Abstract
Cells employ highly dynamic signaling networks to drive biological decision processes. Perturbations to these signaling networks may attract cells to new malignant signaling and phenotypic states, termed cancer network attractors, that result in cancer development. As different cancer cells reach these malignant states by accumulating different molecular alterations, uncovering these mechanisms represents a grand challenge in cancer biology. Addressing this challenge will require new systems-based strategies that capture the intrinsic properties of cancer signaling networks and provide deeper understanding of the processes by which genetic lesions perturb these networks and lead to disease phenotypes. Network biology will help circumvent fundamental obstacles in cancer treatment, such as drug resistance and metastasis, empowering personalized and tumor-specific cancer therapies.
Similar content being viewed by others
Main
Cells are constantly computing decisions based on the integration of different cues that reach them at various times. In contrast to single-cell organisms, in multicellular organisms, cellular decisions should, ultimately, benefit the organism as a whole, even if that implies that an individual cell will have to decide to commit suicide. In line with this unique feature, signaling networks have evolved during multicellular evolution to allow cells to integrate cues and make decisions that ensure cooperative behavior between them. By hijacking these mechanisms, cancer cells escape cooperative rules and transition from a game governed by Nash equilibria1,2 between all cells into a new scenario where cancer cells decide their behavior purely based on their own benefit, or as phrased by Hanahan and Weinberg3, “become masters of their own destinies.” Given the central role played by signaling networks in the integration of cues to compute any cellular responses, we argue that cancer is not simply a disease with a genetic basis, but is one ultimately driven by perturbations at the signaling network level, and that both the 'cue-signal-response' rules of cellular decision-making and the switch in strategy from cooperative to selfish are major, hitherto understudied, hallmarks of cancer3,4.
In this article, we dissect the strategies cancer cells use to become 'selfish' and drive disease. We first review how genetic lesions can lead to altered protein function, which can result in changes to the structure and dynamics of signaling networks and ultimately cellular phenotype. Next, we describe five general properties of cancer signaling networks (Fig. 1) and define five challenges in cancer network biology and propose strategies to overcome them (Fig. 2). By meeting these challenges, network biology may fundamentally advance not only basic biology but also patient treatment. Finally, we describe how a combination of relatively new technologies could become a potent cocktail for the discovery of network drugs, and we discuss the practical implementation of personalized and tumor-specific cancer therapy.
(a) Analogous mutations. Two different tumors may achieve the same signaling and phenotypic outcome with two different mutations (b) Synthetic oncogenes. Mutations that are not oncogenic on their own can cooperate when appearing together to drive tumor formation11; by analogy to synthetic lethality, we call the genes harboring cooperative mutations, synthetic oncogenes. (c) Multivariate nature of signaling networks. The response of a cell to a specific cue depends on, and can only be predicted by taking into account, the state of the cellular signaling networks25. This dependency, known as the multivariate nature of signaling networks, is often neglected when classifying mutations and genes as oncogenes or tumor suppressors and cancer drivers or passengers. (d) Dynamic networks. Although signaling networks are often represented as static, it is clear that they are highly dynamic entities. Given that the role of signaling networks in computing cellular responses is highly dependent on it, and that cancer mutations will perturb it, this dynamic nature is a critical property of cancer signaling networks. (e) Signaling network landscapes. The different states that a signaling network occupies can be represented as a landscape (with stable steady states or attractors represented as valleys and unstable steady states represented as hills), where the cell constantly gets pushed by signaling cues31,32,39,40. These states drive cellular and disease phenotypes and represent network drug targets.
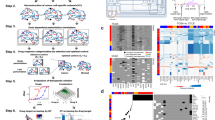
(a) Functional consequences of cancer mutations. Using an ensemble of protein-function features (e.g., ATP binding, substrate specificity, activation of the protein kinase or phospho-tyrosine binding), which together represent a comprehensive description of a protein's molecular functions, will enable more accurate and predictive evaluation of cancer mutations. (b) Modeling of disease networks. Although experimental and computational tools for modeling molecular networks exist, creating more comprehensive, sensitive and accurate new tools especially designed to model disease-associated networks still represents a big challenge in network biology. (c) Network-attacking mutations and cancer network attractors. Network-attacking mutations are mutations that lead to a new cellular phenotype by perturbing signaling networks either at the network structure or the network dynamics level. Network-attacking mutations transform signaling networks, generating new possible network states by changing the number and/or stability of steady states in the signaling landscape31,32,39,40. These acquired signaling capabilities lead to alterations in the cell's normal 'cue-signal-output' flow and thereby drive disease phenotypes (see Fig. 3 for further details). (d) Tumor subpopulations and micro-environment. The field is only beginning to comprehend the complex interactions that exist between different co-evolving tumor cell subpopulations and between those cells and the tumor microenvironment, both of which strongly influence tumor progression. (e) Network-aware and temporal drugs. As predicted by R.L. and Pawson66 several years ago, new pharmaceutical strategies that target networks instead of single proteins are becoming available47,48. We predict this trend will not only continue, but also include recent advances that highlight the possibility to 'cure' networks using time- and order-dependent therapies68. In coming years, the discovery of resistant, metastatic, tissue or cell-specific networks could lead to an even greater advance in the field of network medicine (Fig. 5).
