Abstract
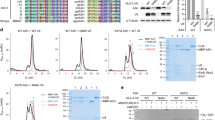
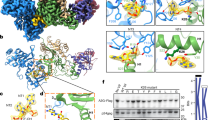
The human APOBEC3G (apolipoprotein B messenger-RNA-editing enzyme, catalytic polypeptide-like 3G) protein is a single-strand DNA deaminase that inhibits the replication of human immunodeficiency virus-1 (HIV-1), other retroviruses and retrotransposons1,2,3,4,5,6. APOBEC3G anti-viral activity is circumvented by most retroelements, such as through degradation by HIV-1 Vif7. APOBEC3G is a member of a family of polynucleotide cytosine deaminases, several of which also target distinct physiological substrates. For instance, APOBEC1 edits APOB mRNA and AID deaminates antibody gene DNA8,9,10. Although structures of other family members exist, none of these proteins has elicited polynucleotide cytosine deaminase or anti-viral activity11,12,13,14,15,16. Here we report a solution structure of the human APOBEC3G catalytic domain. Five α-helices, including two that form the zinc-coordinating active site, are arranged over a hydrophobic platform consisting of five β-strands. NMR DNA titration experiments, computational modelling, phylogenetic conservation and Escherichia coli-based activity assays combine to suggest a DNA-binding model in which a brim of positively charged residues positions the target cytosine for catalysis. The structure of the APOBEC3G catalytic domain will help us to understand functions of other family members and interactions that occur with pathogenic proteins such as HIV-1 Vif.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chelico, L., Pham, P., Calabrese, P. & Goodman, M. F. APOBEC3G DNA deaminase acts processively 3' → 5' on single-stranded DNA. Nat. Struct. Mol. Biol. 13, 392–399 (2006)
Esnault, C. et al. APOBEC3G cytidine deaminase inhibits retrotransposition of endogenous retroviruses. Nature 433, 430–433 (2005)
Harris, R. S. et al. DNA deamination mediates innate immunity to retroviral infection. Cell 113, 803–809 (2003)
Mangeat, B. et al. Broad antiretroviral defence by human APOBEC3G through lethal editing of nascent reverse transcripts. Nature 424, 99–103 (2003)
Sheehy, A. M., Gaddis, N. C., Choi, J. D. & Malim, M. H. Isolation of a human gene that inhibits HIV-1 infection and is suppressed by the viral Vif protein. Nature 418, 646–650 (2002)
Zhang, H. et al. The cytidine deaminase CEM15 induces hypermutation in newly synthesized HIV-1 DNA. Nature 424, 94–98 (2003)
Yu, X. et al. Induction of APOBEC3G ubiquitination and degradation by an HIV-1 Vif–Cul5–SCF complex. Science 302, 1056–1060 (2003)
Conticello, S. G., Langlois, M. A., Yang, Z. & Neuberger, M. S. DNA deamination in immunity: AID in the context of its APOBEC relatives. Adv. Immunol. 94, 37–73 (2007)
Di Noia, J. M. & Neuberger, M. S. Molecular mechanisms of antibody somatic hypermutation. Annu. Rev. Biochem. 76, 1–22 (2007)
Wedekind, J. E., Dance, G. S., Sowden, M. P. & Smith, H. C. Messenger RNA editing in mammals: new members of the APOBEC family seeking roles in the family business. Trends Genet. 19, 207–216 (2003)
Betts, L., Xiang, S., Short, S. A., Wolfenden, R. & Carter, C. W. Cytidine deaminase. The 2.3 Å crystal structure of an enzyme: transition-state analog complex. J. Mol. Biol. 235, 635–656 (1994)
Harris, R. S., Petersen-Mahrt, S. K. & Neuberger, M. S. RNA editing enzyme APOBEC1 and some of its homologs can act as DNA mutators. Mol. Cell 10, 1247–1253 (2002)
Johansson, E., Mejlhede, N., Neuhard, J. & Larsen, S. Crystal structure of the tetrameric cytidine deaminase from Bacillus subtilis at 2.0 Å resolution. Biochemistry 41, 2563–2570 (2002)
Ko, T. P. et al. Crystal structure of yeast cytosine deaminase. Insights into enzyme mechanism and evolution. J. Biol. Chem. 278, 19111–19117 (2003)
Prochnow, C., Bransteitter, R., Klein, M. G., Goodman, M. F. & Chen, X. S. The APOBEC-2 crystal structure and functional implications for the deaminase AID. Nature 445, 447–451 (2007)
Xie, K. et al. The structure of a yeast RNA-editing deaminase provides insight into the fold and function of activation-induced deaminase and APOBEC-1. Proc. Natl Acad. Sci. USA 101, 8114–8119 (2004)
Iwatani, Y., Takeuchi, H., Strebel, K. & Levin, J. G. Biochemical activities of highly purified, catalytically active human APOBEC3G: correlation with antiviral effect. J. Virol. 80, 5992–6002 (2006)
