Abstract
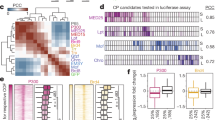
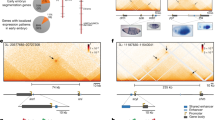
The binding of some transcription factors has been shown to diverge substantially between closely related species. Here we show that the binding of the developmental transcription factor Twist is highly conserved across six Drosophila species, revealing strong functional constraints at its enhancers. Conserved binding correlates with sequence motifs for Twist and its partners, permitting the de novo discovery of their combinatorial binding. It also includes over 10,000 low-occupancy sites near the detection limit, which tend to mark enhancers of later developmental stages. These results suggest that developmental enhancers can be highly evolutionarily constrained, presumably because of their complex combinatorial nature.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Schmidt, D. et al. Five-vertebrate ChIP-seq reveals the evolutionary dynamics of transcription factor binding. Science 328, 1036–1040 (2010).
Borneman, A.R. et al. Divergence of transcription factor binding sites across related yeast species. Science 317, 815–819 (2007).
Odom, D.T. et al. Tissue-specific transcriptional regulation has diverged significantly between human and mouse. Nat. Genet. 39, 730–732 (2007).
Davidson, E.H. & Erwin, D.H. Gene regulatory networks and the evolution of animal body plans. Science 311, 796–800 (2006).
Clark, A.G. et al. Evolution of genes and genomes on the Drosophila phylogeny. Nature 450, 203–218 (2007).
Stark, A. et al. Discovery of functional elements in 12 Drosophila genomes using evolutionary signatures. Nature 450, 219–232 (2007).
Baylies, M.K. & Bate, M. Twist: a myogenic switch in Drosophila. Science 272, 1481–1484 (1996).
Jiang, J., Kosman, D., Ip, Y.T. & Levine, M. The dorsal morphogen gradient regulates the mesoderm determinant twist in early Drosophila embryos. Genes Dev. 5, 1881–1891 (1991).
Leptin, M. Twist and snail as positive and negative regulators during Drosophila mesoderm development. Genes Dev. 5, 1568–1576 (1991).
Sandmann, T. et al. A core transcriptional network for early mesoderm development in Drosophila melanogaster. Genes Dev. 21, 436–449 (2007).
Zeitlinger, J. et al. Whole-genome ChIP-chip analysis of Dorsal, Twist, and Snail suggests integration of diverse patterning processes in the Drosophila embryo. Genes Dev. 21, 385–390 (2007).
Ip, Y.T., Park, R.E., Kosman, D., Yazdanbakhsh, K. & Levine, M. Dorsal-Twist interactions establish snail expression in the presumptive mesoderm of the Drosophila embryo. Genes Dev. 6, 1518–1530 (1992).
Castanon, I. & Baylies, M.K. A Twist in fate: evolutionary comparison of Twist structure and function. Gene 287, 11–22 (2002).
Technau, U. & Scholz, C.B. Origin and evolution of endoderm and mesoderm. Int. J. Dev. Biol. 47, 531–539 (2003).
Zinzen, R.P., Senger, K., Levine, M. & Papatsenko, D. Computational models for neurogenic gene expression in the Drosophila embryo. Curr. Biol. 16, 1358–1365 (2006).
Zeitlinger, J. et al. Program-specific distribution of a transcription factor dependent on partner transcription factor and MAPK signaling. Cell 113, 395–404 (2003).
Sandmann, T. et al. A temporal map of transcription factor activity: mef2 directly regulates target genes at all stages of muscle development. Dev. Cell 10, 797–807 (2006).
Richards, S. et al. Comparative genome sequencing of Drosophila pseudoobscura: chromosomal, gene, and cis-element evolution. Genome Res. 15, 1–18 (2005).
Bradley, R.K. et al. Binding site turnover produces pervasive quantitative changes in transcription factor binding between closely related Drosophila species. PLoS Biol. 8, e1000343 (2010).
Tamura, K., Subramanian, S. & Kumar, S. Temporal patterns of fruit fly (Drosophila) evolution revealed by mutation clocks. Mol. Biol. Evol. 21, 36–44 (2004).
