Abstract
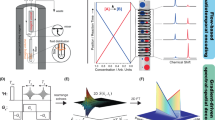
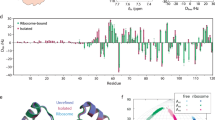
Macromolecular complex formation is governed by two opposing constraints of specificity and speed1,2. Kinetic3,4,5,6 and theoretical considerations suggest that significant rate enhancement can be achieved either by reducing the dimensionality of the search process1,7 or by the creation of a short-range attractive potential around the target site2. This implies the existence of transient intermediates involving non-specific binding modes. Here we show that intermolecular paramagnetic relaxation enhancement (PRE) provides a means of directly detecting the presence of, and investigating the nature of, low population transient intermediates under equilibrium conditions. Applying this approach, we characterize the search process whereby a sequence-specific transcription factor (the homeodomain of HOXD9) binds to non-cognate DNA sites as a means of enhancing the rate of specific association. The PRE data in the fast exchange regime reveal the presence of transient intermediates formed in a stochastic manner at non-cognate sites whose structure is similar to that of the specific complex. Two distinct search processes involving intra- as well as intermolecular translocations can be delineated. The intermolecular PRE method is general and can be readily applied to investigations of transient intermediates in many other macromolecular binding processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Berg, O. G. & von Hippel, P. H. Diffusion-controlled macromolecular interactions. Annu. Rev. Biophys. Biophys. Chem. 14, 131–160 (1985)
Zhou, H.-X. & Szabo, A. Enhancement of association rates by nonspecific binding to DNA and cell membranes. Phys. Rev. Lett. 93, 178101 (2004)
von Hippel, P. H. & Berg, O. G. Facilitated target location in biological systems. J. Biol. Chem. 264, 675–678 (1989)
Halford, S. E. & Marko, J. F. How do site-specific DNA-binding proteins find their targets? Nucleic Acids Res. 32, 3040–3052 (2004)
Schreiber, G. & Fersht, A. R. Rapid, electrostatically assisted association of proteins. Nature Struct. Biol. 3, 427–431 (1996)
Vijayakumar, M. et al. Electrostatic enhancement of diffusion-controlled protein-protein association: comparison of theory and experiment on barnase and barstar. J. Mol. Biol. 278, 1015–1024 (1998)
Adam, G. & Delbruck, M. in Structural Chemistry and Molecular Biology (eds Rich, A. & Davidson, N.) 198–215 (Freeman & Co., San Francisco, 1968)
Solomon, I. Relaxation processes in a system of two spins. Phys. Rev. 99, 559–565 (1955)
Bloembergen, N. & Morgan, L. O. Proton relaxation times in paramagnetic solutions. Effects of election spin relaxation. J. Chem. Phys. 34, 842–850 (1961)
Iwahara, J., Schwieters, C. D. & Clore, G. M. Ensemble approach for NMR structure refinement against 1H paramagnetic relaxation enhancement data arising from a flexible paramagnetic group attached to a macromolecule. J. Am. Chem. Soc. 126, 5879–5896 (2004)
Iwahara, J., Schwieters, C. D. & Clore, G. M. Characterization of nonspecific protein-DNA interactions by 1H paramagnetic relaxation enhancement. J. Am. Chem. Soc. 126, 12800–12808 (2004)
Iwahara, J., Anderson, D. E., Murphy, E. C. & Clore, G. M. EDTA-derivatized deoxythymidine as a tool for rapid determination of protein binding polarity to DNA by intermolecular paramagnetic relaxation enhancement. J. Am. Chem. Soc. 125, 6634–6635 (2003)
Gehring, W. J. et al. Homeodomain-DNA recognition. Cell 78, 211–223 (1994)
Billeter, M. et al. Determination of the nuclear magnetic resonance solution structure of an Antennapedia homeodomain-DNA complex. J. Mol. Biol. 234, 1084–1093 (1993)
Fraenkel, E. & Pabo, C. O. Comparison of X-ray and NMR structures for the Antennapedia homeodomain-DNA complex. Nature Struct. Biol. 5, 692–697 (1998)
Clore, G. M. & Garrett, D. S. R-factor, free R and complete cross-validation for dipolar coupling refinement of NMR structures. J. Am. Chem. Soc. 121, 9008–9012 (1999)
Iwahara, J. & Clore, G. M. Direct observation of enhanced translocation of a homeodomain between DNA cognate sites by NMR exchange spectroscopy. J. Am. Chem. Soc. 128, 404–405 (2006)
Affolter, M., Percival-Smith, A., Müller, M., Leupin, W. & Gehring, W. J. DNA binding properties of the purified Antennapedia homeodomain. Proc. Natl Acad. Sci. USA 87, 4093–4097 (1990)
Catron, K. M., Iler, N. & Abate, C. Nucleotides flanking a conserved TAAT core dictate the DNA binding specificity of three murine homeodomain proteins. Mol. Cell. Biol. 13, 2354–2365 (1993)
Misteli, T. Protein dynamics: implications for nuclear architecture and gene expression. Science 291, 843–847 (2001)
Lewin, B. Genes VII (Oxford Univ. Press, Oxford, 2000)
Gillespie, J. R. & Shortle, D. Characterization of long-range structure in the denatured state of staphylococcal nuclease. I. Paramagnetic relaxation enhancement by nitroxide spin labels. J. Mol. Biol. 268, 158–169 (1997)
Dvoretsky, A., Gaponenko, V. & Rosevear, P. R. Derivation of structural restraints using a thiol-reactive chelator. FEBS Lett. 528, 189–192 (2002)
Gabdoulline, R. R. & Wade, R. C. Biomolecular diffusional association. Curr. Opin. Struct. Biol. 12, 204–213 (2002)
Donaldson, L. W. et al. Structural characterization of proteins with an attached ATCUN motif by paramagnetic relaxation enhancement NMR spectroscopy. J. Am. Chem. Soc. 123, 9843–9847 (2001)
Schwieters, C. D., Kuszewski, J., Tjandra, N. & Clore, G. M. The Xplor-NIH NMR molecular structure determination package. J. Magn. Reson. 160, 65–73 (2003)
McConnell, H. M. Reaction rates by nuclear magnetic resonance. J. Chem. Phys. 28, 430–431 (1958)
Acknowledgements
This work was supported by the Intramural Program of the NIH, the NIDDK, and in part by the AIDS Targeted Antiviral Program of the Office of the Director of the NIH (G.M.C.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains details of lineshape simulations, residual dipolar coupling data, NMR exchange experiments and PRE profiles to be linked to the online version of the paper. (PDF 1671 kb)
Rights and permissions
About this article
Cite this article
Iwahara, J., Clore, G. Detecting transient intermediates in macromolecular binding by paramagnetic NMR. Nature 440, 1227–1230 (2006). https://doi.org/10.1038/nature04673
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04673
This article is cited by
-
Does PCNA diffusion on DNA follow a rotation-coupled translation mechanism?
Nature Communications (2020)
-
Racemic phosphorothioate as a tool for NMR investigations of protein-DNA complexes
Journal of Biomolecular NMR (2020)
-
Paramagnetic NMR in drug discovery
Journal of Biomolecular NMR (2020)
-
Role of Macromolecular Crowding on the Intracellular Diffusion of DNA Binding Proteins
Scientific Reports (2018)
-
Protein docking refinement by convex underestimation in the low-dimensional subspace of encounter complexes
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.