Chen, K. M. et al. Extensive mutagenesis experiments corroborate a structural model for the DNA deaminase domain of APOBEC3G. FEBS Lett. 581, 4761–4766 (2007)
Navarro, F. et al. Complementary function of the two catalytic domains of APOBEC3G. Virology 333, 374–386 (2005)
Newman, E. N. et al. Antiviral function of APOBEC3G can be dissociated from cytidine deaminase activity. Curr. Biol. 15, 166–170 (2005)
Losey, H. C., Ruthenburg, A. J. & Verdine, G. L. Crystal structure of Staphylococcus aureus tRNA adenosine deaminase TadA in complex with RNA. Nat. Struct. Mol. Biol. 13, 153–159 (2006)
Huthoff, H. & Malim, M. H. Cytidine deamination and resistance to retroviral infection: towards a structural understanding of the APOBEC proteins. Virology 334, 147–153 (2005)
Chung, S. J., Fromme, J. C. & Verdine, G. L. Structure of human cytidine deaminase bound to a potent inhibitor. J. Med. Chem. 48, 658–660 (2005)
Zhang, K. L. et al. Model structure of human APOBEC3G. PLoS ONE 2, e378 (2007)
Mariani, R. et al. Species-specific exclusion of APOBEC3G from HIV-1 virions by Vif. Cell 114, 21–31 (2003)
Teh, A. H. et al. The 1.48 Å resolution crystal structure of the homotetrameric cytidine deaminase from mouse. Biochemistry 45, 7825–7833 (2006)
Xiang, S., Short, S. A., Wolfenden, R. & Carter, C. W. The structure of the cytidine deaminase-product complex provides evidence for efficient proton transfer and ground-state destabilization. Biochemistry 36, 4768–4774 (1997)
Devany, M., Kotharu, N. P. & Matsuo, H. Solution NMR structure of the C-terminal domain of the human protein DEK. Protein Sci. 13, 2252–2259 (2004)
Matsuo, H. et al. Structure of translation factor eIF4E bound to m7GDP and interaction with 4E-binding protein. Nat. Struct. Biol. 4, 717–724 (1997)
Kim, S., Cullis, D. N., Feig, L. A. & Baleja, J. D. Solution structure of the Reps1 EH domain and characterization of its binding to NPF target sequences. Biochemistry 40, 6776–6785 (2001)
Philo, J. S. Improved methods for fitting sedimentation coefficient distributions derived by time-derivative techniques. Anal. Biochem. 354, 238–246 (2006)
Philo, J. S. A method for directly fitting the time derivative of sedimentation velocity data and an alternative algorithm for calculating sedimentation coefficient distribution functions. Anal. Biochem. 279, 151–163 (2000)
Schuck, P. On the analysis of protein self-association by sedimentation velocity analytical ultracentrifugation. Anal. Biochem. 320, 104–124 (2003)
Stafford, W. F. & Sherwood, P. J. Analysis of heterologous interacting systems by sedimentation velocity: curve fitting algorithms for estimation of sedimentation coefficients, equilibrium and kinetic constants. Biophys. Chem. 108, 231–243 (2004)
Matsuo, H., Kupce, E., Li, H. & Wagner, G. Increased sensitivity in HNCA and HN(CO)CA experiments by selective C beta decoupling. J. Magn. Reson. B. 113, 91–96 (1996)
Ikura, M., Kay, L. E. & Bax, A. A novel approach for sequential assignment of 1H, 13C, and 15N spectra of proteins: heteronuclear triple-resonance three-dimensional NMR spectroscopy. Application to calmodulin. Biochemistry 29, 4659–4667 (1990)
Kay, L., Ikura, M., Tschudin, R. & Bax, A. Three-dimensional triple-resonance NMR spectroscopy of isotopically enriched proteins. J. Magn. Reson. 89, 496–514 (1990)
Wittekind, M. & Mueller, J. HNCACB, a high-sensitivity 3D NMR experiment to correlate amide-proton and nitrogen resonances with the alpha- and beta- carbon resonances in proteins. J. Magn. Reson. B. 101, 201–205 (1993)
Yamazaki, T., Lee, W., Arrowsmith, C. H., Muhandiram, D. R. & Kay, L. E. A suite of triple resonance NMR experiments for the backbone assignment of 15N, 13C, 2H labeled proteins with high sensitivity. J. Am. Chem. Soc. 116, 11655–11666 (1994)
Matsuo, H., Kupce, E., Li, H. & Wagner, G. Use of selective C alpha pulses for improvement of HN(CA)CO-D and HN(COCA)NH-D experiments. J. Magn. Reson. B. 111, 194–198 (1996)
Matsuo, H., Li, H. & Wagner, G. A sensitive HN(CA)CO experiment for deuterated proteins. J. Magn. Reson. B. 110, 112–115 (1996)
Clubb, R. T., Thanabal, V. & Wagner, G. A constant-time three-dimensional triple-resonance pulse scheme to correlate intraresidue 1HN, 15N, and 13C′ chemical shifts in 15N-13C-labeled proteins. J. Magn. Reson. 97, 213–217 (1992)
