Abstract
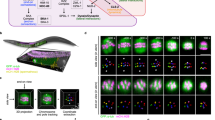
Two distinct chromosome architectures are prevalent among eukaryotes: monocentric, in which localized centromeres restrict kinetochore assembly to a single chromosomal site, and holocentric, in which diffuse kinetochores form along the entire chromosome length. During mitosis, both chromosome types use specialized chromatin, containing the histone H3 variant CENP-A1,2,3, to direct kinetochore assembly4,5,6,7. For the segregation of recombined homologous chromosomes during meiosis8,9, monocentricity is thought to be crucial for limiting spindle-based forces to one side of a crossover and to prevent recombined chromatids from being simultaneously pulled towards both spindle poles. The mechanisms that allow holocentric chromosomes to avert this fate remain uncharacterized. Here, we show that markedly different mechanisms segregate holocentric chromosomes during meiosis and mitosis in the nematode Caenorhabditis elegans. Immediately prior to oocyte meiotic segregation, outer-kinetochore proteins were recruited to cup-like structures on the chromosome surface via a mechanism that is independent of CENP-A. In striking contrast to mitosis, both oocyte meiotic divisions proceeded normally following depletion of either CENP-A or the closely associated centromeric protein CENP-C. These findings highlight a pronounced difference between the segregation of holocentric chromosomes during meiosis and mitosis and demonstrate the potential to uncouple assembly of outer-kinetochore proteins from CENP-A chromatin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Amor, D. J., Kalitsis, P., Sumer, H. & Choo, K. H. Building the centromere: from foundation proteins to 3D organization. Trends Cell Biol. 14, 359–368 (2004).
Mellone, B. G. & Allshire, R. C. Stretching it: putting the CEN(P-A) in centromere. Curr. Opin. Genet. Dev. 13, 191–198 (2003).
Henikoff, S., Furuyama, T. & Ahmad, K. Histone variants, nucleosome assembly and epigenetic inheritance. Trends Genet. 20, 320–326 (2004).
Howman, E. V. et al. Early disruption of centromeric chromatin organization in centromere protein A (Cenpa) null mice. Proc. Natl Acad. Sci. USA 97, 1148–1153 (2000).
Blower, M. D. & Karpen, G. H. The role of Drosophila CID in kinetochore formation, cell-cycle progression and heterochromatin interactions. Nature Cell Biol. 3, 730–739 (2001).
Buchwitz, B. J., Ahmad, K., Moore, L. L., Roth, M. B. & Henikoff, S. A histone-H3-like protein in C. elegans. Nature 401, 547–548 (1999).
Oegema, K., Desai, A., Rybina, S., Kirkham, M. & Hyman, A. A. Functional analysis of kinetochore assembly in Caenorhabditis elegans. J. Cell Biol. 153, 1209–1226 (2001).
Hauf, S. & Watanabe, Y. Kinetochore orientation in mitosis and meiosis. Cell 119, 317–327 (2004).
Petronczki, M., Siomos, M. F. & Nasmyth, K. Un menage a quatre: the molecular biology of chromosome segregation in meiosis. Cell 112, 423–440 (2003).
Pimpinelli, S. & Goday, C. Unusual kinetochores and chromatin diminution in Parascaris. Trends Genet. 5, 310–315 (1989).
Maddox, P. S., Oegema, K., Desai, A. & Cheeseman, I. M. “Holo”er than thou: chromosome segregation and kinetochore function in C. elegans. Chromosome Res. 12, 641–653 (2004).
Moore, L. L. & Roth, M. B. HCP-4, a CENP-C-like protein in Caenorhabditis elegans, is required for resolution of sister centromeres. J. Cell Biol. 153, 1199–1208 (2001).
Howe, M., McDonald, K. L., Albertson, D. G. & Meyer, B. J. HIM-10 is required for kinetochore structure and function on Caenorhabditis elegans holocentric chromosomes. J. Cell Biol. 153, 1227–1238 (2001).
Cheeseman, I. M. et al. A conserved protein network controls assembly of the outer kinetochore and its ability to sustain tension. Genes Dev. 18, 2255–2268 (2004).
Albertson, D. G. & Thomson, J. N. The kinetochores of Caenorhabditis elegans. Chromosoma 86, 409–428 (1982).
O'Toole, E. T. et al. Morphologically distinct microtubule ends in the mitotic centrosome of Caenorhabditis elegans. J. Cell Biol. 163, 451–456 (2003).
Albertson, D. G. & Thomson, J. N. Segregation of holocentric chromosomes at meiosis in the nematode, Caenorhabditis elegans. Chromosome Res. 1, 15–26 (1993).
Comings, D. E. & Okada, T. A. Holocentric chromosomes in Oncopeltus: kinetochore plates are present in mitosis but absent in meiosis. Chromosoma 37, 177–192 (1972).
Desai, A. et al. KNL-1 directs assembly of the microtubule-binding interface of the kinetochore in C. elegans. Genes Dev. 17, 2421–2435 (2003).
Stein, L. D. et al. The genome sequence of Caenorhabditis briggsae: a platform for comparative genomics. PLoS Biol 1, E45 (2003).
