Abstract
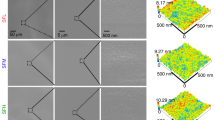
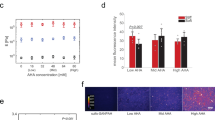
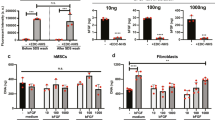
Stem cells regulate their fate by binding to, and contracting against, the extracellular matrix. Recently, it has been proposed that in addition to matrix stiffness and ligand type, the degree of coupling of fibrous protein to the surface of the underlying substrate, that is, tethering and matrix porosity, also regulates stem cell differentiation. By modulating substrate porosity without altering stiffness in polyacrylamide gels, we show that varying substrate porosity did not significantly change protein tethering, substrate deformations, or the osteogenic and adipogenic differentiation of human adipose-derived stromal cells and marrow-derived mesenchymal stromal cells. Varying protein–substrate linker density up to 50-fold changed tethering, but did not affect osteogenesis, adipogenesis, surface–protein unfolding or underlying substrate deformations. Differentiation was also unaffected by the absence of protein tethering. Our findings imply that the stiffness of planar matrices regulates stem cell differentiation independently of protein tethering and porosity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pelham, R. J. Jr & Wang, Y. Cell locomotion and focal adhesions are regulated by substrate flexibility. Proc. Natl Acad. Sci. USA 94, 13661–13665 (1997).
Gilbert, P. M. et al. Substrate elasticity regulates skeletal muscle stem cell self-renewal in culture. Science 329, 1078–1081 (2010).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Choi, Y. S., Vincent, L. G., Lee, A. R., Dobke, M. K. & Engler, A. J. Mechanical derivation of functional myotubes from adipose-derived stem cells. Biomaterials 33, 2482–2491 (2012).
Saha, K. et al. Substrate modulus directs neural stem cell behavior. Biophys. J. 95, 4426–4438 (2008).
Rowlands, A. S., George, P. A. & Cooper-White, J. J. Directing osteogenic and myogenic differentiation of MSCs: Interplay of stiffness and adhesive ligand presentation. Am. J. Physiol. Cell Physiol. 295, C1037–C1044 (2008).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
McBeath, R., Pirone, D. M., Nelson, C. M., Bhadriraju, K. & Chen, C. S. Cell shape, cytoskeletal tension, and RhoA regulate stem cell lineage commitment. Dev. Cell 6, 483–495 (2004).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Kilian, K. A., Bugarija, B., Lahn, B. T. & Mrksich, M. Geometric cues for directing the differentiation of mesenchymal stem cells. Proc. Natl Acad. Sci. USA 107, 4872–4877 (2010).
Huebsch, N. et al. Harnessing traction-mediated manipulation of the cell/matrix interface to control stem-cell fate. Nature Mater. 9, 518–526 (2010).
Khetan, S. et al. Degradation-mediated cellular traction directs stem cell fate in covalently crosslinked three-dimensional hydrogels. Nature Mater. 12, 458–465 (2013).
Baneyx, G., Baugh, L. & Vogel, V. Fibronectin extension and unfolding within cell matrix fibrils controlled by cytoskeletal tension. Proc. Natl Acad. Sci. USA 99, 5139–5143 (2002).
Smith, M. L. et al. Force-induced unfolding of fibronectin in the extracellular matrix of living cells. PLoS Biol. 5, e268 (2007).
Trappmann, B. et al. Extracellular-matrix tethering regulates stem-cell fate. Nature Mater. 11, 642–649 (2012).
Rossman, J. S. & Dym, C. L. Introduction to Engineering Mechanics: A Continuum Approach (CRC Press, 2008).
Plotnikov, S. V., Pasapera, A. M., Sabass, B. & Waterman, C. M. Force fluctuations within focal adhesions mediate ECM-rigidity sensing to guide directed cell migration. Cell 151, 1513–1527 (2012).
Holle, A. W. et al. In situ mechanotransduction via vinculin regulates stem cell differentiation. Stem Cells 31, 2467–2477 (2013).
Lutolf, M. P., Gilbert, P. M. & Blau, H. M. Designing materials to direct stem-cell fate. Nature 462, 433–441 (2009).
Guilak, F. et al. Control of stem cell fate by physical interactions with the extracellular matrix. Cell Stem Cell 5, 17–26 (2009).
Discher, D. E., Mooney, D. J. & Zandstra, P. W. Growth factors, matrices, and forces combine and control stem cells. Science 324, 1673–1677 (2009).
Stellwagen, N. C. Apparent pore size of polyacrylamide gels: Comparison of gels cast and run in tris-acetate-EDTA and tris-borate-EDTA buffers. Electrophoresis 19, 1542–1547 (1998).
Del Alamo, J. C. et al. Three-dimensional quantification of cellular traction forces and mechanosensing of thin substrata by fourier traction force microscopy. PLoS ONE 8, e69850 (2013).
