Abstract
In budding yeast, chromatin mobility increases after a DNA double-strand break (DSB). This increase is dependent on Mec1, the yeast ATR kinase, but the targets responsible for this phenomenon are unknown. Here we report that the Mec1-dependent phosphorylation of Cep3, a kinetochore component, is required to stimulate chromatin mobility after DNA breaks. Cep3 phosphorylation counteracts a constraint on chromosome movement imposed by the attachment of centromeres to the spindle pole body. A second constraint, imposed by the tethering of telomeres to the nuclear periphery, is also relieved after chromosome breakage. A non-phosphorylatable Cep3 mutant that impairs DSB-induced chromatin mobility is proficient in DSB repair, suggesting that break-induced chromatin mobility may be dispensable for homology search. Rather, we propose that the relief of centromeric constraint promotes cell cycle arrest and faithful chromosome segregation through the engagement of the spindle assembly checkpoint.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Marshall, W. F. et al. Interphase chromosomes undergo constrained diffusional motion in living cells. Curr. Biol. 7, 930–939 (1997).
Chubb, J. R., Boyle, S., Perry, P. & Bickmore, W. A. Chromatin motion is constrained by association with nuclear compartments in human cells. Curr. Biol. 12, 439–445 (2002).
Vazquez, J., Belmont, A. S. & Sedat, J. W. Multiple regimes of constrained chromosome motion are regulated in the interphase Drosophila nucleus. Curr. Biol. 11, 1227–1239 (2001).
Heun, P., Laroche, T., Shimada, K., Furrer, P. & Gasser, S. M. Chromosome dynamics in the yeast interphase nucleus. Science 294, 2181–2186 (2001).
Seeber, A., Dion, V. & Gasser, S. M. Checkpoint kinases and the INO80 nucleosome remodeling complex enhance global chromatin mobility in response to DNA damage. Genes Dev. 27, 1999–2008 (2013).
Chung, D. K. et al. Perinuclear tethers license telomeric DSBs for a broad kinesin- and NPC-dependent DNA repair process. Nat. Commun. 6, 7742 (2015).
Dion, V. & Gasser, S. M. Chromatin movement in the maintenance of genome stability. Cell 152, 1355–1364 (2013).
Krawczyk, P. M. et al. Chromatin mobility is increased at sites of DNA double-strand breaks. J. Cell Sci. 125, 2127–2133 (2012).
Lottersberger, F., Karssemeijer, R. A., Dimitrova, N. & de Lange, T. 53BP1 and the LINC complex promote microtubule-dependent DSB mobility and DNA repair. Cell 163, 880–893 (2015).
Dimitrova, N., Chen, Y. C., Spector, D. L. & de Lange, T. 53BP1 promotes non-homologous end joining of telomeres by increasing chromatin mobility. Nature 456, 524–528 (2008).
Cho, N. W., Dilley, R. L., Lampson, M. A. & Greenberg, R. A. Interchromosomal homology searches drive directional ALT telomere movement and synapsis. Cell 159, 108–121 (2014).
Kruhlak, M. J. et al. Changes in chromatin structure and mobility in living cells at sites of DNA double-strand breaks. J. Cell Biol. 172, 823–834 (2006).
Soutoglou, E. et al. Positional stability of single double-strand breaks in mammalian cells. Nat. Cell Biol. 9, 675–682 (2007).
Roukos, V. et al. Spatial dynamics of chromosome translocations in living cells. Science 341, 660–664 (2013).
Dion, V., Kalck, V., Horigome, C., Towbin, B. D. & Gasser, S. M. Increased mobility of double-strand breaks requires Mec1, Rad9 and the homologous recombination machinery. Nat. Cell Biol. 14, 502–509 (2012).
Mine-Hattab, J. & Rothstein, R. Increased chromosome mobility facilitates homology search during recombination. Nat. Cell Biol. 14, 510–517 (2012).
Aylon, Y. & Kupiec, M. DSB repair: the yeast paradigm. DNA Repair 3, 797–815 (2004).
Renkawitz, J., Lademann, C. A. & Jentsch, S. Mechanisms and principles of homology search during recombination. Nat. Rev. Mol. Cell Biol. 15, 369–383 (2014).
