Abstract
Cold induces expression of a number of genes that encode proteins that enhance tolerance to freezing temperatures in plants1,2. A cis-acting element responsive to cold and drought, the C-repeat/dehydration-responsive element (C/DRE), was identified in the Arabidopsis thaliana stress-inducible genes RD29A3 and COR15a4 and found in other cold-inducible genes in various plants1,5. C/DRE-binding factor/DRE-binding protein (CBF/DREB) is an essential component of the cold-acclimation response1,2, but the signaling pathways and networks are mostly unknown. Here we used targeted genetic approach to isolate A. thaliana mutants with altered cold-responsive gene expression (acg) and identify ACG1 as a negative regulator of the CBF/DREB pathway. acg1 flowered late and had elevated expression of FLOWERING LOCUS C (FLC)6, a repressor of flowering encoding a MADS-box protein. We showed that acg1 is a null allele of the autonomous pathway gene FVE. FVE encodes a homolog of the mammalian retinoblastoma-associated protein, a component of a histone deacetylase (HDAC) complex involved in transcriptional repression7,8. We also showed that plants sense intermittent cold stress through FVE and delay flowering with increasing expression of FLC. Dual roles of FVE in regulating the flowering time and the cold response may have an evolutionary advantage for plants by increasing their survival rates.
Similar content being viewed by others
Main
To circumvent the problems associated with the complicated interactive effects of various cis-acting elements in the whole promoter region in studying signaling pathways and networks, we generated transgenic A. thaliana with four copies of the synthetic C/DRE derived from the COR15a promoter, fused to a GUS reporter gene with a minimal promoter containing a TATA box (4C/DRE-GUS), that responds to cold but not to other stress conditions, such as abscisic acid or high salinity (Fig. 1a)9. Light signaling mediated by phytochrome B is necessary for cold-induced gene expression through the C/DRE9. We mutagenized 4C/DRE-GUS plants by ethylmethane sulfonate and screened the resulting M2 population for mutants with constitutive GUS expression at 23 °C or under cold stress but without light by carrying out histochemical GUS assays on two excised leaves (Fig. 1b). We observed dwarfism and growth retardation in most mutants (data not shown) and delay in flowering in some mutants. We studied the acg1 mutant, which had the most severe delay in flowering, to explore the genetic link between cold-stress signaling and the phase transition to flowering. Genetic analysis of the acg1 mutant showed that aberrant GUS expression at 23 °C was due to a single recessive nuclear mutation that cosegregated with the late-flowering phenotype (Fig. 1c). acg1 mutants had higher GUS expression at 23 °C or 3 °C in the dark than 4C/DRE-GUS plants (hereafter called wild-type; Fig. 1d). Cold treatment in the light further enhanced GUS expression in acg1 relative to wild-type plants.
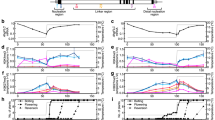
(a) Diagram of the chimeric construct used for generating 4C/DRE-GUS transgenic A. thaliana. WT, wild-type. (b) A typical example of screening that shows putative 4C/DRE-GUS mutants with constitutive GUS expression. (c) Genetic analysis of acg1 mutants by backcrossing with wild-type plants. ND, not determined. (d) Fluorometric quantification of GUS activities of acg1 mutants and wild-type plants (WT). Specific activities (units) are shown as pmol reaction product (4-methylumbelliferone) generated per min per mg of total protein. Data are mean ± standard error of three independent treatments.
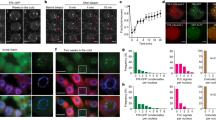
To investigate alteration of cold-responsive gene expression in acg1 mutants, we examined the time course of GUS induction in response to cold in acg1 and wild-type plants using northern-blot analysis (Fig. 2a). The level of GUS mRNA was higher in acg1 mutants than in wild-type plants before cold treatment and began in increase in acg1 mutants, but not wild-type plants, 4 h after cold treatment. By 6–8 h after cold treatment, the level of GUS mRNA was much higher in mutants than in wild-type plants. Longer cold treatment substantially increased GUS mRNA levels in mutants and wild-type plants to similar levels.