Ruoslahti, E. & Pierschbacher, M. D. New perspectives in cell adhesion: RGD and integrins. Science 238, 491–497 (1987).
Engler, A. et al. Substrate compliance versus ligand density in cell on gel responses. Biophys. J. 86, 617–628 (2004).
Murrell, M., Kamm, R. & Matsudaira, P. Substrate viscosity enhances correlation in epithelial sheet movement. Biophys. J. 101, 297–306 (2011).
Bangasser, B. L., Rosenfeld, S. S. & Odde, D. J. Determinants of maximal force transmission in a motor-clutch model of cell traction in a compliant microenvironment. Biophys. J. 105, 581–592 (2013).
Fu, J. et al. Mechanical regulation of cell function with geometrically modulated elastomeric substrates. Nature Methods 7, 733–736 (2010).
Khatiwala, C. B., Peyton, S. R., Metzke, M. & Putnam, A. J. The regulation of osteogenesis by ECM rigidity in MC3T3-E1 cells requires MAPK activation. J. Cell. Physiol. 211, 661–672 (2007).
Young, J. L. & Engler, A. J. Hydrogels with time-dependent material properties enhance cardiomyocyte differentiation in vitro. Biomaterials 32, 1002–1009 (2011).
Chopra, A. et al. Reprogramming cardiomyocyte mechanosensing by crosstalk between integrins and hyaluronic acid receptors. J. Biomech. 45, 824–831 (2012).
Kong, H. J., Polte, T. R., Alsberg, E. & Mooney, D. J. FRET measurements of cell-traction forces and nano-scale clustering of adhesion ligands varied by substrate stiffness. Proc. Natl Acad. Sci. USA 102, 4300–4305 (2005).
Choi, Y. S. et al. The alignment and fusion assembly of adipose-derived stem cells on mechanically patterned matrices. Biomaterials 33, 6943–6951 (2012).
Carpenter, A. E. et al. CellProfiler: Image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Kaushik, G. et al. Measuring passive myocardial stiffness in Drosophila melanogaster to investigate diastolic dysfunction. J. Cell. Mol. Med. 16, 1656–1662 (2012).
Bonanni, B. et al. Single molecule recognition between cytochrome C 551 and gold-immobilized azurin by force spectroscopy. Biophys. J. 89, 2783–2791 (2005).
Chirasatitsin, S. & Engler, A. J. Detecting cell-adhesive sites in extracellular matrix using force spectroscopy mapping. J. Phys. Condens. Matter 22, 194102 (2010).
Fuhrmann, A., Anselmetti, D., Ros, R., Getfert, S. & Reimann, P. Refined procedure of evaluating experimental single-molecule force spectroscopy data. Phys. Rev. E 77, 031912 (2008).
Acknowledgements
The authors thank M. Joens, J. Kasuboski and J. Fitzpatrick at the Waitt Advanced Biophotonics Center at the Salk Institute (supported by NCI P30 Cancer Center Support Grant CA014195-40 and NINDS P30 Neuroscience Center Core Grant NS072031-03A1 and by the W. M. Keck Foundation) for technical assistance with microscopy, and W. Murphy (University of Wisconsin), T. McDevitt (Georgia Tech) and J. C. del Alamo (UC San Diego) for helpful conversations. The National Institutes of Health (DP02OD006460 to A.J.E.), the Human Frontiers Science Program (RGY0064/2010 to A.J.E.), the National Science Foundation Graduate Research Fellowship Program (to J.H.W., L.G.V. and H.T-W.), the Siebel Scholars Program, and the Achievement Rewards for College Scientists (to L.G.V.) supported this work.
Author information
Authors and Affiliations
Contributions
All authors contributed to the design of experiments. Y.S.C., K.C.H. and S.C. characterized the hydrogel substrates. H.T-W. characterized and performed experiments with the FRET probe. J.H.W., L.G.V. and A.F. conducted all other experiments and performed the data analysis. J.H.W., L.G.V. and A.J.E. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 6452 kb)
Rights and permissions
About this article
Cite this article
Wen, J., Vincent, L., Fuhrmann, A. et al. Interplay of matrix stiffness and protein tethering in stem cell differentiation. Nature Mater 13, 979–987 (2014). https://doi.org/10.1038/nmat4051
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat4051
This article is cited by
-
Zyxin regulates embryonic stem cell fate by modulating mechanical and biochemical signaling interface
Communications Biology (2023)
-
Computational indentation in weakly cross-linked polymer networks
International Journal of Advances in Engineering Sciences and Applied Mathematics (2023)
-
Soft nanofiber modified micropatterned substrates enhance native-like endothelium maturation via CXCR4/calcium-mediated actin cytoskeleton assembly
Nano Research (2023)
-
Interplay between mechanics and signalling in regulating cell fate
Nature Reviews Molecular Cell Biology (2022)
-
Hydrogel protection strategy to stabilize water-splitting photoelectrodes
Nature Energy (2022)