Hubbard, E. J. & Greenstein, D. The Caenorhabditis elegans gonad: a test tube for cell and developmental biology. Dev. Dyn. 218, 2–22 (2000).
MacQueen, A. J., Colaiacovo, M. P., McDonald, K. & Villeneuve, A. M. Synapsis-dependent and -independent mechanisms stabilize homolog pairing during meiotic prophase in C. elegans. Genes Dev. 16, 2428–2442 (2002).
Nabeshima, K., Villeneuve, A. M. & Colaiacovo, M. P. Crossing over is coupled to late meiotic prophase bivalent differentiation through asymmetric disassembly of the SC. J. Cell Biol. 168, 683–689 (2005).
Moore, L. L., Morrison, M. & Roth, M. B. HCP-1, a protein involved in chromosome segregation, is localized to the centromere of mitotic chromosomes in Caenorhabditis elegans. J. Cell Biol. 147, 471–480 (1999).
Rogers, E., Bishop, J. D., Waddle, J. A., Schumacher, J. M. & Lin, R. The aurora kinase AIR-2 functions in the release of chromosome cohesion in Caenorhabditis elegans meiosis. J. Cell Biol. 157, 219–229 (2002).
Bernard, P., Maure, J. F. & Javerzat, J. P. Fission yeast Bub1 is essential in setting up the meiotic pattern of chromosome segregation. Nature Cell Biol. 3, 522–526 (2001).
Kitajima, T. S., Kawashima, S. A. & Watanabe, Y. The conserved kinetochore protein shugoshin protects centromeric cohesion during meiosis. Nature 427, 510–517 (2004).
Oegema, K. & Hyman, A. in Wormbook (ed. The C. elegans Research Community). http://www.wormbook.org (2005).
Fukushige, T., Hendzel, M. J., Bazett-Jones, D. P. & McGhee, J. D. Direct visualization of the elt-2 gut-specific GATA factor binding to a target promoter inside the living Caenorhabditis elegans embryo. Proc. Natl Acad. Sci. USA 96, 11883–11888 (1999).
Hillers, K. J. & Villeneuve, A. M. Chromosome-wide control of meiotic crossing over in C. elegans. Curr. Biol. 13, 1641–1647 (2003).
Chan, R. C., Severson, A. F. & Meyer, B. J. Condensin restructures chromosomes in preparation for meiotic divisions. J. Cell Biol. 167, 613–625 (2004).
Edgar, L. G. Blastomere culture and analysis. Methods Cell Biol. 48, 303–321 (1995).
Laiken, S. L., Gross, C. A. & Von Hippel, P. H. Equilibrium and kinetic studies of Escherichia coli lac repressor-inducer interactions. J. Mol. Biol. 66, 143–155 (1972).
Acknowledgements
We thank J. McGhee for strain JM93 and members of the Oegema and Desai laboratories for support and discussions. J.M. is supported by the University of California, San Diego Genetics Training Grant; P.S.M. is the Fayez Sarofim Fellow of the Damon Runyon Cancer Research; K.O. is a Pew Scholar in the Biomedical Sciences; A.D. is the Connie and Bob Lurie Scholar of the Damon Runyon Cancer Research Foundation. This work was supported, in part, by a grant from the National Institutes of Health to A.D. (R01GM074215-01); A.D. and K.O. also receive salary and additional support from the Ludwig Institute for Cancer Research. A.D. and K.O. made the preliminary observations that led to this study. P.S.M. helped to generate and characterize the GFP–CPAR-1 strain. F.H. analysed the relative expression of CENP-A-like proteins. J.M. performed all of the other experiments, and J.M., K.O. and A.D. jointly prepared the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information Guide plus figures S1, S2, S3, S4 and S5 (PDF 591 kb)
Supplementary Information
Supplementary Movie S1 (MOV 598 kb)
Supplementary Information
Supplementary Movie S2 (MOV 1645 kb)
Supplementary Information
Supplementary Movie S3 (MOV 900 kb)
Supplementary Information
Supplementary Movie S4 (MOV 1380 kb)
Supplementary Information
Supplementary Movie S5 (MOV 604 kb)
Rights and permissions
About this article
Cite this article
Monen, J., Maddox, P., Hyndman, F. et al. Differential role of CENP-A in the segregation of holocentric C. elegans chromosomes during meiosis and mitosis. Nat Cell Biol 7, 1248–1255 (2005). https://doi.org/10.1038/ncb1331
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1331
This article is cited by
-
Kinetochore component function in C. elegans oocytes revealed by 4D tracking of holocentric chromosomes
Nature Communications (2023)
-
Chromosomal evolution in Cryptangieae Benth. (Cyperaceae): Evidence of holocentrism and pseudomonads
Protoplasma (2023)
-
Epigenetic, genetic and maternal effects enable stable centromere inheritance
Nature Cell Biology (2022)
-
Meiotic regulation of the Ndc80 complex composition and function
Current Genetics (2021)
-
Unequal contribution of two paralogous CENH3 variants in cowpea centromere function
Communications Biology (2020)