(a) Northern-blot analyses of cold-responsive gene expression in acg1 mutants and wild-type plants (WT). Plants were incubated at 1 °C with varying times in the light. The X-ray film exposure time of northern blots of the GUS mRNA from the samples treated for 6–48 h was three times shorter than that of the samples treated for 1–4 h. Actin7 (ACT7) mRNA was used as a loading control for RNA samples. (b) Electrolyte leakage of leaves from acg1 mutants and wild-type plants at different temperatures with or without cold acclimation (incubation at 1 °C for 2 d in the light). Data are mean ± standard error of at least 10 different samples. Broken lines, wild-type plants; solid lines, acg1 mutants; filled circles, wild-type plants without cold acclimation; filled triangles, acg1 mutants without cold acclimation; open circles, wild-type plants with cold acclimation; open triangles, acg1 mutants with cold acclimation.
To test whether the mutation affected expression of COR genes containing the C/DRE, we analyzed induction kinetics of COR15a and COR47 and observed an expression pattern similar to that of GUS, although expression of COR47 was not as robust as that of COR15a. We could not detect the transcripts of three CBF genes at 23 °C in acg1, suggesting that the constitutive expression of GUS and COR in acg1 at normal temperatures was not caused by enhanced expression of CBF. We observed similarly increased levels of CBF transcripts in response to cold in wild-type plants and mutants, but the transient increase in CBF expression occurred earlier in acg1 mutants than in wild-type plants. Thus, enhanced expression of COR15a in response to cold may be due, in part, to earlier expression of CBF. These results indicate that the acg1 mutation impairs the repression of the CBF/DREB pathway.
To test whether the acg1 mutation had an effect on freezing tolerance, we conducted an electrolyte leakage test10 on 2-week-old acg1 mutant and wild-type plants before and after preincubation at 4 °C for 2 d (Fig. 2b). Nonacclimated acg1 and wild-type plants showed similar tolerance to freezing, but when cold-acclimated, acg1 mutants had greater freezing tolerance than wild-type plants, in the range of −7 to −9 °C. This difference is statistically significant and may be due to the enhanced upregulation of COR genes in response to cold.
acg1 mutants flowered later than wild-type plants under both long days and short days, and the flowering time of acg1 mutants was reduced to wild-type level by vernalization treatment (Fig. 3a–c), suggestive of a mutation in the autonomous pathway. In A. thaliana, mutations in autonomous pathway genes cause elevated FLC expression and late flowering, but vernalization offsets this effect11. FLC expression was higher in acg1 mutants than in Columbia-0 (Col-0) plants, similar to the level of expression seen in fve-3 plants (Fig. 3d). fca-9 (ref. 12) plants, which flowered later than acg1 and fve-3 (ref. 13) plants, had a higher FLC level, but co-1 (ref. 14) plants, which are late-flowering only in long-day photoperiods, had a similar FLC level as Col-0. FLC negatively regulates the expression of SUPPRESSOR OF OVEREXPRESSION OF CO 1 (SOC1), encoding a MADS-box transcription factor, and FLOWERING LOCUS T (FT), encoding a protein with similarity to a Raf kinase inhibitor, both of which activate flower meristem identity genes15. We observed decreased levels of SOC1 and FT in acg1 mutants as well as in fve-3 and fca-9 plants. These results and complementation tests (Supplementary Fig. 1 online) suggest that late flowering in acg1 mutants is due to elevated FLC expression and that acg1 is an allele of fve, an autonomous pathway gene13,16. Expression of COR15a at 23 °C was enhanced in fve-3, fve-4, fve-4 flc-3 plants, as in acg1 mutants, but expression in flc-3 plants (Fig. 3e) was similar to that seen in Col-0 plants, suggesting that the effect of FVE on expression of cold-responsive genes is independent of FLC function.
(a) 6-week-old wild-type (WT; left) and acg1 (right) plants grown under long-day conditions (16 h light–8 h dark). (b) The rosette leaf numbers of acg1 mutants and wild-type plants (WT) vernalized for 6 weeks (solid column) or not vernalized (open column) and grown under long-day conditions. Time to flowering was measured as the number of rosette leaves formed on the main shoot. (c) The rosette leaf numbers of acg1 mutants and wild-type plants (WT) grown under short-day conditions (8 h light–16 h dark). Counting of the rosette leaf number of acg1 mutants was stopped after growing the plants for 7 months. (d) RT-PCR analysis of expression of FLC, SOC1 and FT in Col-0, acg1 and other late-flowering mutants. β-tubulin (TUBB) mRNA was used as a loading control for RNA samples. (e) Northern-blot analysis of COR15a expression in fve-3, fve-4, fve-4 flc-3 and flc-3 mutants compared with acg1 and Col-0 plants.
