Abstract
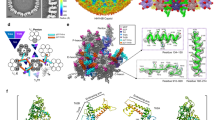
Retroviruses are the aetiological agents of a range of human diseases including AIDS and T-cell leukaemias. They follow complex life cycles, which are still only partly understood at the molecular level. Maturation of newly formed retroviral particles is an essential step in production of infectious virions, and requires proteolytic cleavage of Gag polyproteins in the immature particle to form the matrix, capsid and nucleocapsid proteins present in the mature virion1. Capsid proteins associate to form a dense viral core2 that may be spherical, cylindrical or conical depending on the genus of the virus3. Nonetheless, these assemblies all appear to be composed of a lattice formed from hexagonal rings, each containing six capsid monomers2,4,5. Here, we describe the X-ray structure of an individual hexagonal assembly from N-tropic murine leukaemia virus (N-MLV). The interface between capsid monomers is generally polar, consistent with weak interactions within the hexamer. Similar architectures are probably crucial for the regulation of capsid assembly and disassembly in all retroviruses. Together, these observations provide new insights into retroviral uncoating and how cellular restriction factors may interfere with viral replication.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Swanstrom, R. & Wills, J. W. in Synthesis, Assembly and Processing of Viral Proteins (eds Coffin, J. M., Hughes, S. H. & Varmus, H. E.) (Cold Spring Harbor Laboratory Press, New York, 1997)
Briggs, J. A., Wilk, T., Welker, R., Krausslich, H. G. & Fuller, S. D. Structural organization of authentic, mature HIV-1 virions and cores. EMBO J. 22, 1707–1715 (2003)
Nermut, M. V. & Hockley, D. J. Comparative morphology and structural classification of retroviruses. Curr. Top. Microbiol. Immunol. 214, 1–24 (1996)
Li, S., Hill, C. P., Sundquist, W. I. & Finch, J. T. Image reconstructions of helical assemblies of the HIV-1 CA protein. Nature 407, 409–413 (2000)
Ganser, B. K., Cheng, A., Sundquist, W. I. & Yeager, M. Three-dimensional structure of the M-MuLV CA protein on a lipid monolayer: a general model for retroviral capsid assembly. EMBO J. 22, 2886–2892 (2003)
Kingston, R. L. et al. Structure and self-association of the Rous sarcoma virus capsid protein. Struct. Fold. Des. 8, 617–628 (2000)
Cornilescu, C. C., Bouamr, F., Yao, X., Carter, C. & Tjandra, N. Structural analysis of the N-terminal domain of the human T-cell leukemia virus capsid protein. J. Mol. Biol. 306, 783–797 (2001)
Gamble, T. R. et al. Crystal structure of human cyclophilin A bound to the amino-terminal domain of HIV-1 capsid. Cell 87, 1285–1294 (1996)
Gitti, R. K. et al. Structure of the amino-terminal core domain of the HIV-1 capsid protein. Science 273, 231–235 (1996)
Jin, Z., Jin, L., Peterson, D. L. & Lawson, C. L. Model for lentivirus capsid core assembly based on crystal dimers of EIAV p26. J. Mol. Biol. 286, 83–93 (1999)
Khorasanizadeh, S., Campos-Olivas, R. & Summers, M. F. Solution structure of the capsid protein from the human T-cell leukemia virus type-I. J. Mol. Biol. 291, 491–505 (1999)
Nandhagopal, N. et al. Dimeric Rous sarcoma virus capsid protein structure relevant to immature Gag assembly. J. Mol. Biol. 335, 275–282 (2004)
Tang, C., Ndassa, Y. & Summers, M. F. Structure of the N-terminal 283-residue fragment of the immature HIV-1 Gag polyprotein. Nature Struct. Biol. 9, 537–543 (2002)
Gross, I., Hohenberg, H., Huckhagel, C. & Krausslich, H. G. N-Terminal extension of human immunodeficiency virus capsid protein converts the in vitro assembly phenotype from tubular to spherical particles. J. Virol. 72, 4798–4810 (1998)