From genomic lesions to functional network perturbations
Tumor cells often harbor hundreds to thousands of genetic lesions. But based on the observation that some of these genetic lesions are repeatedly observed in several cancers (e.g., BRAF V600E, present in >50% of all malignant melanomas5), it has been hypothesized that only a few genetic lesions are causally implicated in cancer development ('drivers'), whereas the majority have no functional consequences ('passengers')6.
Although this classification has had some use in identifying mutations that are highly prevalent, it is now apparent that a tumor is not, under any circumstances, a static and uniform population of malignant cells. Rather, it is a dynamic ensemble of subpopulations with different abnormalities undergoing molecular evolution7,8,9. Two fundamental principles of cancer signaling networks can explain why a binary driver/passenger classification may be too simplistic to accommodate the complex dynamic nature of tumors. First, different tumors can develop similar phenotypes by acquiring mutations in different proteins10, in what we term analogous mutations (Fig. 1a). Second, it has been shown that two different mutations not capable of causally driving cancer by themselves are able to do so when they appear in combination within the same cells or even within two neighboring cells11, in what could be described as two passengers becoming drivers or, as we refer to them, synthetic oncogenes (Fig. 1b). Thus, patient-to-patient heterogeneity can be driven by the presence of different mutations in the same or in different proteins that lead to a similar signaling state and phenotypic outcome.
Altogether, the intrinsic heterogeneity of tumors makes it a pressing challenge for cancer network biologists to develop tools to identify the extent to which combinations of cancer mutations affect protein function and cellular and phenotypic states (Fig. 2a,b). Even though several such tools have been developed (reviewed in ref. 12), existing methods are mainly based on protein structure and/or sequence conservation. This is at odds with recent findings that show that cancer mutations tend not to cluster on the most conserved protein regions. In kinases, for example, mutations typically hit the kinase activation segment, a functional, yet largely nonconserved protein region13. Because cancer cells would obtain the greatest fitness advantage from mutations that target the most-functional residues, we reason that a better understanding of the functionality of protein residues would allow more accurate predictions of the consequences of cancer mutations. Functional residues have been defined as those residues required for a protein to perform its molecular function(s), in the sense that they cannot be freely changed without directly affecting the role(s) of the protein14. Here we extend this definition to include a more fine-grained and precise definition of protein function as an ensemble of protein features that together describe the different functional capabilities of proteins (e.g., ATP binding, substrate specificity, protein activation or phospho-tyrosine binding). This new definition would not only adapt well to current studies of sequence-function associations15,16, but also lead to a better description of the effects of a mutation affecting such residues (Fig. 2a,b).
An insightful example of how to explore this sequence-function relationship in protein domains was carried out by researchers in the Ranganathan and Yaffe laboratories who, using methods from statistical mechanics, generated synthetic WW domains de novo that maintained fold and function17,18. Further supporting a complex sequence-function relationship, additional studies from the Ranganathan laboratory demonstrated that, in addition to protein architecture described as combinations of modules such as globular domains and linear motifs19,20,21, protein domains themselves often have well-defined sectors formed by sparse networks of residues often linking spatially distant regions that contribute cooperatively but unequally to its function22,23. Although some targeted studies analyzing several cancer mutations in a single kinase have been conducted24, similar approaches to those used for WW domains should be pursued to generate high-throughput experimental studies of cancer mutations in the context of signaling networks. These would help gain a better understanding of which amino acid residues can be changed freely without affecting the protein and network function and, most importantly, which cannot.