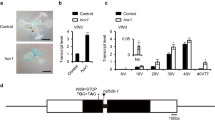
FVE was isolated by positional cloning17. Linkage analysis confirmed that acg1 might be fve, an allele of FVE (Fig. 4a). acg1 or fve-3 has a point mutation that converts Trp46 or Trp142, respectively, into a premature stop codon (Fig. 4b). The level of FVE mRNA was much lower in acg1 mutants than in Col-0 and wild-type plants, suggestive of mRNA instability due to a premature termination (Fig. 4c). The protein encoded by FVE shows high sequence homology to the mammalian retinoblastoma-associated protein (RbAp; Supplementary Fig. 2 online) and is predicted to contain five WD-40 repeats, domains that are found in multiprotein complexes involved in numerous cellular processes18. In mammals, the retinoblastoma (Rb) protein represses transcription by recruiting HDAC involved in repressing gene expression by deacetylation of histone residues7. RbAp interacts with HDAC associated with Rb and has a role in transcriptional repression8. These mechanisms seem to be conserved in plants, although the biological roles of the homologous proteins are not known19,20. A loss-of-function mutation in FVE could release the transcriptional repression mediated by HDAC, resulting in aberrant expression of GUS and COR at normal temperatures. This alteration of COR expression could affect a negative feedback loop of CBF expression, shifting it to an earlier time. Release of the transcriptional repression and earlier CBF expression could enhance the response of COR expression to cold in acg1 mutants (Fig. 2a). FVE mRNA was not affected by cold (Fig. 4d).
(a) Linkage analysis of the late-flowering phenotype of acg1 with FVE. Homozygous acg1 mutants in the segregated F2 population obtained from crossing acg1 with the ecotype Landsberg erecta were used for mapping. (b) Scheme of domain structures of the protein encoded by FVE and positions of nonsense mutations in acg1 and fve-3. Numbers above the thick column denote the amino acid numbers. WD40 indicates WD-40-repeat domain predicted by a Motif Scan program25. (c) Northern blot showing the FVE mRNA level in Col-0, wild-type (WT) and acg1 plants. (d) Northern-blot analysis of FVE expression in response to cold in Col-0 plants. Plants were incubated at 1 °C for varying periods of time (h). COR15a was analyzed for monitoring cold-inducible gene expression.
Because the same mutation caused both enhanced response of cold-induced gene expression and late flowering, we tested whether cold stress affected flowering time by treating plants at 4 °C for 2 h or 5 h every day during the entire growth period. Intermittent cold treatment substantially delayed flowering in wild-type plants but had no effect on acg1 and fve mutants (Fig. 5a). Moreover, intermittent cold treatment increased the level of FLC (Fig. 5b) and thereby decreased SOC1 expression (Fig. 5c). Genetic and expression analyses of various flowering-time mutants have indicated that a low ambient growth temperature (such as 16 °C) is sensed through a genetic pathway requiring both FCA and FVE21. Our results suggest that plants sense intermittent cold stress through FVE and delay flowering time by increasing the level of FLC. Consistent with a proposed role of FVE as a component of HDAC, the fve mutant had an increase in acetylation of FLC, as shown by chromatin immunoprecipitation analysis of the acetylation state of histone H4 at FLC22. Dual roles of FVE in regulating the flowering time and the stress response to cold may provide evolutionary fitness to plants, particularly in enduring the cold spell of early spring.
Methods
Plant materials and growth conditions.
We used the ecotype Columbia-0 for all experiments. We obtained the late-flowering mutants fve-3, fca-9, ld-1 and co-1 from Arabidopsis Biological Resource Center and fld-3, fve-4, fve-4 flc-3 and flc-3 seeds from R. Amasino (University of Wisconsin, Madison). We grew plants for 10 d on agar medium under long-day conditions (16 h light–8 h dark) at 23 °C and used them for cold treatment at 1–4 °C or for expression analysis of floral pathway genes. For flowering time analysis, we grew plants on soil under long-day (16 h light–8 h dark) or short-day conditions (8 h light–16 h dark). For analysis of the effect of intermittent cold stress on the flowering time, we incubated plants on soil every day at 4 °C for 0, 2 or 5 h, starting 4 h after dawn.
Mutagenesis, mutant isolation and genetic analysis.