Horigome, C. et al. SWR1 and INO80 chromatin remodelers contribute to DNA double-strand break perinuclear anchorage site choice. Mol. Cell 55, 626–639 (2014).
Gasser, S. M. Visualizing chromatin dynamics in interphase nuclei. Science 296, 1412–1416 (2002).
Verdaasdonk, J. S. et al. Centromere tethering confines chromosome domains. Mol. Cell 52, 819–831 (2013).
Hediger, F., Neumann, F. R., Van Houwe, G., Dubrana, K. & Gasser, S. M. Live imaging of telomeres: yKu and Sir proteins define redundant telomere-anchoring pathways in yeast. Curr. Biol. 12, 2076–2089 (2002).
Gartenberg, M. R., Neumann, F. R., Laroche, T., Blaszczyk, M. & Gasser, S. M. Sir-mediated repression can occur independently of chromosomal and subnuclear contexts. Cell 119, 955–967 (2004).
Hill, A. & Bloom, K. Genetic manipulation of centromere function. Mol. Cell. Biol. 7, 2397–2405 (1987).
Smolka, M. B., Albuquerque, C. P., Chen, S. H. & Zhou, H. Proteome-wide identification of in vivo targets of DNA damage checkpoint kinases. Proc. Natl Acad. Sci. USA 104, 10364–10369 (2007).
Chen, S. H., Albuquerque, C. P., Liang, J., Suhandynata, R. T. & Zhou, H. A proteome-wide analysis of kinase-substrate network in the DNA damage response. J. Biol. Chem. 285, 12803–12812 (2010).
Lisby, M., Rothstein, R. & Mortensen, U. H. Rad52 forms DNA repair and recombination centers during S phase. Proc. Natl Acad. Sci. USA 98, 8276–8282 (2001).
Lisby, M., Mortensen, U. H. & Rothstein, R. Colocalization of multiple DNA double-strand breaks at a single Rad52 repair centre. Nat. Cell Biol. 5, 572–577 (2003).
Lechner, J. & Carbon, J. A 240 kd multisubunit protein complex, CBF3, is a major component of the budding yeast centromere. Cell 64, 717–725 (1991).
Lechner, J. A zinc finger protein, essential for chromosome segregation, constitutes a putative DNA binding subunit of the Saccharomyces cerevisiae kinetochore complex, Cbf3. EMBO J. 13, 5203–5211 (1994).
Purvis, A. & Singleton, M. R. Insights into kinetochore–DNA interactions from the structure of Cep3Delta. EMBO Rep. 9, 56–62 (2008).
Pinsky, B. A., Kung, C., Shokat, K. M. & Biggins, S. The Ipl1-Aurora protein kinase activates the spindle checkpoint by creating unattached kinetochores. Nat. Cell Biol. 8, 78–83 (2006).
Dorn, J. F. et al. Yeast kinetochore microtubule dynamics analyzed by high-resolution three-dimensional microscopy. Biophys. J. 89, 2835–2854 (2005).
Jeggo, P. A. & Downs, J. A. Roles of chromatin remodellers in DNA double strand break repair. Exp. Cell Res. 329, 69–77 (2014).
Agmon, N., Liefshitz, B., Zimmer, C., Fabre, E. & Kupiec, M. Effect of nuclear architecture on the efficiency of double-strand break repair. Nat. Cell Biol. 15, 694–699 (2013).
Rieder, C. L., Cole, R. W., Khodjakov, A. & Sluder, G. The checkpoint delaying anaphase in response to chromosome monoorientation is mediated by an inhibitory signal produced by unattached kinetochores. J. Cell Biol. 130, 941–948 (1995).
Sanchez, Y. et al. Control of the DNA damage checkpoint by chk1 and rad53 protein kinases through distinct mechanisms. Science 286, 1166–1171 (1999).
Wang, H., Liu, D., Wang, Y., Qin, J. & Elledge, S. J. Pds1 phosphorylation in response to DNA damage is essential for its DNA damage checkpoint function. Genes Dev. 15, 1361–1372 (2001).
Dotiwala, F., Harrison, J. C., Jain, S., Sugawara, N. & Haber, J. E. Mad2 prolongs DNA damage checkpoint arrest caused by a double-strand break via a centromere-dependent mechanism. Curr. Biol. 20, 328–332 (2010).