Nègre, N. et al. A comprehensive map of insulator elements for the Drosophila genome. PLoS Genet. 6, e1000814 (2010).
MacArthur, S. et al. Developmental roles of 21 Drosophila transcription factors are determined by quantitative differences in binding to an overlapping set of thousands of genomic regions. Genome Biol. 10, R80 (2009).
Visel, A., Rubin, E.M. & Pennacchio, L.A. Genomic views of distant-acting enhancers. Nature 461, 199–205 (2009).
Bulger, M. & Groudine, M. Enhancers: the abundance and function of regulatory sequences beyond promoters. Dev. Biol. 339, 250–257 (2010).
Degenhardt, K.R. et al. Distinct enhancers at the Pax3 locus can function redundantly to regulate neural tube and neural crest expressions. Dev. Biol. 339, 519–527 (2010).
Perry, M.W., Boettiger, A.N., Bothma, J.P. & Levine, M. Shadow enhancers foster robustness of Drosophila gastrulation. Curr. Biol. 20, 1562–1567 (2010).
Frankel, N. et al. Phenotypic robustness conferred by apparently redundant transcriptional enhancers. Nature 466, 490–493 (2010).
O'Meara, M.M. et al. Cis-regulatory mutations in the Caenorhabditis elegans homeobox gene locus cog-1 affect neuronal development. Genetics 181, 1679–1686 (2009).
Hong, J.W., Hendrix, D.A. & Levine, M.S. Shadow enhancers as a source of evolutionary novelty. Science 321, 1314 (2008).
Zinzen, R.P., Girardot, C., Gagneur, J., Braun, M. & Furlong, E.E. Combinatorial binding predicts spatio-temporal cis-regulatory activity. Nature 462, 65–70 (2009).
García-Zaragoza, E., Mas, J.A., Vivar, J., Arredondo, J.J. & Cervera, M. CF2 activity and enhancer integration are required for proper muscle gene expression in Drosophila. Mech. Dev. 125, 617–630 (2008).
Markstein, M. et al. A regulatory code for neurogenic gene expression in the Drosophila embryo. Development 131, 2387–2394 (2004).
Li, X.Y. et al. Transcription factors bind thousands of active and inactive regions in the Drosophila blastoderm. PLoS Biol. 6, e27 (2008).
Qian, S., Capovilla, M. & Pirrotta, V. Molecular mechanisms of pattern formation by the BRE enhancer of the Ubx gene. EMBO J. 12, 3865–3877 (1993).
Kasowski, M. et al. Variation in transcription factor binding among humans. Science 328, 232–235 (2010).
Zheng, W., Zhao, H., Mancera, E., Steinmetz, L.M. & Snyder, M. Genetic analysis of variation in transcription factor binding in yeast. Nature 464, 1187–1191 (2010).
Wilczynski, B. & Furlong, E.E. Dynamic CRM occupancy reflects a temporal map of developmental progression. Mol. Syst. Biol. 6, 383 (2010).
Conboy, C.M. et al. Cell cycle genes are the evolutionarily conserved targets of the E2F4 transcription factor. PLoS ONE 2, e1061 (2007).
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nat. Genet. 42, 631–634 (2010).
Mikkelsen, T.S. et al. Comparative epigenomic analysis of murine and human adipogenesis. Cell 143, 156–169 (2010).
Carroll, S.B. Evo-devo and an expanding evolutionary synthesis: a genetic theory of morphological evolution. Cell 134, 25–36 (2008).
Davidson, E.H. & Britten, R.J. Regulation of gene expression: possible role of repetitive sequences. Science 204, 1052–1059 (1979).
Arendt, D. The evolution of cell types in animals: emerging principles from molecular studies. Nat. Rev. Genet. 9, 868–882 (2008).
Cande, J., Goltsev, Y. & Levine, M.S. Conservation of enhancer location in divergent insects. Proc. Natl. Acad. Sci. USA 106, 14414–14419 (2009).