We mutagenized 4C/DRE-GUS seeds (approximately 20,000 seeds) with ethylmethane sulfonate and harvested M2 seeds. We planted M2 seeds individually in 150-mm × 20-mm plates (69 seedlings per plate) containing agar medium and grew M2 seedlings for 10 d under long-day conditions. We excised two leaves from each seedling and then carried out histochemical GUS assays in the 96-well plates. We then transferred putative mutants with GUS staining in the leaves to soil to set seeds (M3). We carried out GUS assays for 5–10 seedlings of M3 to isolate mutants that inherited GUS expression. We backcrossed acg1 mutants with wild-type plants and tested the resulting F1 seeds for both the late-flowering phenotype and GUS expression. We scored the F2 populations obtained from self-pollinated F1 plants for segregation with the late-flowering phenotype. We tested the late-flowering F3 populations for GUS expression by the histochemical method. For genetic mapping of the acg1 mutation, we crossed acg1 with the ecotype Landsberg erecta. We selected homozygous acg1 mutants in the segregated F2 population based on the late-flowering phenotype. We mapped the mutation using SSLP markers HYB7, HYB8 and PLS2. Primer sequences are available on request.
Analysis of GUS expression.
We carried out histochemical assays of GUS activity by incubating the treated seedlings in 5-bromo-4-chloro-3-indolyl glucuronide (DUCHEFA) at 37 °C for 24 h and removing the chlorophyll from green tissues by incubation in 100% ethanol, as described23. We carried out quantitative GUS assays in tissue extracts by fluorometric measurement of 4-methylumbelliferone produced from the β-D-glucuronide precursor23.
Analysis of gene expression.
We extracted total RNA using TRI Reagent (Molecular Research Center) and carried out northern-blot analysis as previously described9. We analyzed expression of the genes FLC, SOC1, FT and TUBB (encoding β-tubulin) by RT-PCR as described24. DNA probes used for northern-blot analysis and primer sequences and conditions for amplification are available on request.
Electrolyte leakage test.
We determined freezing-induced electrolyte leakage from fully developed rosette leaves from 2-week-old plants grown in soil as described10. We placed one excised leaflet in a 5-ml test tube containing 100 μl of deionized water in a refrigerated circulator bath (PolyScience) at 0 °C and programmed the temperature of the bath to decrease to −9 °C in 1-°C increments over 30 min. We used at least 10 samples for each temperature.
Sequencing of DNA from mutants.
We extracted genomic DNA of mutants using DNeasy plant mini-kit (Qiagen) and amplified the FVE genomic region by PCR using two sets of specific primers, HYB15F and MJH6-2, and HYB6 and HYB15R. Primer sequences are available on request. We sequenced six independent PCR products for each amplified DNA fragment.
Note: Supplementary information is available on the Nature Genetics website.
References
Thomashow, M.F. Plant cold acclimation: freezing tolerance genes and regulatory mechanisms. Ann. Rev. Plant Physiol. Plant Mol. Biol. 50, 571–599 (1999).
Shinozaki, K. & Yamaguchi-Shinozaki, K. Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr. Opin. Plant Biol. 3, 353–360 (2000).
Yamaguchi-Shinozaki, K. & Shinozaki, K. A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low temperature, or high-salt stress. Plant Cell 6, 251–264 (1994).
Baker, S.S., Wilhelm, K.S. & Thomashow, M.F. The 5′-region of Arabidopsis thaliana cor15a has cis-acting elements that confer cold-, drought- and ABA-regulated gene expression. Plant Mol. Biol. 24, 701–713 (1994).
Jaglo, K.R. et al. Components of the Arabidopsis C-repeat/dehydration- responsive element binding factor cold-response pathway are conserved in Brassica napus and other plant species. Plant Physiol. 127, 910–917 (2001).
Michaels, S.D. & Amasino, R.M. FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11, 949–956 (1999).
Brehm, A. et al. Retinoblastoma protein recruits histone deacetylase to repress transcription. Nature 391, 597–601 (1998).
Nicolas, E. et al. RbAp48 belongs to the histone deacetylase complex that associates with the retinoblastoma protein. J. Biol. Chem. 275, 9797–9804 (2000).
Kim, H.-J., Kim, Y.-K., Park, J.-Y. & Kim, J. Light signalling mediated by phytochrome plays an important role in cold-induced gene expression through the C-repeat/dehydration responsive element (C/DRE) in Arabidopsis thaliana. Plant J. 29, 693–704 (2002).