Musacchio, A. & Salmon, E. D. The spindle-assembly checkpoint in space and time. Nat. Rev. Mol. Cell Biol. 8, 379–393 (2007).
Yuen, K. W. et al. Systematic genome instability screens in yeast and their potential relevance to cancer. Proc. Natl Acad. Sci. USA 104, 3925–3930 (2007).
Warren, C. D. et al. S-phase checkpoint genes safeguard high-fidelity sister chromatid cohesion. Mol. Biol. Cell 15, 1724–1735 (2004).
Renkawitz, J., Lademann, C. A., Kalocsay, M. & Jentsch, S. Monitoring homology search during DNA double-strand break repair in vivo. Mol. Cell 50, 261–272 (2013).
Neumann, F. R. et al. Targeted INO80 enhances subnuclear chromatin movement and ectopic homologous recombination. Genes Dev. 26, 369–383 (2012).
Chambers, A. L. et al. The INO80 chromatin remodeling complex prevents polyploidy and maintains normal chromatin structure at centromeres. Genes Dev. 26, 2590–2603 (2012).
Lee, C.-S. et al. Chromosome position determines the success of double-strand break repair. Proc. Natl Acad. Sci. USA 113, E146–E154 (2015).
Dick, A. E. & Gerlich, D. W. Kinetic framework of spindle assembly checkpoint signalling. Nat. Cell Biol. 15, 1370–1377 (2013).
Collin, P., Nashchekina, O., Walker, R. & Pines, J. The spindle assembly checkpoint works like a rheostat rather than a toggle switch. Nat. Cell Biol. 15, 1378–1385 (2013).
Rohner, S., Gasser, S. M. & Meister, P. Modules for cloning-free chromatin tagging in Saccharomyces cerevisae. Yeast 25, 235–239 (2008).
Sage, D., Neumann, F. R., Hediger, F., Gasser, S. M. & Unser, M. Automatic tracking of individual fluorescence particles: application to the study of chromosome dynamics. IEEE Trans. Image Process. 14, 1372–1383 (2005).
Wybenga-Groot, L. E. et al. Structural basis of Rad53 kinase activation by dimerization and activation segment exchange. Cell Signal. 26, 1825–1836 (2014).
Acknowledgements
We are grateful to R. Szilard and members of the Durocher laboratory for critical reading of the manuscript. We also thank S. Jaspersen (Stowers Institute for Medical Research, USA), P. Hieter (University of British Columbia, Canada), G. Brown (University of Toronto, Canada), M. Meneghini (University of Toronto, Canada), R. Rothstein (Columbia University, USA), S. Gasser (Friedrich Miescher Institute, Switzerland), S. Biggins (Fred Hutchinson Cancer Research Center, USA) and M. Kupiec (Tel Aviv University, Israel) for sharing strains and plasmids. J.S. is supported by a CIHR Doctoral award. D.D. is the Thomas Kierans Chair in Mechanisms of Cancer Development and a Canada Research Chair (Tier 1) in the Molecular Mechanisms of Genome Integrity. This work was supported by CIHR grant FDN143343 (to D.D.) and MOP123468 (to L.P.) and a Grant-in-Aid from the Krembil Foundation to L.P. and D.D.
Author information
Authors and Affiliations
Contributions
J.S. carried out the experiments and wrote the manuscript. G.D.G. and M.B. generated MATLAB scripts for nuclear alignment and MSD analysis. W.Z. and M.-C.L. helped initiate the project and set up the microscopy system for measuring chromatin mobility. L.P. supervised G.D.G. and M.B. and provided microscopy advice. D.D. supervised the project and wrote the manuscript with J.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 3 System for chromatin mobility analysis in budding yeast.
(a) Representative images of mobility strains. Chromatin is tracked by integrating LacOx256 arrays in cells expressing LacI-GFP and Nup49-mCherry. Scale bar represents 2 μm. (b) X-axis displacement of a single tracked MAT locus over time in 1 × 1 or 2 × 2 binned images. (c) Growth comparison of unexposed and imaged cells. Cells were held in a DeltaVision microscope at 30 °C in SD media supplemented with raffinose and imaged at the indicated time points. Scale bar represents 4 μm. (d) MSD analysis of the MAT locus in wild-type cells before and 3 h after DSB induction. (e) MSD analysis of the MAT locus in wild-type cells or in the INO80 mutant arp8Δ. All MSD data represent the mean ± s.e.m. of cells pooled from 3 independent experiments. See Supplementary Table 1 for mobility parameters and the precise number of cells analysed.