From network perturbations to cellular phenotypes
The characterization of cellular signaling processes has largely focused on identifying the function of individual genes and proteins. A notable exception is a landmark study25 on the context dependence of the Jun-activated kinase (JNK) in apoptosis. Before this work, paradoxical results suggested that JNK had a pro-apoptotic function26, an anti-apoptotic function27 or even a lack of involvement in apoptosis28. The systematic approach undertaken by Janes et al.25 revealed that the phosphorylation status of JNK (and thus its catalytic activity) was not sufficient to determine apoptotic commitment; instead, activation of JNK could lead to both apoptosis and proliferation depending on the cellular signaling network state at the time of activation. Thus, this work demonstrated that a protein's cellular role is not a static property but rather can only be defined dynamically—that is, its role depends on the context of the network it is operating within. Similar context dependencies have been confirmed for other kinases, such as Erk and MK2. Because of this, which is referred to as the multivariate property of signaling networks (Fig. 1c), we suggest that it is essential to study cellular context at the systems level.
Although these multivariate molecular networks seem to have evolved a complex structure that makes them robust against deletion of a few proteins29, they are highly dynamic. Thus, a more accurate description of signaling networks should take into account the fact that a single static network does not exist unchanged over time. Instead, a cell contains a dynamic ensemble of networks whose different permutations are manifested in the cell depending on the different cues the cell is presented with (Fig. 1d). This dynamic nature of signaling networks could, at least in part, explain why all mutant proteins do not seem to be expressed at a given point in time30, if a substantial part of the proteome is so dynamic that it is expressed only when the cell senses a specific cue.
Moreover, according to a general principle of complex systems introduced in the 1980s31,32, dynamic cellular networks can only exist in a finite number of states, owing to the constraints that interactions between nodes impose on one another. These network states can be represented as landscapes, where most-probable and least-probable states are represented as valleys and mountains, respectively (Fig. 1e). Cells are continuously exploring this landscape and are pushed from one state to another by different environmental or intracellular cues.
Implications for cancer research
The multivariate nature of signaling networks has profound implications for cancer research. Just as it is inaccurate to assign a static function (e.g., apoptotic or anti-apoptotic) to a single protein, it is clear that static interpretations of mutations, that is, driver or passenger mutations, are also misleading. For example, given that the phenotypic role of JNK strongly depends on network state, it is clear that a mutation in JNK (and thus probably any other mutation) should not be statically labeled as a driver or passenger or as an oncogene or tumor suppressor, as such classifications are context dependent (e.g., disease or cell-type specific). Several examples, such as Myc33 or WT1 (ref. 34) gene products that act as both tumor suppressors and oncogenes, support this idea. These results underscore the importance of assessing mutations based on their effects on signaling networks and of developing novel classification methods to do so. Along these lines, MAP2K4 (one of the protein kinases that can phosphorylate and activate JNK) has been shown to be recurrently lost or mutated in several cancers35,36,37,38. These represent prime examples of mutations that may display ambivalent phenotypic impact similar to JNK.
Motivated by the example of MAP2K4 and many other mutated kinases38, we maintain that mutations capable of affecting signaling networks—which we call network-attacking mutations (Fig. 2c)—are more likely to affect phenotype than other mutations. Thus, we discuss a general strategy in which mutations in individual cancers are assessed based on, first, the likelihood they will affect protein function, and second, the cellular role of the signaling network that they are operating within (Fig. 3). Our strategy extends the concepts introduced by Waddington and elaborated by Kauffman and Huang et al.31,32,39,40, where cancer mutations are turned into perturbations capable of reshaping these landscapes. We represent the cellular response or phenotype as another dimension where each network state (every point in the landscape) is constantly projected to and translated into a cellular decision or phenotypic outcome.
Proteins are the key elements of signaling networks as a result of their ability to integrate external cues and direct the information flow toward a specific cellular outcome (e.g., epidermal growth factor (EGF) leading to proliferation or tumor necrosis factor alpha (TNF-α) leading to apoptosis). Network-attacking mutations affect the 'cue-signal-output' cellular information flow by affecting either the dynamics (middle), for example, by keeping proteins constitutively active, or the structure (right), by affecting protein specificity, of the signaling networks. Signaling networks can be represented as a landscape with the most likely network states represented as valleys (stable steady states or attractors) and the least likely network states as mountains (unstable steady states). Network-attacking mutations dysregulate signaling networks by perturbing the number and/or stability of steady states in the landscape, effectively creating new cancer-specific attractors that only cancer cells will be able to reach.
