Abstract
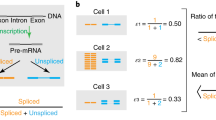
Transcription is a slow and expensive process: in eukaryotes, approximately 20 nucleotides can be transcribed per second1,2 at the expense of at least two ATP molecules per nucleotide3. Thus, at least for highly expressed genes, transcription of long introns, which are particularly common in mammals, is costly. Using data on the expression of genes that encode proteins in Caenorhabditis elegans and Homo sapiens, we show that introns in highly expressed genes are substantially shorter than those in genes that are expressed at low levels. This difference is greater in humans, such that introns are, on average, 14 times shorter in highly expressed genes than in genes with low expression, whereas in C. elegans the difference in intron length is only twofold. In contrast, the density of introns in a gene does not strongly depend on the level of gene expression. Thus, natural selection appears to favor short introns in highly expressed genes to minimize the cost of transcription and other molecular processes, such as splicing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Ucker, D.S. & Yamamoto, K.R. Early events in the stimulation of mammary tumor virus RNA synthesis by glucocorticoids. Novel assays of transcription rates. J. Biol. Chem. 259, 7416–7420 (1984).
Izban, M.G. & Luse, D.S. Factor-stimulated RNA polymerase-II transcribes at physiological elongation rates on naked DNA but very poorly on chromatin templates. J. Biol. Chem. 267, 13647–13655 (1992).
Lehninger, A.L., Nelson, D.L. & Cox, M.M. Principles of Biochemistry 615–644 (Worth, New York, 1982).
Deutsch, M. & Long, M. Intron-exon structures of eukaryotic model organisms. Nucleic Acids Res. 27, 3219–3228 (1999).
Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Ogata, H., Fujibuchi, W. & Kanehisa, M. The size differences among mammalian introns are due to the accumulation of small deletions. FEBS Lett. 390, 99–103 (1996).
Moriyama, E.N., Petrov, D.A. & Hartl, D.L. Genome size and intron size in Drosophila. Mol. Biol. Evol. 15, 770–773 (1998).
Hill, A.A., Hunter, C.P., Tsung, B.T., Tucker-Kellogg, G. & Brown, E.L. Genomic analysis of gene expression in C. elegans. Science 290, 809–812 (2000).
Eyre-Walker, A. Synonymous codon bias is related to gene length in Escherichia coli: selection for translational accuracy? Mol. Biol. Evol. 13, 864–872 (1996).
Duret, L. & Mouchiroud, D. Expression pattern and, suprisingly, gene length shape codon usage in Caenorhabditis, Drosophila, and Arabidopsis. Proc. Natl Acad. Sci. USA 96, 4482–4487 (1999).
Coghlan, A. & Wolfe, H. Relationship of codon bias to mRNA concentration and protein length in Saccharomyces cerevisiae. Yeast 16, 1131–1145 (2000).
Nixon, J.E. et al. A spliceosomal intron in Giardia lamblia. Proc. Natl Acad. Sci. USA 99, 3701–3705 (2002).
Ophir, R. & Graur, D. Patterns and rates of indel evolution in processed pseudogenes from humans and murids. Gene 205, 191–202 (1997).
Petrov, D.A., Lozovskaya, E.R. & Hartl, D.L. High intrinsic rate of DNA loss in Drosophila. Nature 384, 346–349 (1998).
Robertson, H.M. The large srh family of chemoreceptor genes in Caenorhabditis nematodes reveals processes of genome evolution involving large duplications and deletion and intron gains and losses. Genome Res. 10, 192–203 (2000).
Boulikas, T. Evolutionary consequences of nonrandom damage and repair of chromatin domains. J. Mol. Evol. 35, 156–180 (1992).
Sullivan, D.T. DNA excision repair and transcription: implications for genome evolution. Curr. Opin. Genet. Dev. 5, 786–791 (1995).
Reinke, V. et al. A global profile of germline gene expression in C. elegans. Mol. Cell 6, 605–616 (2000).
Zhou, Z. et al. The protein Aly links pre-messenger-RNA splicing to nuclear export in metazoans. Nature 407, 401–405 (2000).
Carvalho, A.B. & Clark, A.G. Intron size and natural selection. Nature 401, 344 (1999).
Comeron, J.M. & Kreitman, M. The correlation between intron length and recombination in Drosophila. Dynamic equilibrium between mutational and selective forces. Genetics 156, 1175–1190 (2000).
Hurst, L.D., Brunton, C.F.A. & Smith, N.G.C. Small introns tend to occur in GC-rich regions in some but not all vertebrates. Trends Genet. 15, 437–439 (1999).
Carels, N. & Bernardi, G. Two classes of genes in plants. Genetics 154, 1819–1825 (2000).
Hurst, L.D. & McVean, G. Imprinted genes have few and short introns. Nature Genet. 12, 234–237 (1996).
Akashi, H. Gene expression and molecular evolution. Curr. Opin. Genet. Dev. 11, 660–666 (2001).
Okubo, K. et al. Large scale cDNA sequencing for analysis of quantitative and qualitative aspects of gene expression. Nature Genet. 2, 173–179 (1992).
Lee, N.H. et al. Comparative expressed-sequence-tag analysis of differential gene expression profiles in PC-12 cells before and after nerve growth factor treatment. Proc. Natl Acad. Sci. USA 92, 8303–8307 (1995).
Bortoluzzi, S. & Danieli, G.A. Towards an in silico analysis of transcription patterns. Trends Genet. 15, 118–119 (1999).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
The C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
Acknowledgements
We are grateful to A. Kondrashov, I. Rogozin and A. Feldman for reading the manuscript and P. Bouman, J. Cherry, J. Blumensteil and T. Kim for discussion.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Castillo-Davis, C., Mekhedov, S., Hartl, D. et al. Selection for short introns in highly expressed genes. Nat Genet 31, 415–418 (2002). https://doi.org/10.1038/ng940
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng940
This article is cited by
-
Protein phosphatase 1 regulatory subunit 15 A promotes translation initiation and induces G2M phase arrest during cuproptosis in cancers
Cell Death & Disease (2024)
-
An effect of large-scale deletions and duplications on transcript expression
Functional & Integrative Genomics (2023)
-
Intron size minimisation in teleosts
BMC Genomics (2022)
-
Genomic insights into positive selection during barley domestication
BMC Plant Biology (2022)
-
STREAMING-tag system reveals spatiotemporal relationships between transcriptional regulatory factors and transcriptional activity
Nature Communications (2022)