Abstract
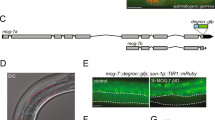
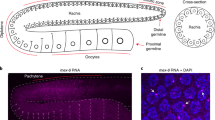
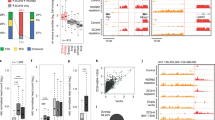
Mutations that enhance the response to double-stranded RNA (dsRNA) have revealed components of the RNA interference (RNAi) pathway or related small RNA pathways. To explore these small RNA pathways, we screened for Caenorhabditis elegans mutants displaying an enhanced response to exogenous dsRNAs. Here we describe the isolation of mutations in two adjacent, divergently transcribed open reading frames (eri-6 and eri-7) that fail to complement. eri-6 and eri-7 produce separate pre-messenger RNAs (pre-mRNAs) that are trans-spliced to form a functional mRNA, eri-6/7. Trans-splicing of eri-6/7 is mediated by a direct repeat that flanks the eri-6 gene. Adenosine to inosine editing within untranslated regions of eri-6 and eri-7 pre-mRNAs reveals a double-stranded pre-mRNA intermediate, forming in the nucleus before splicing occurs. The ERI-6/7 protein is a superfamily I helicase that both negatively regulates the exogenous RNAi pathway and functions in an endogenous RNAi pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Mello, C. C. & Conte, D. Revealing the world of RNA interference. Nature 431, 338–342 (2004)
Zamore, P. D. & Haley, B. Ribo-gnome: the big world of small RNAs. Science 309, 1519–1524 (2005)
Bernstein, E., Caudy, A. A., Hammond, S. M. & Hannon, G. J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001)
Parker, J. S. & Barford, D. Argonaute: a scaffold for the function of short regulatory RNAs. Trends Biochem. Sci. 31, 622–630 (2006)
Duchaine, T. F. et al. Functional proteomics reveals the biochemical niche of C. elegans DCR-1 in multiple small-RNA-mediated pathways. Cell 124, 343–354 (2006)
Simmer, F. et al. Loss of the putative RNA-directed RNA polymerase RRF-3 makes C. elegans hypersensitive to RNAi. Curr. Biol. 12, 1317–1319 (2002)
Kennedy, S., Wang, D. & Ruvkun, G. A conserved siRNA-degrading RNase negatively regulates RNA interference in C. elegans . Nature 427, 645–649 (2004)
Lee, R. C., Hammell, C. M. & Ambros, V. Interacting endogenous and exogenous RNAi pathways in Caenorhabditis elegans . RNA 12, 589–597 (2006)
Alder, M. N., Dames, S., Gaudet, J. & Mango, S. E. Gene silencing in Caenorhabditis elegans by transitive RNA interference. RNA 9, 25–32 (2003)
Pak, J. & Fire, A. Distinct populations of primary and secondary effectors during RNAi in C. elegans . Science 315, 241–244 (2007)
Sijen, T. et al. On the role of RNA amplification in dsRNA-triggered gene silencing. Cell 107, 465–476 (2001)
Kamath, R. S. et al. Systematic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature 421, 231–237 (2003)
Tavernarakis, N., Wang, S. L., Dorovkov, M., Ryazanov, A. & Driscoll, M. Heritable and inducible genetic interference by double-stranded RNA encoded by transgenes. Nature Genet. 24, 180–183 (2000)
Timmons, L., Court, D. L. & Fire, A. Ingestion of bacterially expressed dsRNAs can produce specific and potent genetic interference in Caenorhabditis elegans . Gene 263, 103–112 (2001)
McIntire, S. L., Reimer, R. J., Schuske, K., Edwards, R. H. & Jorgensen, E. M. Identification and characterization of the vesicular GABA transporter. Nature 389, 870–876 (1997)
von Mende, N., Bird, D. M., Albert, P. S. & Riddle, D. L. dpy-13: a nematode collagen gene that affects body shape. Cell 55, 567–576 (1988)
Costa, M. et al. A putative catenin–cadherin system mediates morphogenesis of the Caenorhabditis elegans embryo. J. Cell Biol. 141, 297–308 (1998)
Beitel, G. J., Tuck, S., Greenwald, I. & Horvitz, H. R. The Caenorhabditis elegans gene lin-1 encodes an ETS-domain protein and defines a branch of the vulval induction pathway. Genes Dev. 9, 3149–3162 (1995)
Tabara, H. et al. The rde-1 gene, RNA interference, and transposon silencing in C. elegans . Cell 99, 123–132 (1999)
Fraser, A. G. et al. Functional genomic analysis of C. elegans chromosome I by systematic RNA interference. Nature 408, 325–330 (2000)
WormBase Release. WS170. 〈http://ws170.wormbase.org/〉 (2007)
Wicks, S. R., Yeh, R. T., Gish, W. R., Waterston, R. H. & Plasterk, R. H. Rapid gene mapping in Caenorhabditis elegans using a high density polymorphism map. Nature Genet. 28, 160–164 (2001)
Garcia-Muse, T. & Boulton, S. J. Meiotic recombination in Caenorhabditis elegans . Chromosome Res. 15, 607–621 (2007)
Bass, B. L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71, 817–846 (2002)
Shaner, N. C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nature Biotechnol. 22, 1567–1572 (2004)
Knight, S. W. & Bass, B. L. The role of RNA editing by ADARs in RNAi. Mol. Cell 10, 809–817 (2002)
