, Kroner A, Wolf C, Knop M, Muller-Sarnowski F et al. Evidence for VAV2 and ZNF433 as susceptibility genes for multiple sclerosis. J Neuroimmunol 2010; 227: 162–166.
Sanna S, Pitzalis M, Zoledziewska M, Zara I, Sidore C, Murru R et al. Variants within the immunoregulatory CBLB gene are associated with multiple sclerosis. Nat Genet 2010; 42: 495–497.
Baranzini SE, Galwey NW, Wang J, Khankhanian P, Lindberg R, Pelletier D et al. Pathway and network-based analysis of genome-wide association studies in multiple sclerosis. Hum Mol Genet 2009; 18: 2078–2090.
Cotsapas C, Voight BF, Rossin E, Lage K, Neale BM, Wallace C et al. Pervasive sharing of genetic effects in autoimmune disease. PLoS Genet 2011; 7: e1002254.
Rioux JD, Goyette P, Vyse TJ, Hammarstrom L, Fernando MM, Green T et al. Mapping of multiple susceptibility variants within the MHC region for 7 immune-mediated diseases. Proc Natl Acad Sci USA 2009; 106: 18680–18685.
Couturier N, Bucciarelli F, Nurtdinov RN, Debouverie M, Lebrun-Frenay C, Defer G et al. Tyrosine kinase 2 variant influences T lymphocyte polarization and multiple sclerosis susceptibility. Brain 2011; 134 (Pt 3): 693–703.
De Jager PL, Baecher-Allan C, Maier LM, Arthur AT, Ottoboni L, Barcellos L et al. The role of the CD58 locus in multiple sclerosis. Proc Natl Acad Sci USA 2009; 106: 5264–5269.
Gregory AP, Dendrou CA, Attfield KE, Haghikia A, Xifara DK, Butter F et al. TNF receptor 1 genetic risk mirrors outcome of anti-TNF therapy in multiple sclerosis. Nature 2012; 488: 508–511.
Gregory SG, Schmidt S, Seth P, Oksenberg JR, Hart J, Prokop A et al. Interleukin 7 receptor alpha chain (IL7R) shows allelic and functional association with multiple sclerosis. Nat Genet 2007; 39: 1083–1091.
Maier LM, Lowe CE, Cooper J, Downes K, Anderson DE, Severson C et al. IL2RA genetic heterogeneity in multiple sclerosis and type 1 diabetes susceptibility and soluble interleukin-2 receptor production. PLoS Genet 2009; 5: e1000322.
De Jager PL, Chibnik LB, Cui J, Reischl J, Lehr S, Simon KC et al. Integration of genetic risk factors into a clinical algorithm for multiple sclerosis susceptibility: a weighted genetic risk score. Lancet Neurol 2009; 8: 1111–1119.
Gourraud PA, McElroy JP, Caillier SJ, Johnson BA, Santaniello A, Hauser SL et al. Aggregation of multiple sclerosis genetic risk variants in multiple and single case families. Ann Neurol 2011; 69: 65–74.
Sawcer S, Ban M, Wason J, Dudbridge F . What role for genetics in the prediction of multiple sclerosis? Ann Neurol 2010; 67: 3–10.
Patsopoulos NA, Esposito F, Reischl J, Lehr S, Bauer D, Heubach J et al. Genome-wide meta-analysis identifies novel multiple sclerosis susceptibility loci. Ann Neurol 2011; 70: 897–912.
Lango Allen H, Estrada K, Lettre G, Berndt SI, Weedon MN, Rivadeneira F et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 2010; 467: 832–838.
Barcellos LF, Oksenberg JR, Green AJ, Bucher P, Rimmler JB, Schmidt S et al. Genetic basis for clinical expression in multiple sclerosis. Brain 2002; 125 (Pt 1): 150–158.
Goodkin DE, Doolittle TH, Hauser SS, Ransohoff RM, Roses AD, Rudick RA . Diagnostic criteria for multiple sclerosis research involving multiply affected families. Arch Neurol 1991; 48: 805–807.
Johnson AD, Handsaker RE, Pulit SL, Nizzari MM, O'Donnell CJ, de Bakker PI . SNAP: a web-based tool for identification and annotation of proxy SNPs using HapMap. Bioinformatics 2008; 24: 2938–2939.
Acknowledgements
This study was supported by the National Institutes of Health (grant RO1NS26799 RO1NS19142), the National Multiple Sclerosis Society (grant RG2901) and the Cambridge NIHR Biomedical Research Centre, the Institut National de la Santé et de la Recherche Médicale (INSERM), the Fondation d’Aide pour la Recherche sur la Sclérose En Plaques (ARSEP), the Association Française contre les Myopathies (AFM) and GIS-IBISA. The research leading to these results has received funding from the program ‘Investissements d’avenir’ ANR-10-IAIHU-06. We acknowledge use of the cohort of the REFGENSEP and thank ICM, CIC Pitié-Salpêtrière, Généthon and REFGENSEP’s members for their help and support. NI is a postdoctoral fellow supported by Japan Society for Promotion of Science. VD received a travel grant from ARSEP. P-AG is a recipient of the Nancy Davis Foundation young investigator award. We acknowledge the assistance of Stacy Caillier, Adam Santaniello, Vinod Bakthavachalam Jinyang Wang and Matthew Chan in sample and data management.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Genes and Immunity website
Rights and permissions
About this article
Cite this article
Isobe, N., Damotte, V., Lo Re, V. et al. Genetic burden in multiple sclerosis families. Genes Immun 14, 434–440 (2013). https://doi.org/10.1038/gene.2013.37
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2013.37
Keywords
This article is cited by
-
Genetic susceptibility to multiple sclerosis: interactions between conserved extended haplotypes of the MHC and other susceptibility regions
BMC Medical Genomics (2021)
-
Aberrant STAT phosphorylation signaling in peripheral blood mononuclear cells from multiple sclerosis patients
Journal of Neuroinflammation (2018)