Grzesiek, S., Anglister, J. & Bax, A. Correlation of backbone amide and aliphatic side-chain resonances in 13C/15N-enriched proteins by isotropic mixing of 13C magnetization. J. Magn. Reson. B. 101, 114–119 (1993)
Clore, G. M., Bax, A., Driscoll, P. C., Wingfield, P. T. & Gronenborn, A. M. Assignment of the side-chain 1H and 13C resonances of interleukin-1 beta using double- and triple-resonance heteronuclear three-dimensional NMR spectroscopy. Biochemistry 29, 8172–8184 (1990)
Zhang, O., Kay, L. E., Olivier, J. P. & Forman-Kay, J. Backbone 1H and 15N resonance assignments of the N-terminal SH3 domain of drk in folded and unfolded states using enhanced-sensitivity pulsed field gradient NMR techniques. J. Biomol. NMR 4, 845–858 (1994)
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995)
Keller, R. Optimizing the process of nuclear magnetic resonance spectrum analysis and computer aided resonance assignment. PhD thesis, Swiss Fed. Inst. Tech. Zurich (2004)
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 13, 289–302 (1999)
Herrmann, T., Guntert, P. & Wuthrich, K. Protein NMR structure determination with automated NOE-identification in the NOESY spectra using the new software ATNOS. J. Biomol. NMR 24, 171–189 (2002)
Herrmann, T., Guntert, P. & Wuthrich, K. Protein NMR structure determination with automated NOE assignment using the new software CANDID and the torsion angle dynamics algorithm DYANA. J. Mol. Biol. 319, 209–227 (2002)
Brunger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998)
Desmet, J., De Maeyer, M., Hazas, B. & Lasters, I. The dead-end elimination theorem and its use in protein side-chain positioning. Nature 356, 539–542 (1992)
Goldstein, R. F. Efficient rotamer elimination applied to protein side-chains and related spin glasses. Biophys. J. 66, 1335–1340 (1994)
Acknowledgements
We thank R. LaRue, N. Martemyanova, M. Stenglein and S. Wagner for assistance, laboratory members for discussions, V. Pathak for sharing unpublished information, and J. Lipscomb and K. Walters for comments on the manuscript. Key instrumentation was provided by the University of Minnesota NMR Facility (NSF) and Supercomputing Institute, the University of Wisconsin NMRfam (NIH) and the University of Connecticut Analytical Ultracentrifugation Facility. This work was supported by grants from the National Institutes of Health (A.F., H.M. and R.S.H.), the Medica Foundation (MN Partnership for Biotechnology and Medical Genomics (H.M. and R.S.H.)), the University of Minnesota (H.M. and R.S.H.) and the Searle Scholarship Program (R.S.H.).
Author Contributions H.M. and R.S.H. conceived the experimental designs, wrote the manuscript and assisted with experimentation. K.C., E.H., P.G., Y.L., A.F. and K.S. primarily contributed to protein purification, NMR data analyses, activity assays, site-directed mutagenesis, A3G-2K3A–DNA complex modelling and immunoblotting/purification optimization experiments, respectively. All authors contributed to data analyses, figure constructions and manuscript revisions.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
The file contains Supplementary Discussion with additional references, Supplementary Tables S1-S3 and Supplementary Figures S1-S6 with Legends. (PDF 3551 kb)
Rights and permissions
About this article
Cite this article
Chen, KM., Harjes, E., Gross, P. et al. Structure of the DNA deaminase domain of the HIV-1 restriction factor APOBEC3G. Nature 452, 116–119 (2008). https://doi.org/10.1038/nature06638
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06638
This article is cited by
-
Inhibition of base editors with anti-deaminases derived from viruses
Nature Communications (2022)
-
A rationally engineered cytosine base editor retains high on-target activity while reducing both DNA and RNA off-target effects
Nature Methods (2020)
-
The KT Jeang Prize 2019: Reuben S. Harris
Retrovirology (2019)
-
Off-target RNA mutation induced by DNA base editing and its elimination by mutagenesis
Nature (2019)
-
Crystal structure of the catalytic domain of HIV-1 restriction factor APOBEC3G in complex with ssDNA
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.