Flames, N. & Hobert, O. Gene regulatory logic of dopamine neuron differentiation. Nature 458, 885–889 (2009).
Deato, M.D. & Tjian, R. Switching of the core transcription machinery during myogenesis. Genes Dev. 21, 2137–2149 (2007).
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
Pennacchio, L.A. et al. In vivo enhancer analysis of human conserved non-coding sequences. Nature 444, 499–502 (2006).
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–345 (2005).
Ettwiller, L. et al. The discovery, positioning and verification of a set of transcription-associated motifs in vertebrates. Genome Biol. 6, R104 (2005).
Del Bene, F. et al. In vivo validation of a computationally predicted conserved Ath5 target gene set. PLoS Genet. 3, 1661–1671 (2007).
Yin, Z., Xu, X.L. & Frasch, M. Regulation of the twist target gene tinman by modular cis-regulatory elements during early mesoderm development. Development 124, 4971–4982 (1997).
Kim, J., Kerr, J.Q. & Min, G.S. Molecular heterochrony in the early development of Drosophila. Proc. Natl. Acad. Sci. USA 97, 212–216 (2000).
Rothwell, W.F. & Sullivan, W. Drosophila Protocols. 141 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2000).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Kheradpour, P., Stark, A., Roy, S. & Kellis, M. Reliable prediction of regulator targets using 12 Drosophila genomes. Genome Res. 17, 1919–1931 (2007).
Acknowledgements
We would like to thank E. Furlong (European Molecular Biology Laboratory (EMBL)) and M. Levine (University of California Berkeley) for generously providing Twist antibodies; B. Dickson, W. Lugmayr, R. Revilla (IMP), C. Schlötterer (University of Veterinary Medicine Vienna), M. Levine (University of California Berkeley) and R. Krumlauf (Stowers Institute for Medical Research) for discussions, help and advice. A.F.B. was supported by the Austrian Ministry for Science and Research through the Genome Research in Austria (GEN-AU) Bioinformatics Integration Network III. J.Z. is a Pew scholar. A.S. is supported by an European Research Council (ERC) Starting Grant from the European Community's Seventh Framework Programme (FP7/2007-2013)/ERC grant agreement no. 242922.
Author information
Authors and Affiliations
Contributions
Q.H. performed the ChIP experiments and library preparation, and J.J., A.P., M.G. and J.Z. established the ChIP-Seq pipeline. B.P. and J.P. raised the different Drosophila species, harvested the embryos and staged them, A.F.B. and A.S. analyzed the data, and Q.H., A.F.B., A.S. and J.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–19 and Supplementary Tables 1–6, 8–10 and 14–16. (PDF 2930 kb)
Supplementary Table 7
Conservation of D. melanosgaster Twist binding peaks in six Drosophila species (XLS 390 kb)
Supplementary Table 11
Quantitative changes of Twist binding peaks in all six Drosophila species (XLS 2360 kb)
Supplementary Table 12
GO analysis of invariant vs. variant peaks (XLS 446 kb)
Supplementary Table 13
GO analysis of genes near Twist binding peaks (XLS 588 kb)
Rights and permissions
About this article
Cite this article
He, Q., Bardet, A., Patton, B. et al. High conservation of transcription factor binding and evidence for combinatorial regulation across six Drosophila species. Nat Genet 43, 414–420 (2011). https://doi.org/10.1038/ng.808
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.808
This article is cited by
-
Prediction and analysis of cis-regulatory elements in Dorsal and Ventral patterning genes of Tribolium castaneum and its comparison with Drosophila melanogaster
Molecular and Cellular Biochemistry (2024)
-
Functional annotations of three domestic animal genomes provide vital resources for comparative and agricultural research
Nature Communications (2021)
-
Genome-wide identification of Drosophila dorso-ventral enhancers by differential histone acetylation analysis
Genome Biology (2016)
-
Common binding by redundant group B Sox proteins is evolutionarily conserved in Drosophila
BMC Genomics (2015)
-
ChIP-nexus enables improved detection of in vivo transcription factor binding footprints
Nature Biotechnology (2015)