We postulate that network-attacking mutations affect the cell not by perturbing how the signaling landscape is projected to the phenotypic dimension, but by changing the ensemble of dynamic networks that can be manifested in a cell and, in consequence, the number and stability of steady states in the signaling landscape, thus creating new attractor states that only cancer cells can occupy, also known as cancer network attractors (Fig. 3). This has additional implications for other mechanisms, such as oncogene and non-oncogene addiction41, where cancer cells would be trapped in cancer attractor states and could escape from them by reverting the genomic aberration that initially caused the perturbed landscape. Given the high degree of determinism that exists between signaling networks, landscapes and phenotypes, we argue that network-attacking mutations are at the heart of all new decision-making capabilities acquired by cancer cells. Consequently, in our view, the study of both network-attacking mutations and new attractor states acquired by cancer cells, that is, cancer network attractors, deserves the highest priority from the field. Such studies should be performed through systematic and quantitative sampling of cell dynamics at multiple levels (e.g., genomic or epigenetic, proteomic and phenotypic), followed by nonlinear interpolation and integrative computational modeling (Fig. 4).
In more traditional biological approaches, where only one or a few genes or proteins are sampled across a limited set of conditions, there has been limited success in deriving predictive models across conditions or cell types that would require comprehensive sampling. In contrast, network biology relies on systematic sampling across combinations of states that result in increased performance of a network model. Unlike classic approaches, in which the system is stimulated with single specific cellular cues (e.g., growth factor), in the network biology approach, the multivariate nature of signaling networks and the nonlinear relationship between signaling input and output can be successfully elucidated by interrogating the system with multiple orthogonal cues.
The first network-attacking cancer mutation, described more than 15 years ago42, was a point mutation in the kinase domain of RET (M918T), which leads to a switch in peptide specificity. In line with their importance, network-attacking mutations have attracted more attention in recent years43,44,45,46,47,48. Moreover, information has been accumulating steadily about how specificity in signaling networks and modular protein domains emerges49,50,51, leading to the definition of determinants of specificity in protein domains52,53. These determinants, sometimes referred to as specificity-determining residues, are residues that can lead to substrate specificity changes after mutation. Notably, direct mutagenesis of these determinants of specificity has been used to rewire the entire histidine kinase signaling system in bacteria in a predictive manner54. Recent follow-up work indicates that mutations in determinants of specificity prevent cross-talk and allow protein family expansions55, in a process similar to the one powered by negative selection over Src homology 3 (SH3) protein domains that show similar specificity56. We propose that similar studies in human signaling networks, coupled with mapping of cancer mutations on these determinants of specificity, would shed new light on whether signaling rewiring is a general principle of oncogenesis and tumor progression, knowledge of which would in turn be critical as molecular therapies target proteins and their networks and not genes.
Despite the fact that the number of known cancer network-attacking mutations is still relatively low, recent findings suggest that in-frame mutations are enriched on interaction interfaces57, which implies they are also likely to affect determinants of specificity. Moreover, many fusion proteins have been discovered that likely directly rewire or create new network states58. Given the rate at which cancer mutations are being reported and the development of new computational methods for systematically identifying these mutations (Fig. 2b), we predict a steep increase in the number of network-attacking mutations that will be uncovered in the coming years.
Personalized cancer network biology
Led by recent advances in sequencing technologies, the amount of data on cancer genome mutations is growing exponentially59. Current efforts from the Cancer Genome Atlas and Cancer Genome Project, now under the umbrella of the International Cancer Genome Consortium60, will facilitate the annotation and collection of cancer genome data. We foresee similar waves of technological progress and the generation of new consortiums in the cancer proteomics fields in the near future. The establishment of the Clinical Proteomic Tumor Analysis Consortium (http://proteomics.cancer.gov/programs/cptacnetwork), and the implementation of new approaches61 and labeling techniques62 optimized for patient samples are encouraging advances in this direction.
These advances, however, will need to be coordinated with new algorithmic and experimental high-throughput methods (e.g., high-content screening) capable of interpreting this flood of information because the functional interpretation of the data is currently the main bottleneck in the field of personalized cancer network biology. Computational integration of large quantitative data sets is also becoming increasingly important, and thus there is a growing requirement for supercomputing infrastructure with large algorithmic dynamic range (e.g., next-generation large shared memory systems). Benchmarking and validation of systematic workflows and algorithms is already receiving increasing attention through initiatives, such as the DREAM challenge63 and IMPROVER64.
Two emerging areas in network biology that are likely to contribute to the future of cancer research are the study of cell-cell interactions (Fig. 2d) and drugs specifically designed to interfere with diseased network dynamics (that is, network drugs; Fig. 2e).