Wang, D. et al. Somatic misexpression of germline P granules and enhanced RNA interference in retinoblastoma pathway mutants. Nature 436, 593–597 (2005)
Tabara, H., Yigit, E., Siomi, H. & Mello, C. C. The dsRNA binding protein RDE-4 interacts with RDE-1, DCR-1, and a DExH-box helicase to direct RNAi in C. elegans . Cell 109, 861–871 (2002)
Ketting, R. F., Haverkamp, T. H., van Luenen, H. G. & Plasterk, R. H. Mut-7 of C. elegans, required for transposon silencing and RNA interference, is a homolog of Werner syndrome helicase and RNaseD. Cell 99, 133–141 (1999)
Yigit, E. et al. Analysis of the C. elegans Argonaute family reveals that distinct Argonautes act sequentially during RNAi. Cell 127, 747–757 (2006)
Meister, G. et al. Identification of novel argonaute-associated proteins. Curr. Biol. 15, 2149–2155 (2005)
Dalmay, T., Horsefield, R., Braunstein, T. H. & Baulcombe, D. C. SDE3 encodes an RNA helicase required for post-transcriptional gene silencing in Arabidopsis . EMBO J. 20, 2069–2078 (2001)
Tomari, Y. et al. RISC assembly defects in the Drosophila RNAi mutant armitage. Cell 116, 831–841 (2004)
Labrador, M. et al. Protein encoding by both DNA strands. Nature 409, 1000 (2001)
Dorn, R., Reuter, G. & Loewendorf, A. Transgene analysis proves mRNA trans-splicing at the complex mod(mdg4) locus in Drosophila . Proc. Natl Acad. Sci. USA 98, 9724–9729 (2001)
Horiuchi, T., Giniger, E. & Aigaki, T. Alternative trans-splicing of constant and variable exons of a Drosophila axon guidance gene, lola . Genes Dev. 17, 2496–2501 (2003)
Felix, M. A. Genomes: a helpful cousin for our favourite worm. Curr. Biol. 14, R75–R77 (2004)
Obbard, D. J., Jiggins, F. M., Halligan, D. L. & Little, T. J. Natural selection drives extremely rapid evolution in antiviral RNAi genes. Curr. Biol. 16, 580–585 (2006)
Wilkins, C. et al. RNA interference is an antiviral defence mechanism in Caenorhabditis elegans . Nature 436, 1044–1047 (2005)
Schott, D. H., Cureton, D. K., Whelan, S. P. & Hunter, C. P. An antiviral role for the RNA interference machinery in Caenorhabditis elegans . Proc. Natl Acad. Sci. USA 102, 18420–18424 (2005)
Stiernagle, T. in WormBook (ed. the C. elegans research community) 10.1895/wormbook.1.101. 1 〈http://www.wormbook.org〉 (2006)
Sempere, L. F., Dubrovsky, E. B., Dubrovskaya, V. A., Berger, E. M. & Ambros, V. The expression of the let-7 small regulatory RNA is controlled by ecdysone during metamorphosis in Drosophila melanogaster . Dev. Biol. 244, 170–179 (2002)
Caplen, N. J. et al. Rescue of polyglutamine-mediated cytotoxicity by double-stranded RNA-mediated RNA interference. Hum. Mol. Genet. 11, 175–184 (2002)
Acknowledgements
We thank: S. Kennedy for advice and initiating the enhanced RNAi screen; M. Finney for advice on advanced PCR; C. Zhang for mg441 Eri characterization; Y. Kohara, D. Thierry-Mieg and J. Thierry-Mieg for EST sequences and clones; the Mitani laboratory for deletion strains; the Caenorhabditis Genetics Center for strains; Ruvkun laboratory members for reading the manuscript and many discussions; the laboratories of J. Kaplan and F. Ausubel for discussions; and the Leukemia and Lymphoma Society, and EMBO for funding to S.E.J.F.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1–12 and Legends, Supplementary Methods, Supplementary Tables 1–3, a Supplementary Discussion and Supplementary Notes (PDF 3058 kb)
Rights and permissions
About this article
Cite this article
Fischer, S., Butler, M., Pan, Q. et al. Trans-splicing in C. elegans generates the negative RNAi regulator ERI-6/7. Nature 455, 491–496 (2008). https://doi.org/10.1038/nature07274
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07274
This article is cited by
-
Evolutionary honing in and mutational replacement: how long-term directed mutational responses to specific environmental pressures are possible
Theory in Biosciences (2023)
-
Genes that are Used Together are More Likely to be Fused Together in Evolution by Mutational Mechanisms: A Bioinformatic Test of the Used-Fused Hypothesis
Evolutionary Biology (2023)
-
Conservation and diversification of small RNA pathways within flatworms
BMC Evolutionary Biology (2017)
-
Simplification, Innateness, and the Absorption of Meaning from Context: How Novelty Arises from Gradual Network Evolution
Evolutionary Biology (2017)
-
Genomes of parasitic nematodes (Meloidogyne hapla, Meloidogyne incognita, Ascaris suum and Brugia malayi) have a reduced complement of small RNA interference pathway genes: knockdown can reduce host infectivity of M. incognita
Functional & Integrative Genomics (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.