Abstract
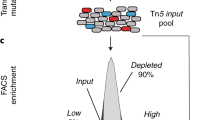
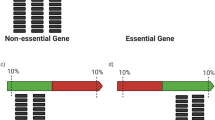
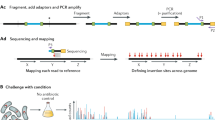
We describe a method of genome-wide analysis of quantitative growth phenotypes using insertional mutagenesis and DNA microarrays. We applied the method to assess the fitness contributions of Escherichia coli gene domains under specific growth conditions. A transposon library was subjected to competitive growth selection in Luria–Bertani (LB) and in glucose minimal media. Transposon-containing genomic DNA fragments from the selected libraries were compared with the initial unselected transposon insertion library on DNA microarrays to identify insertions that affect fitness. Genes involved in the biosynthesis of nutrients not provided in the growth medium were found to be significantly enriched in the set of genes containing negatively selected insertions. The data also identify fitness contributions of several uncharacterized genes, including putative transcriptional regulators and enzymes. The applicability of this high-resolution array selection in other species is discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Riley, M. Genes and proteins of Escherichia coli K-12. Nucleic Acids Res. 26, 54 (1998).
Smith, V., Chou, K.N., Lashkari, D., Botstein, D. & Brown, P.O. Functional analysis of the genes of yeast chromosome V by genetic footprinting. Science 274, 2069–2074 (1996).
Winzeler, E.A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Akerley, B.J. et al. Systematic identification of essential genes by in vitro mariner mutagenesis. Proc. Natl. Acad. Sci. USA 95, 8927–8932 (1998).
Wong, S.M. & Mekalanos, J.J. Genetic footprinting with mariner-based transposition in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 97, 10191–10196 (2000).
Hutchison, C.A. et al. Global transposon mutagenesis and a minimal Mycoplasma genome. Science 286, 2165–2169 (1999).
Hare, R.S. et al. Genetic footprinting in bacteria. J. Bacteriol. 183, 1694–1706 (2001).
Schena, M., Shalon, D., Davis, R.W. & Brown, P.O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Iyer, V.R. et al. Genomic binding sites of the yeast cell-cycle transcription factors SBF and MBF. Nature 409, 533–538 (2001).
Ren, B. et al. Genome-wide location and function of DNA binding proteins. Science 290, 2306–2309 (2000).
Pollack, J.R. et al. Genome-wide analysis of DNA copy-number changes using cDNA microarrays. Nat. Genet. 23, 41–46 (1999).
Alexeyev, M.F. & Shokolenko, I.N. Mini-Tn10 transposon derivatives for insertion mutagenesis and gene delivery into the chromosome of gram-negative bacteria. Gene 160, 59–62 (1995).
Bender, J. & Kleckner, N. IS10 transposase mutations that specifically alter target site recognition. EMBO J. 11, 741–750 (1992).
Tavazoie, S. & Church, G.M. Quantitative whole-genome analysis of DNA–protein interactions by in vivo methylase protection in E. coli. Nat. Biotechnol. 16, 566–571 (1998).
Robison, K., McGuire, A.M. & Church, G.M. A comprehensive library of DNA-binding site matrices for 55 proteins applied to the complete Escherichia coli K-12 genome. J. Mol. Biol. 284, 241–254 (1998).
Arigoni, F. et al. A genome-based approach for the identification of essential bacterial genes. Nat. Biotechnol. 16, 851–856 (1998).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Edwards, J.S. & Palsson, B.O. The Escherichia coli MG1655 in silico metabolic genotype: its definition, characteristics, and capabilities. Proc. Natl. Acad. Sci. USA 97, 5528–5533 (2000).
Edwards, J.S., Ibarra, R.U. & Palsson, B.O. In silico predictions of Escherichia coli metabolic capabilities are consistent with experimental data. Nat. Biotechnol. 19, 125–130 (2001).
Zwaal, R.R., Broeks, A., van Meurs, J., Groenen, J.T. & Plasterk, R.H. Target-selected gene inactivation in Caenorhabditis elegans by using a frozen transposon insertion mutant bank. Proc. Natl. Acad. Sci. USA 90, 7431–7435 (1993).
Giaever, G. et al. Genomic profiling of drug sensitivities via induced haploinsufficiency. Nat. Genet. 21, 278–283 (1999).
Hicks, G.G. et al. Functional genomics in mice by tagged sequence mutagenesis. Nat. Genet. 16, 338–344 (1997).
Zambrowicz, B.P. et al. Disruption and sequence identification of 2,000 genes in mouse embryonic stem cells. Nature 392, 608–611 (1998).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular cloning: a laboratory manual. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY; 1989).
Ausubel, F.M. et al. Current protocols in molecular biology. (Wiley Interscience, New York, NY; 1994).
Marton, M.J. et al. Drug target validation and identification of secondary drug target effects using DNA microarrays. Nat. Med. 4, 1293–1301 (1998).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W., & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Livesey, F.J., Furukawa, T., Steffen, M.A., Church, G.M. & Cepko, C.L. Microarray analysis of the transcriptional network controlled by the photoreceptor homeobox gene Crx. Curr. Biol. 10, 301–310 (2000).
Tavazoie, S., Hughes, J.D., Campbell, M.J., Cho, R.J. & Church, G.M. Systematic determination of genetic network architecture. Nat. Genet. 22, 281–285 (1999).
Acknowledgements
We would like to thank Martin Steffen and Xiaohua Huang for their tireless efforts to produce the E. coli microarrays, Barak Cohen and Tzachi Pilpel for helpful discussion, and members of the Church lab for their support and encouragement. We would also like to thank the referees for their insightful comments. This work was supported by grants from the US Department of Energy and the National Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
Badarinarayana, V., Estep, P., Shendure, J. et al. Selection analyses of insertional mutants using subgenic-resolution arrays. Nat Biotechnol 19, 1060–1065 (2001). https://doi.org/10.1038/nbt1101-1060
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt1101-1060
This article is cited by
-
Construction of high-density transposon mutant library of Staphylococcus aureus using bacteriophage ϕ11
Journal of Microbiology (2022)
-
Transposon insertion site sequencing (TIS) of Pseudomonas aeruginosa
Journal of Microbiology (2021)
-
Biosensor-guided improvements in salicylate production by recombinant Escherichia coli
Microbial Cell Factories (2019)
-
Combining Shigella Tn-seq data with gold-standard E. coli gene deletion data suggests rare transitions between essential and non-essential gene functionality
BMC Microbiology (2016)
-
Transcriptomic Analysis of 3-Hydroxypropanoic Acid Stress in Escherichia coli
Applied Biochemistry and Biotechnology (2016)