Abstract
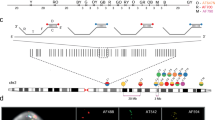
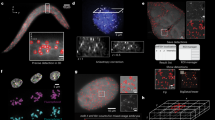
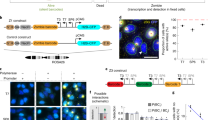
We describe a method for stretching DNA, which, when combined with fluorescent hybridization procedures, forms a new mapping technology that produces a high resolution, vivid, multi–colour image and map. Restriction fragments and cosmid probes were successfully mapped by this procedure with validation by standard restriction mapping. A long range map of a >200 kilobase region containing five copies of the amplified dihydrofolate reductase gene was easily generated within two days. This DNA mapping procedure offers a significant and rapid alternative to a variety of standard mapping procedures.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Green, E.D. & Olson, M.V. Chromosomal region of the cystic fibrosis gene in yeast artificial chromosomes: a model for human genome mapping. Science 250, 94–98 (1990).
Cox, D.R., Burmeister, M., Price, E.R., Kim, S. & Myers, R.M. Radiation hybrid mapping: a somatic cell genetic method for constructing high-resolution maps of mammalian chromosomes. Science 250, 245–250 (1990).
Ballanne-Chantelot, B. et al. Mapping the whole human genome by fingerprinting yeast artificial chromosomes. Cell 70, 1059–1068 (1992).
Lichter, P. et al. High-resolution mapping of human chromosome 11 by in situ hybridization with cosmid clones. Science 247, 64–69 (1990).
Trask, B., Pinkel, D. & van den Engh, G. The proximity of DNA sequences in interphase cell nuclei is correlated to genomic distance and permits ordering of cosmids spanning 250 kilobase pairs. Genomics 5, 710–717 (1989).
Trask, B.J., Massa, H., Kenwrick, S. & Gitschier, J. Mapping of human chromosome Xo28 by two-color fluorescence in situ hybridization of DNA sequences to interphase cell nuclei. Am. J. hum. Genet. 48, 1–15 (1991).
Lawrence, J.B., Singer, R.H. & McNeil, J.A. Interphase and metaphase resolution of different distances within the human dystrophin gene. Science 249, 928–932 (1990).
van den Engh, G., Sachs, R. & Trask, B.J. Estimating genomic distances from DNA sequence location in cell nuclei by a random walk model. Science 257, 1410–1412 (1992).
Heng, H.H.Q., Squire, J. & Tsui, L.-C. High-resolution mapping of mammalian genes by in situ hybridization to free chromatin. Proc. natn. Acad. Sci. U.S.A. 89, 9509–9513 (1992).
Wiegant, J. et al. High-resolution in situ hybridization using DNA halo preparations. Hum. molec. Genet. 1, 587–591 (1992).
Lawrence, J.B., Carter, K.C. & Gerdes, M.J. Extending the capabilities of interphase chromatin mapping. Nature Genet. 2, 171–172 (1992).
Windle, B., Draper, B.W., Yin, Y., O'Gorman, S. & Wahl, G.M. A central role for chromosome breakage in gene amplification, deletion formation, and amplicon integration. Genes Dev. 5, 160–174 (1991).
Evans, G.A. & Wahl, G.M. Cosmid vectors for genomic walking and rapid restriction mapping. Meth. Enzymol. 152, 604–610 (1987).
Ma, C., Looney, J.E., Leu, T.H. & Hamlin, J.L. Organization and genesis of dihydrofolate reductase amplicons in the genome of a methotrexate-resistant Chinese hamster ovary cell line. Molec. cell. Biol. 8, 2316–2327 (1988).
Amler, L.C. & Schwab, M. Amplified N-myc in human neuroblastoma cells is often arranged as clustered tandem repeats of differently recombined DNA. Molec. cell. Biol. 9, 4903–4913 (1989).
Pinkel, D., Straume, T. & Gray, J.W. Cytogenetic analysis using quantitative, high sensitivity, fluorescence hybridization. Proc. natn. Acad. Sci. U.S.A. 83, 2934–2938 (1986).
Urlaub, G. et al. Effect of gamma rays at the dihydrofolate reductase locus: deletions and inversions. Somat. Cell molec. Genet. 12, 555–566 (1986).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Parra, I., Windle, B. High resolution visual mapping of stretched DNA by fluorescent hybridization. Nat Genet 5, 17–21 (1993). https://doi.org/10.1038/ng0993-17
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0993-17
This article is cited by
-
ELOF1 is a transcription-coupled DNA repair factor that directs RNA polymerase II ubiquitylation
Nature Cell Biology (2021)
-
Warsaw Breakage Syndrome associated DDX11 helicase resolves G-quadruplex structures to support sister chromatid cohesion
Nature Communications (2020)
-
Loss of SIM2s inhibits RAD51 binding and leads to unresolved replication stress
Breast Cancer Research (2019)
-
Discrimination of Adsorbed Double-Stranded and Single-Stranded DNA Molecules on Surfaces by Fluorescence Emission Spectroscopy Using Acridine Orange Dye
Journal of Fluorescence (2017)