R.L. and collaborators65 studied cell-cell interactions by isotopically labeling two distinct subpopulations of cells, one expressing ephrin-B1+ and the other Eph-B2+, and carrying out a comprehensive phospho-proteomic analysis. This strategy facilitated the first measurements of phosphorylation events during the interaction of two cell subpopulations. The proliferative behavior of cancer cells is still poorly understood in part because it is difficult to experimentally study the transmission of proliferative factors from one cell to its neighbors3. Therefore, we argue that a similar isotopic labeling strategy could be used to investigate the cooperation between cells with different oncogenic lesions that together (that is, synthetic oncogenes; Figs. 1b and 2d) lead to tumor formation11.
Combination drugs that interfere with disease networks (so-called network medicine66) have been shown to lead to a better response than single-hit therapies by causing secondary perturbations to signaling networks47,48,67. Recent work by the Yaffe laboratory represents a clear leap forward within the field of network medicine68,69. Following network modeling, Yaffe and colleagues68 managed to decode the signaling network dynamics that drive resistance to DNA-damaging chemotherapy. This information was used to sensitize otherwise resistant triple-negative breast cancer cells to conventional DNA-damaging chemotherapy by administering doxorubicin (Adriamycin, Doxil) and erlotinib (Tarceva) in an order- and time-dependent fashion. This could be considered the first example of temporal network drugs (Figs. 2e and 5).
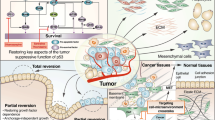
The goal of personalized cancer network biology is to be able to treat each tumor with the best combination of drugs tailored to that tumor. Ideally, early diagnosis should be followed by the development of tumor-specific cell lines and xenograft models, cancer genome sequencing, and proteomic and phenotypic analysis. Combinations of network drugs should then be tried in the tumor-specific cell line and xenograft model and eventually transferred back to the patient. Continuing to treat the tumor-specific cell culture with the same network drug combination as is used in the patient may be useful for understanding potential resistance and/or metastasis.
We predict that personalized or even tumor-specific cancer therapy will become a reality in the foreseeable future, starting from early diagnosis of the disease, followed by next-generation sequencing, proteomic analysis, high-throughput profiling of phenotypic cell states in the tumor and design of patient-specific combinations of network drugs with resistance follow-up (Fig. 5). Relatively new techniques, such as single-cell and high-depth sequencing70,71, imaging72 and cytometry time-of-flight73, could prove especially valuable for monitoring the number, properties and behavior of different tumor subclones (Fig. 2d). Ideally, network drugs, such as the aforementioned order- and time-dependent combination68, should then be chosen based on the interpretation of sequencing as well as the proteomic and phenotypic analysis of tumor cells and tested on the tumor-specific cell lines and xenograft model. The best-performing combination should ultimately be transferred back to the patient (Fig. 5). This whole process should take the shortest time possible to avoid the evolution of the tumor in the patient and the consequent loss of relationship between the primary tumor and the cell line. Tumor-specific cell lines would be kept and treated with the same drugs used in the patient to monitor tumor evolution and treat for resistance and/or metastasis as soon as there is enough evidence of it (Fig. 5). Ideally, every patient and paired xenograft or cell line should have a complete electronic record showing the treatment history to facilitate retrospective and cross-disease studies74,75.
Conclusions
Although we have highlighted some of the challenges that still exist in cancer network biology, substantial progress is also being made. For example, the usage of patient-derived tumor tissue in animal xenograft models to test the response to particular drugs aimed at developing new personalized cancer therapy is rapidly becoming an established technology76. Surgical orthotopic implantation to transplant tumors taken directly from the patient to the corresponding organ of immunodeficient mice77 is currently one of the most promising methods to enable drug screening in patients. In addition, new clinical trials, such as the MD Anderson T9 project78, are under way in which patients are given therapy that targets tumor-specific aberrations. Nevertheless, the implementation of the strategy depicted in Figure 5 would benefit from further developments in technology, funding and legislation. For example, generating models for cancer research that represent human patient diversity79 and mimicking the complexity of tumor microenvironments (J.T.E. and collaborators)80 remain extraordinary challenges (Fig. 2), and further research efforts and investments are required. As cancer biology becomes a 'big data' science, similar to physics, we expect to see more systematic, data-driven research efforts that will uncover and confront many of the tumor complexities that have remained elusive so far.
Despite recent predictions of >13 million cancer deaths in 2030 (ref. 81), as discussed in this Perspective, we foresee that within this timeframe tumor-specific medicine will become a reality, thanks to a new generation of cancer network biologists who will hopefully overcome these challenges, positively contributing to the battle against this devastating disease and the significant reduction of patient suffering.