Ishitani, M., Xiong, L., Lee, H., Stevenson, B. & Zhu, J.-K. HOS1, a genetic locus involved in cold-responsive gene expression in Arabidopsis. Plant Cell 10, 1151–1161 (1998).
Simpson, G.G. & Dean, C. Arabidopsis, the rosetta stone of flowering time? Science 296, 285–289 (2002).
Macknight, R. et al. FCA, a gene controlling flowering time in Arabidopsis, encodes a protein containing RNA-binding domains. Cell 89, 737–745 (1997).
Koornneef, M., Hanhart, C.J. & Van deer Veen, J.H. A genetic and physiological analysis of late flowering mutants in Arabidopsis thaliana. Mol. Gen. Genet. 229, 57–66 (1991).
Putterill, J., Robson, F., Lee, K., Simon, R. & Coupland, G. The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc finger transcription factors. Cell 80, 847–857 (1995).
Samach, A. et al. Distinct roles of CONSTANS target genes in reproductive development of Arabidopsis. Science 288, 1613–1616 (2000).
Sheldon, C.C. et al. The control of flowering by vernalization. Curr. Opin. Plant Biol. 3, 418–422 (2000).
Ausín, I., Alonso-Blanco, C., Jarillo, J.A., Ruiz-García, L. & Martínez-Zapater, J.M. Regulation of flowering time by FVE, a retinoblastoma-associated protein. Nat. Genet. advance online publication, 25 January 2004 (doi:10.1038/ng1295).
Smith, T.F., Gaitatzes, C., Saxena, K. & Neer, E.J. The WD repeat: a common architecture for diverse functions. Trends Biochem. Sci. 24, 181–185 (1999).
Rossi, V. et al. A maize histone deacetylase and retinoblastoma-related protein physically interact and cooperate in repressing gene transcription. Plant Mol. Biol. 51, 401–413 (2003).
Wu, K., Tian, L., Zhou, C., Brown, D. & Miki, B. Repression of gene expression by Arabidopsis HD2 histone deacetylases. Plant J. 34, 241–247 (2003).
Blázquez, M.A., Ahn, J.H. & Weigel, D. A thermosensory pathway controlling flowering time in Arabidopsis thaliana. Nat. Genet. 33, 168–171 (2003).
He, Y., Michaels, S.D. & Amasino, R.M. Regulation of flowering time by histone acetylation in Arabidopsis. Science advance online publication, 30 October 2003 (doi:10.1126/science.1091109).
Jefferson, R.A. & Wilson, K.J. The GUS gene fusion system. Plant Mol. Biol. Manual B14, 1–33 (1991).
Lee, H. et al. The AGAMOUS-LIKE 20 MADS domain protein integrates floral inductive pathways in Arabidopsis. Genes Dev. 14, 2366–2376 (2000).
Falquet, L. et al. The PROSITE database, its status in 2002. Nucleic Acids Res. 30, 235–238 (2002).
Acknowledgements
We thank R. Amasino for fld-3, fve-4, fve-4 flc-3 and flc-3 seeds and for comments on the manuscript and J.M. Martínez-Zapater for sharing unpublished data. This work was supported by grants from Plant Diversity Research Center of 21st Century Frontier Research Program (J.K. and I.L.) funded by Ministry of Science and Technology of Korean Government and from Agricultural Plant Stress Research Center (J.K.) funded by Korea Science and Engineering Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Kim, HJ., Hyun, Y., Park, JY. et al. A genetic link between cold responses and flowering time through FVE in Arabidopsis thaliana. Nat Genet 36, 167–171 (2004). https://doi.org/10.1038/ng1298
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1298
This article is cited by
-
Two B-box domain proteins, BBX28 and BBX29, regulate flowering time at low ambient temperature in Arabidopsis
Plant Molecular Biology (2021)
-
Plant abiotic stress response and nutrient use efficiency
Science China Life Sciences (2020)
-
Phalaenopsis flowering locus VE regulates floral organ maturation
Plant Cell Reports (2018)
-
Research progress on the autonomous flowering time pathway in Arabidopsis
Physiology and Molecular Biology of Plants (2017)
-
A cis cold memory element and a trans epigenome reader mediate Polycomb silencing of FLC by vernalization in Arabidopsis
Nature Genetics (2016)