Abstract
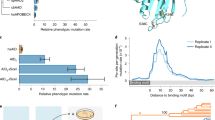
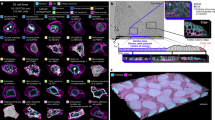
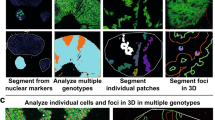
The way in which cells adopt different morphologies is not fully understood. Cell shape could be a continuous variable or restricted to a set of discrete forms. We developed quantitative methods to describe cell shape and show that Drosophila haemocytes in culture are a heterogeneous mixture of five discrete morphologies. In an RNAi screen of genes affecting the morphological complexity of heterogeneous cell populations, we found that most genes regulate the transition between discrete shapes rather than generating new morphologies. In particular, we identified a subset of genes, including the tumour suppressor PTEN, that decrease the heterogeneity of the population, leading to populations enriched in rounded or elongated forms. We show that these genes have a highly conserved function as regulators of cell shape in both mouse and human metastatic melanoma cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Thiery, J.P., Acloque, H., Huang, R. Y. & Nieto, M. A. Epithelial-mesenchymal transitions in development and disease. Cell 139, 871–890 (2009).
Keren, K. et al. Mechanism of shape determination in motile cells. Nature 453, 475–480 (2008).
Mogilner, A. & Keren, K. The shape of motile cells. Curr. Biol. 19, R762–R771 (2009).
Guarino, M., Rubino, B. & Ballabio, G. The role of epithelial-mesenchymal transition in cancer pathology. Pathology 39, 305–318 (2007).
Sanz-Moreno, V. et al. Rac activation and inactivation control plasticity of tumour cell movement. Cell 135, 510–523 (2008).
Wolf, K. et al. Compensation mechanism in tumour cell migration: mesenchymal-amoeboid transition after blocking of pericellular proteolysis. J. Cell Biol. 160, 267–277 (2003).
Bakal, C., Aach, J., Church, G. & Perrimon, N. Quantitative morphological signatures define local signalling networks regulating cell morphology. Science 316, 1753–1756 (2007).
Yin, Z. et al. Using iterative cluster merging with improved gap statistics to perform online phenotype discovery in the context of high-throughput RNAi screens. BMC Bioinformatics 9, 264 (2008).
Waddington, C. H. The Strategy of Genes (Allen Unwin, 1957).
Gadea, G., Sanz-Moreno, V., Self, A., Godi, A. & Marshall, C. J. DOCK10-mediated Cdc42 activation is necessary for amoeboid invasion of melanoma cells. Curr. Biol. 18, 1456–1465 (2008).
Sahai, E. & Marshall, C. J. Differing modes of tumour cell invasion have distinct requirements for Rho/ROCK signalling and extracellular proteolysis. Nat. Cell Biol. 5, 711–719 (2003).
Sanz-Moreno, V. et al. ROCK and JAK1 signalling cooperate to controlactomyosin contractility in tumour cells and stroma. Cancer Cell 20, 229–245 (2011).
Tan, C., Stronach, B. & Perrimon, N. Roles of myosin phosphatase during Drosophila development. Development 130, 671–681 (2003).
Viros, A. et al. Improving melanoma classification by integrating genetic and morphologic features. PLoS Med. 5, e120 (2008).
Sanz-Moreno, V. & Marshall, C. J. The plasticity of cytoskeletal dynamics underlying neoplastic cell migration. Curr. Opin. Cell Biol. 22, 690–696 (2010).
Ridley, A. J. et al. Cell migration: integrating signals from front to back. Science 302, 1704–1709 (2003).
Waddington, C. H. Canalization of development and genetic assimilation of acquired characters. Nature 183, 1654–1655 (1959).
Kauffman, S. A. The Origins of Order. Self-Organization and Selection in Evolution (Oxford Univ. Press, 1993).
Dhomen, N. et al. Oncogenic Braf induces melanocyte senescence and melanoma in mice. Cancer Cell 15, 294–303 (2009).
Dankort, D. et al. Braf(V600E) cooperates with Pten loss to induce metastatic melanoma. Nat. Genet. 41, 544–552 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Acknowledgements
We are indebted to the Drosophila RNAi Screening Center staff at Harvard Medical School for their invaluable assistance. We especially thank I. Flockhart for assistance with data management. We thank J. Wang, X. Zhou and P. Bradley for their initial involvement in this study. We are grateful to N. Dhomen and R. Marais for melanoma cell lines. Work was financially supported in part by NCI grants (Grants R01CA121225 and U54CA149196) to S.T.C.W. and CRUK grants to C.B. (Grant 13478) and C.J.M. (Grant C107/A10433). N.P. is an Investigator of the Howard Hughes Medical Institute. A.S. is a Marie Curie Intra-European Fellow. C.J.M. is a Gibb Life Fellow of CRUK. C.B. is a Research Career Development Fellow of the Wellcome Trust.
Author information
Authors and Affiliations
Contributions
Z.Y. performed the bulk of statistical analysis of RNAi screening data and wrote the Supplementary Note. A.S. designed and performed all RNAi and cell line characterization experiments in mouse and human melanoma cells and contributed to writing of the manuscript. H.S. performed the analysis of live-cell melanoma cell imaging experiments and contributed to visualization of statistical results. A.M. performed all mouse work. X.X. and F.L. performed processing of images generated in Drosophila RNAi screen. M.A.G. and L.E. performed experiments describing penetrance of effects of different dsRNAs. A.R.B. contributed to writing and editing of the manuscript and the Supplementary Note. N.P. participated in the initial design of the study. S.T.C.W. coordinated image processing and statistical analysis. C.J.M. participated in design of melanoma experiments and contributed to writing the manuscript. C.B. participated in the design of experiments and statistical analysis, performed the Drosophila RNAi screen, performed the live-cell imaging assays, coordinated experimental and computational analysis, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1609 kb)
Supplementary Table 1
Supplementary Information (XLSX 10 kb)
Supplementary Table 2
Supplementary Information (XLSX 8 kb)
Supplementary Table 3
Supplementary Information (XLSX 107 kb)
Supplementary Table 4
Supplementary Information (XLSX 8 kb)
Supplementary Table 5
Supplementary Information (XLSX 146 kb)
Supplementary Table 6
Supplementary Information (XLSX 10 kb)
Supplementary Table 7
Supplementary Information (XLSX 31 kb)
Supplementary Table 8
Supplementary Information (XLSX 15 kb)
Rights and permissions
About this article
Cite this article
Yin, Z., Sadok, A., Sailem, H. et al. A screen for morphological complexity identifies regulators of switch-like transitions between discrete cell shapes. Nat Cell Biol 15, 860–871 (2013). https://doi.org/10.1038/ncb2764
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2764
This article is cited by
-
Self-supervised deep learning encodes high-resolution features of protein subcellular localization
Nature Methods (2022)
-
Self-supervised classification of subcellular morphometric phenotypes reveals extracellular matrix-specific morphological responses
Scientific Reports (2022)
-
Traject3d allows label-free identification of distinct co-occurring phenotypes within 3D culture by live imaging
Nature Communications (2022)
-
Morphodynamics facilitate cancer cells to navigate 3D extracellular matrix
Scientific Reports (2021)
-
Image-based profiling for drug discovery: due for a machine-learning upgrade?
Nature Reviews Drug Discovery (2021)