Supplementary Figure 4 Cep3-S575 is required for DSB-induced chromatin mobility.
(a,b) Radius of confinement (a) and diffusion coefficients (b) of wild-type and cep3-S575A cells following a DSB. Data represents the mean ± s.e.m., n represent the number of cells pooled from 3 independent experiments (WT control n = 131, WT DSB n = 81, cep3-S575A control n = 189, cep3-S575A DSB n = 111). ∗∗∗P < 0.001; one-way ANOVA. (c) MSD analysis of the MAT locus in wild-type cells and a cep3-S575A mutant generated at the endogenous CEP3 locus by the pop-in/pop-out method. (d) MSD analysis of the MAT locus in wild-type or cep3-S575A/cep3-S575A diploid cells containing an HO cut site on one homolog. All MSD data represent the mean ± s.e.m. of cells pooled from 3 independent experiments. See Supplementary Table 1 for mobility parameters and the precise number of cells analysed.
Supplementary Figure 5 Cep3 phosphorylation requires Mec1/Rad53/Dun1 signalling.
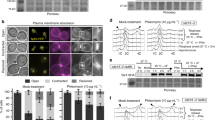
(a) Immunoblot analysis using a Cep3-S575 phosphospecific antibody on whole cell extracts from cells expressing Myc-tagged Cep3 (WT or S575A) in response to a HO-induced DSB. Rad53 was probed to confirm checkpoint activation, Myc was used to control for loading. (b) Immunoblot analysis of asynchronous (ASN), α-factor arrested (G1, or nocodazole arrested (G2/M) cells treated with Zeocin (250 μg ml−1) for 1 h. Levels of the mitotic cyclin Clb2 were ascertained to confirm cell cycle synchronization. Pgk1 and Rad53 were probed to control for loading and to confirm checkpoint activation respectively. (c) MSD analysis of the MAT locus in sml1Δ tel1Δ mutants. (d) MSD analysis of the MAT locus in sml1Δ tel1Δ mec1Δ mutants. (e) MSD analysis of the MAT locus in cep3-S575A rad53Δ mutants. (f) Amino acid sequence surrounding Cep3-S575 and the consensus phosphorylation target sites of the Rad53 and Dun1 kinases. Matching residues are highlighted in red, Ψ represents aliphatic amino acids. (g) Time-course of Cep3-13xMyc phosphorylation in response to Zeocin (250 μg ml−1) determined by immunoblot analysis. (h) MSD analysis of the MAT locus in dun1Δ mutants. (i) Cep3-S575 phosphorylation in dun1Δ cells following 1 h of Zeocin treatment (250 μg ml−1) determined by immunoblot analysis.
Supplementary Figure 6 Mobility of a CEN-bound episome following DSB induction.
(a,b) MSD analysis of a CEN/ARS-LacOx256 episome in wild-type (a) and cep3-S575A (b) cells in response to a DSB 27 kb from CEN4. (c) MSD analysis of a CEN/ARS-LacOx256 episome in response to a DSB 998 kb from CEN4. All MSD data represent the mean ± s.e.m. of cells pooled from 3 independent experiments. See Supplementary Table 1 for mobility parameters and the precise number of cells analysed.
Supplementary Figure 7 Loss of telomeric tethering does not increase the mobility of a broken chromosome arm.
(a) MSD analysis of the MAT locus on Chr III (in cis) and the MAK10 locus on unbroken Chr V in trans to a DSB 27 kb from CEN4. (b) MSD analysis of the MAK10 locus on unbroken Chr V in trans to a DSB 27 kb from CEN4 in sir4Δ cells. (c,d,e) MSD analysis of the MAT locus, in cis to a DSB, in sir4Δ (c), yku70Δ (d), and yku70Δ sir4Δ (e) cells. All MSD data represent the mean ± s.e.m. of cells pooled from 3 independent experiments. See Supplementary Table 1 for mobility parameters and the precise number of cells analysed.