von Schwedler, U. K. et al. Proteolytic refolding of the HIV-1 capsid protein amino-terminus facilitates viral core assembly. EMBO J. 17, 1555–1568 (1998); corrigendum 19, 2391 (2000)
Briggs, J. A. et al. The stoichiometry of Gag protein in HIV-1. Nature Struct. Mol. Biol. 11, 672–675 (2004)
Lanman, J. et al. Identification of novel interactions in HIV-1 capsid protein assembly by high-resolution mass spectrometry. J. Mol. Biol. 325, 759–772 (2003)
Lanman, J. et al. Key interactions in HIV-1 maturation identified by hydrogen-deuterium exchange. Nature Struct. Mol. Biol. 11, 676–677 (2004)
von Schwedler, U. K., Stray, K. M., Garrus, J. E. & Sundquist, W. I. Functional surfaces of the human immunodeficiency virus type 1 capsid protein. J. Virol. 77, 5439–5450 (2003)
Forshey, B. M., von Schwedler, U., Sundquist, W. I. & Aiken, C. Formation of a human immunodeficiency virus type 1 core of optimal stability is crucial for viral replication. J. Virol. 76, 5667–5677 (2002)
Ganser-Pornillos, B. K., von Schwedler, U. K., Stray, K. M., Aiken, C. & Sundquist, W. I. Assembly properties of the human immunodeficiency virus type 1 CA protein. J. Virol. 78, 2545–2552 (2004)
Fassati, A. & Goff, S. P. Characterization of intracellular reverse transcription complexes of Moloney murine leukemia virus. J. Virol. 73, 8919–8925 (1999)
Fassati, A. & Goff, S. P. Characterization of intracellular reverse transcription complexes of human immunodeficiency virus type 1. J. Virol. 75, 3626–3635 (2001)
Karageorgos, L., Li, P. & Burrell, C. Characterization of HIV replication complexes early after cell-to-cell infection. AIDS Res. Hum. Retroviruses 9, 817–823 (1993)
Best, S., Le Tissier, P., Towers, G. & Stoye, J. P. Positional cloning of the mouse retrovirus restriction gene Fv1. Nature 382, 826–829 (1996)
Boone, L. R. et al. Reversal of Fv-1 host range by in vitro restriction endonuclease fragment exchange between molecular clones of N-tropic and B-tropic murine leukemia virus genomes. J. Virol. 48, 110–119 (1983)
Stremlau, M. et al. The cytoplasmic body component TRIM5α restricts HIV-1 infection in Old World monkeys. Nature 427, 848–853 (2004)
Terwilliger, T. C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999)
Cowtan, K. ‘dm’: an automated procedure for phase improvement by density modification. Joint CCP4 ESF-EACBM Newsl. Prot. Crystallogr. 31, 34–38 (1994)
Morris, R. J., Perrakis, A. & Lamzin, V. S. ARP/wARP and automatic interpretation of protein electron density maps. Methods Enzymol. 374, 229–244 (2003)
Acknowledgements
We thank S. Gamblin for assistance in crystal handling.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Mortuza, G., Haire, L., Stevens, A. et al. High-resolution structure of a retroviral capsid hexameric amino-terminal domain. Nature 431, 481–485 (2004). https://doi.org/10.1038/nature02915
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02915
This article is cited by
-
Structure of a Ty1 restriction factor reveals the molecular basis of transposition copy number control
Nature Communications (2021)
-
Origins and evolutionary consequences of ancient endogenous retroviruses
Nature Reviews Microbiology (2019)
-
An RNA-binding compound that stabilizes the HIV-1 gRNA packaging signal structure and specifically blocks HIV-1 RNA encapsidation
Retrovirology (2018)
-
A comparative analysis of the foamy and ortho virus capsid structures reveals an ancient domain duplication
BMC Structural Biology (2018)
-
In vitro assembly of the Rous Sarcoma Virus capsid protein into hexamer tubes at physiological temperature
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.