Author contributions
All correspondence should be addressed to both J.T.E. and R.L.
References
Nash, J.F. Equilibrium points in N-person games. Proc. Natl. Acad. Sci. USA 36, 48–49 (1950).
Nash, J.F. Non-cooperative games. Ann. Math. 54, 286–295 (1951).
Hanahan, D. & Weinberg, R.A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Hanahan, D. & Weinberg, R.A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Davies, H. et al. Mutations of the BRAF gene in human cancer. Nature 417, 949–954 (2002).
Stratton, M.R., Campbell, P.J. & Futreal, P.A. The cancer genome. Nature 458, 719–724 (2009).
Ding, L. et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 481, 506–510 (2012).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Hou, Y. et al. Single-cell exome sequencing and monoclonal evolution of a JAK2-negative myeloproliferative neoplasm. Cell 148, 873–885 (2012).
Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Wu, M., Pastor-Pareja, J.C. & Xu, T. Interaction between RasV12 and scribbled clones induces tumour growth and invasion. Nature 463, 545–548 (2010).
Ng, P.C. & Henikoff, S. Predicting the effects of amino acid substitutions on protein function. Annu. Rev. Genomics Hum. Genet. 7, 61–80 (2006).
Dixit, A. et al. Sequence and structure signatures of cancer mutation hotspots in protein kinases. PLoS ONE 4, e7485 (2009).
Pazos, F. & Bang, J.-W. Computational prediction of functionally important regions in proteins. Curr. Bioinform. 1, 15–23 (2006).
Fowler, D.M. et al. High-resolution mapping of protein sequence-function relationships. Nat. Methods 7, 741–746 (2010).
Jensen, L.J. et al. Ab initio prediction of human orphan protein function from post-translational modifications and localization features. J. Mol. Biol. 319, 1257–1265 (2002).
Socolich, M. et al. Evolutionary information for specifying a protein fold. Nature 437, 512–518 (2005).
Russ, W., Lowery, D., Mishra, P., Yaffe, M. & Ranganathan, R. Natural-like function in artificial WW domains. Nature 437, 579–583 (2005).
Puntervoll, P. et al. ELM server: a new resource for investigating short functional sites in modular eukaryotic proteins. Nucleic Acids Res. 31, 3625–3630 (2003).
Lim, W.A. & Pawson, T. Phosphotyrosine signaling: evolving a new cellular communication system. Cell 142, 661–667 (2010).
Seet, B.T., Dikic, I., Zhou, M.M. & Pawson, T. Reading protein modifications with interaction domains. Nat. Rev. Mol. Cell Biol. 7, 473–483 (2006).
Halabi, N., Rivoire, O., Leibler, S. & Ranganathan, R. Protein sectors: Evolutionary units of three-dimensional structure. Cell 138, 774–786 (2009).
Reynolds, K.A., McLaughlin, R. & Ranganathan, R. Hot spots for allosteric regulation on protein surfaces. Cell 147, 1564–1575 (2011).
Wan, P.T. et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 116, 855–867 (2004).
Janes, K.A. et al. A systems model of signaling identifies a molecular basis set for cytokine-induced apoptosis. Science 310, 1646–1653 (2005).
Lei, K. & Davis, R.J. JNK phosphorylation of Bim-related members of the Bcl2 family induces Bax-dependent apoptosis. Proc. Natl. Acad. Sci. USA 100, 2432–2437 (2003).
Lamb, J.A. et al. JunD mediates survival signaling by the JNK signal transduction pathway. Mol. Cell 11, 1479–1489 (2003).
Abreu-Martin, M.T. et al. Fas activates the JNK pathway in human colonic epithelial cells: lack of a direct role in apoptosis. Am. J. Physiol. 276, G599 (1999).
Jeong, H., Mason, S.P., Barabasi, A.L. & Oltvai, Z.N. Lethality and centrality in protein networks. Nature 411, 41–42 (2001).
Shah, S.P. et al. The clonal and mutational evolution spectrum of primary triplenegative breast cancers. Nature advance online publication, doi:10.1038/nature10933 (4 April 2012).
Kauffman, S. & Levin, S. Towards a general theory of adaptive walks on rugged landscapes. J. Theor. Biol. 128, 11–45 (1987).
Kauffman, S.A. & Weinberger, E.D. The NK model of rugged fitness landscapes and its application to maturation of the immune response. J. Theor. Biol. 141, 211–245 (1989).