Supplementary Figure 8 Mobility of a broken and unbroken chromosome arm.
(a) Schematic of strains to follow the mobility of both chromosome arms following an HO-induced DSB on Chr IV (red triangle). (b) MSD analysis of the Chr IV-R and Chr IV-L arms before and after a DSB. (c) MSD analysis of the uncut Chr IV-L arm in wild-type and cep3-S575A cells. (d) MSD analysis of the cut Chr IV-R arm and the uncut Chr IV-L arm after a DSB in a sir4Δ background. All MSD data represent the mean ± s.e.m. of cells pooled from 3 independent experiments. See Supplementary Table 1 for mobility parameters and the precise number of cells analysed.
Supplementary Figure 9 Cep3-S75A does not affect the kinetics of DSB repair or DNA binding.
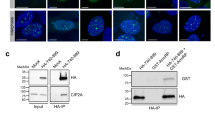
(a) Schematic of the HR repair strain NA60. PCR primers P1 and P2 were used to amplify the homologous recombination cassette containing the HO cut site on Chr IX. Repair of the DSB by HR is accompanied by the appearance of a ClaI site provided by the donor recombination cassette on Chr XIII. (b) Agarose gel of ClaI-digested PCR products in response to a DSB in wild-type and cep3-S575A cells. DNA was extracted from cells harvested at the indicated time points, the cut locus was amplified by PCR, and PCR products were digested with ClaI. Gene conversion (GC) products are cleaved by ClaI and form two lower molecular weight bands. Early repair products can be detected at 2.5 h in both wild-type and cep3-S575A cells. A colony that survived plating on galactose was used as a positive control (+). (c) Chromatin immunoprecipitation (ChIP) of Cep3-13xMyc and Cep3-S575A-13xMyc in response to a DSB at the MAT locus. Fold enrichment at CEN3 is normalized to the non-centromeric control locus TSC11. Data represents the mean ± s.d., n = 5 independent experiments for each condition.
Supplementary Figure 10 Cep3 and INO80 function in a common pathway.
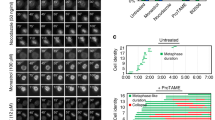
(a,b) MSD analysis of the MAT locus in INO80 mutant arp8Δ cells with CEN3 (a) or CEN5 (b) inactivation. (c) Length of checkpoint arrest in response to an irreparable DSB at the MAT locus. Each dot represents a single cell combined from 2 or more independent experiments for each genotype (WT n = 199 cells, cep3-S575A n = 194, arp8Δ n = 171, cep3-S75A arp8Δ n = 171); red bars represent the median arrest length. (d) Immunoblot analysis of Cep3-S575 phosphorylation in wild-type and cep3-S575Acells, and two clones each of the INO80 mutants arp5Δ and arp8Δ after Zeocin treatment (250 μg ml−1) for 1 h (+) as described in Fig. 3c. All MSD data represent the mean ± s.e.m. of cells pooled from 3 independent experiments. See Supplementary Table 1 for mobility parameters and the precise number of cells analysed.
Supplementary Figure 11 Uncropped scans of immunoblots.
Cropped images for figures are indicated by a red box.
Supplementary information
Supplementary Information
Supplementary Information (PDF 777 kb)
Rights and permissions
About this article
Cite this article
Strecker, J., Gupta, G., Zhang, W. et al. DNA damage signalling targets the kinetochore to promote chromatin mobility. Nat Cell Biol 18, 281–290 (2016). https://doi.org/10.1038/ncb3308
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3308
This article is cited by
-
Fission yeast condensin contributes to interphase chromatin organization and prevents transcription-coupled DNA damage
Genome Biology (2020)
-
Help or hindrance: how do microtubule-based forces contribute to genome damage and repair?
Current Genetics (2020)
-
The Rabl configuration limits topological entanglement of chromosomes in budding yeast
Scientific Reports (2019)
-
DNA double-strand breaks in telophase lead to coalescence between segregated sister chromatid loci
Nature Communications (2019)
-
Chromatin mobility upon DNA damage: state of the art and remaining questions
Current Genetics (2019)