Uribesalgo, I., Benitah, S.A. & Di Croce, L. From oncogene to tumor suppressor: The dual role of Myc in leukemia. Cell Cycle 11, 1757–1764 (2012).
Yang, L., Han, Y., Saurez Saiz, F. & Minden, M.D. A tumor suppressor and oncogene: the WT1 story. Leukemia 21, 868–876 (2007).
Ellis, M.J. et al. Whole-genome analysis informs breast cancer response to aromatase inhibition. Nature 486, 353–360 (2012).
Curtis, C. et al. The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Kan, Z. et al. Diverse somatic mutation patterns and pathway alterations in human cancers. Nature 466, 869–873 (2010).
Greenman, C. et al. Pattern of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007).
Waddington, C.H. The Strategy of the Genes: a Discussion of Some Aspects of Theoretical Biology (Allen & Unwin, 1957).
Huang, S. & Ingber, D.E. Shape-dependent control of cell growth, differentiation, and apoptosis: switching between attractors in cell regulatory networks. Exp. Cell Res. 261, 91–103 (2000).
Luo, J., Solimini, N.L. & Elledge, S.J. Principles of cancer therapy: oncogene and non-oncogene addiction. Cell 136, 823–837 (2009).
Songyang, Z. et al. Catalytic specificity of protein-tyrosine kinases is critical for selective signaling. Nature 373, 536–539 (1995).
Zhong, Q. et al. Edgetic perturbation models of human inherited disorders. Mol. Syst. Biol. 5, 321 (2009).
Dreze, M. et al. 'Edgetic' perturbation of a C. elegans BCL2 ortholog. Nat. Methods 6, 843–849 (2009).
Pe'er, D. & Hacohen, N. Principles and strategies for developing network models in cancer. Cell 144, 864–873 (2011).
Vidal, M., Cusick, M.E. & Barabási, A.-L.L. Interactome networks and human disease. Cell 144, 986–998 (2011).
Schoeberl, B. et al. Therapeutically targeting ErbB3: a key node in ligand-induced activation of the ErbB receptor-PI3K axis. Sci. Signal. 2, ra31 (2009).
Huang, P.H. et al. Quantitative analysis of EGFRvIII cellular signaling networks reveals a combinatorial therapeutic strategy for glioblastoma. Proc. Natl. Acad. Sci. USA 104, 12867–12872 (2007).
Miller, M.L.L. et al. Linear motif atlas for phosphorylation-dependent signaling. Sci. Signal. 1, ra2+ (2008).
Linding, R. et al. Systematic discovery of in vivo phosphorylation networks. Cell 129, 1415–1426 (2007).
Mok, J. et al. Deciphering protein kinase specificity through large-scale analysis of yeast phosphorylation site motifs. Sci. Signal. 3, ra12 (2010).
Brinkworth, R.I., Breinl, R.A. & Kobe, B. Structural basis and prediction of substrate specificity in protein serine/threonine kinases. Proc. Natl. Acad. Sci. USA 100, 74–79 (2003).
Turk, B.E. Understanding and exploiting substrate recognition by protein kinases. Curr. Opin. Chem. Biol. 12, 4–10 (2008).
Skerker, J.M. et al. Rewiring the specificity of two-component signal transduction systems. Cell 133, 1043–1054 (2008).
Capra, E.J., Perchuk, B.S., Skerker, J.M. & Laub, M.T. Adaptive mutations that prevent crosstalk enable the expansion of paralogous signaling protein families. Cell 150, 222–232 (2012).
Zarrinpar, A., Park, S.H. & Lim, W.A. Optimization of specificity in a cellular protein interaction network by negative selection. Nature 426, 676–680 (2003).
Wang, X. et al. Three-dimensional reconstruction of protein networks provides insight into human genetic disease. Nat. Biotechnol. 30, 159–164 (2012).
Brehme, M. et al. Charting the molecular network of the drug target Bcr-Abl. Proc. Natl. Acad. Sci. USA 106, 7414–7419 (2009).
Wong, K.M.M., Hudson, T.J. & McPherson, J.D. Unraveling the genetics of cancer: genome sequencing and beyond. Annu. Rev. Genomics Hum. Genet. 12, 407–430 (2011).
Ledford, H. Big science: the cancer genome challenge. Nature 464, 972–974 (2010).
Bensimon, A., Heck, A.J.R. & Aebersold, R. Mass spectrometry-based proteomics and network biology. Annu. Rev. Biochem. 81, 379–405 (2012).
Geiger, T., Cox, J., Ostasiewicz, P., Wisniewski, J.R. & Mann, M. Super-SILAC mix for quantitative proteomics of human tumor tissue. Nat. Methods 7, 383–385 (2010).
Prill, R.J. et al. Towards a rigorous assessment of systems biology models: the DREAM3 challenges. PLoS ONE 5, e9202 (2010).
Meyer, P. et al. Verification of systems biology research in the age of collaborative competition. Nat. Biotechnol. 29, 811–815 (2011).
Jørgensen, C. et al. Cell-specific information processing in segregating populations of Eph receptor ephrin-expressing cells. Science 326, 1502–1509 (2009).
Pawson, T. & Linding, R. Network medicine. FEBS Lett. 582, 1266–1270 (2008).
Chandarlapaty, S. et al. AKT inhibition relieves feedback suppression of receptor tyrosine kinase expression and activity. Cancer Cell 19, 58–71 (2011).
Lee, M.J. et al. Sequential application of anticancer drugs enhances cell death by rewiring apoptotic signaling networks. Cell 149, 780–794 (2012).
Erler, J.T. & Linding, R. Network medicine strikes a blow against breast cancer. Cell 149, 731–733 (2012).
Navin, N. et al. Tumor evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Pedersen, M.W. et al. Sym004: a novel synergistic anti-epidermal growth factor receptor antibody mixture with superior anticancer efficacy. Cancer Res. 70, 588–597 (2010).
Bendall, S.C. & Nolan, G.P. From single cells to deep phenotypes in cancer. Nat. Biotechnol. 30, 639–647 (2012).
Roque, F.S. et al. Using electronic patient records to discover disease correlations and stratify patient cohorts. PLOS Comput. Biol. 7, e1002141 (2011).
Jensen, P.B., Jensen, L.J. & Brunak, S. Mining electronic health records: towards better research applications and clinical care. Nat. Rev. Genet. 13, 395–405 (2012).
Blumenthal, R.D. & Goldenberg, D.M. Methods and goals for the use of in vitro and in vivo chemosensitivity testing. Mol. Biotechnol. 35, 185–197 (2007).
Hoffman, R.M. Orthotopic mouse models expressing fluorescent proteins for cancer drug discovery. Expert Opin. Drug Discov. 5, 851–866 (2010).
Gonzalez-Angulo, A.M., Hennessy, B.T. & Mills, G.B. Future of personalized medicine in oncology: a systems biology approach. J. Clin. Oncol. 28, 2777–2783 (2010).
Hunter, K.W. Mouse models of cancer: does the strain matter? Nat. Rev. Cancer 12, 144–149 (2012).
Cox, T.R. & Erler, J.T. Remodeling and homeostasis of the extracellular matrix: implications for fibrotic diseases and cancer. Dis. Model. Mech. 4, 165–178 (2011).
WHO. World health organization fact sheet 297 (2012). http://www.who.int/mediacentre/factsheets/fs297/en/
Acknowledgements
We apologize to our colleagues whose work could not be cited due to space limitations. We thank all members of the C-SIG (DTU), the ErlerLab (BRIC), M. Yaffe (MIT) and N. Brunner (KU) for critical input on this manuscript. R.L. is a Lundbeck Foundation Fellow and is supported by a Sapere Aude Starting Grant from The Danish Council for Independent Research and a Career Development Award from Human Frontier Science Program. J.T.E. is supported by a Hallas Møller Stipend from the Novo Nordisk Foundation. Visit http://www.networkbio.org/, http://www.lindinglab.org/ and http://www.erlerlab.org/ for more information on cancer-related network biology.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Creixell, P., Schoof, E., Erler, J. et al. Navigating cancer network attractors for tumor-specific therapy. Nat Biotechnol 30, 842–848 (2012). https://doi.org/10.1038/nbt.2345
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2345
This article is cited by
-
Identification of anticancer drug target genes using an outside competitive dynamics model on cancer signaling networks
Scientific Reports (2021)
-
WASABI: a dynamic iterative framework for gene regulatory network inference
BMC Bioinformatics (2019)
-
Minimal intervening control of biomolecular networks leading to a desired cellular state
Scientific Reports (2019)
-
A bioprinted human-glioblastoma-on-a-chip for the identification of patient-specific responses to chemoradiotherapy
Nature Biomedical Engineering (2019)
-
Systems Pharmacogenomic Landscape of Drug Similarities from LINCS data: Drug Association Networks
Scientific Reports (2019)