Abstract
Protein tagging is widely used in approaches ranging from affinity purification to fluorescence-based detection in live cells. However, an intrinsic limitation of tagging is that the native function of the protein may be compromised or even abolished by the presence of the tag. Here we describe and characterize a set of small, innocuous protein tags (inntags) that we anticipate will find application in a variety of biological techniques.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Shine, J., Fettes, I., Lan, N.C.Y., Roberts, J.L. & Baxter, J.D. Nature 285, 456–463 (1980).
Waugh, D.S. Trends Biotechnol. 23, 316–320 (2005).
Stadler, C. et al. Nat. Methods 10, 315–323 (2013).
Hoffmann, C. et al. Nat. Methods 2, 171–176 (2005).
Tompa, P. Trends Biochem. Sci. 37, 509–516 (2012).
Woestenenk, E.A., Hammarstrom, M., van den Berg, S., Hard, T. & Berglund, H. J. Struct. Funct. Genomics 5, 217–229 (2004).
Schembri, L. et al. Nat. Methods 4, 107–108 (2007).
Chant, A., Kraemer-Pecore, C.M., Watkin, R. & Kneale, G.G. Protein Expr. Purif. 39, 152–159 (2005).
Liu, X. et al. Proc. Natl. Acad. Sci. USA 94, 10669–10674 (1997).
Song, J. & Markley, J.L. Protein Pept. Lett. 14, 265–268 (2007).
Landgraf, D., Okumus, B., Chien, P., Baker, T.A. & Paulsson, J. Nat. Methods 9, 480–482 (2012).
Kopito, R.R. Trends Cell Biol. 10, 524–530 (2000).
Vavouri, T., Semple, J.I., Garcia-Verdugo, R. & Lehner, B. Cell 138, 198–208 (2009).
Sopko, R. et al. Mol. Cell 21, 319–330 (2006).
Cook, M. & Tyers, M. Curr. Opin. Biotechnol. 18, 341–350 (2007).
Morgan, D.O. The Cell Cycle: Principles of Control. (New Science Press, 2007).
Kim, Y.E., Hipp, M.S., Bracher, A., Hayer-Hartl, M. & Hartl, F.U. Annu. Rev. Biochem. 82, 323–355 (2013).
Vergés, E., Colomina, N., Garí, E., Gallego, C. & Aldea, M. Mol. Cell 26, 649–662 (2007).
Ferrezuelo, F. et al. Nat. Commun. 3, 1012 (2012).
Fredriksson, S. et al. Nat. Methods 4, 327–329 (2007).
Zhang, X. et al. Nat. Methods 5, 163–165 (2008).
Hayes, C.N. et al. Bioinformatics 24, 2564–2565 (2008).
Vita, R. et al. Nucleic Acids Res. 38, D854–D862 (2010).
Bernstein, F.C. et al. J. Mol. Biol. 112, 535–542 (1977).
Punta, M. et al. Nucleic Acids Res. 40, D290–D301 (2012).
Saha, S. & Raghava, G.P.S. in Artificial Immune Systems (eds. Nicosia, G. et al.) 197–204 (Springer, 2004).
Humphrey, W., Dalke, A. & Schulten, K. J. Mol. Graph. 14, 33–38 (1996).
The Gene Ontology Consortium. Nucleic Acids Res. 41, D530–D535 (2013).
Xue, B. et al. Biochim. Biophys. Acta 1804, 996–1010 (2010).
Pedraza, N. et al. J. Neurosci. 34, 13988–13997 (2014).
Gallego, C., Garí, E., Colomina, N., Herrero, E. & Aldea, M. EMBO J. 16, 7196–7206 (1997).
Sambrook, J. & Russell, D.W. Molecular cloning: A laboratory manual. CSHL Press, Cold Spring Harbor, NY, (2001).
Yahya, G., Parisi, E., Flores, A., Gallego, C. & Aldea, M. Mol. Cell 53, 115–126 (2014).
Afkarian, M. et al. Mol. Cell. Proteomics 9, 2195–2204 (2010).
Wang, H., Garí, E., Vergés, E., Gallego, C. & Aldea, M. EMBO J. 23, 180–190 (2004).
Llanes, D., Nogal, M., Prados, F. & Viñuela, E. Hybridoma 10, 757–762 (1992).
Slaughter, B.D., Schwartz, J.W. & Li, R. Proc. Natl. Acad. Sci. USA 104, 20320–20325 (2007).
Acknowledgements
We thank A. Cornadó, E. Rebollo and S. Dillon (Harvard University) for technical assistance, M. Azor (AntibodyBcn) for administrative support and C. Mann (Centre de Génétique Moléculaire) for the anti-Cdc28 antibody. This work was funded by the Ministry of Economy and Competitiveness of Spain (INNPACTO IPT-010000-2010-19), Consolider-Ingenio 2010 (CSD2007-15), the Instituto Nacional de Bioinformática (INB) and the European Union (FEDER) and received financial support from Antibody BCN and Immunostep. M.O. acknowledges support from Institució Catalana de Recerca i Estudis Avançats.
Author information
Authors and Affiliations
Contributions
M.V.G., G.Y. and R.O. performed the experiments. L.C. carried out the bioinformatics screen. L.T. and J.C. obtained the monoclonal antibodies. A.I., M.J. and R.J. directed monoclonal antibody selection and preparation, and J.L.G. and M.O. directed the bioinformatics screen. C.G. and M.A. directed the wet lab experiments. C.G., M.O. and M.A. conceived and designed the experiments, and M.A. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
M.A. and M.O. are named on the pending patent application PCT/ES2014/070460 for inntags IT5, IT6 and IT10 for protein-tagging applications. This patent has been licensed by AbBcn Ltd. (AntibodyBcn). L.T., M.J. and A.I. are employed by AbBcn Ltd. J.C. and R.J. are employed by Immunostep Ltd.
Integrated supplementary information
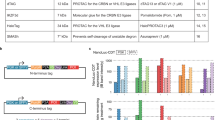
Supplementary Figure 1 Inntags selected by the bioinformatics screen, and expression and aggregation tests.
(a) List of selected structures as inntags. (b) GFP fusions of inntags IT1 to IT12 were overexpressed in HEK293T cells to analyze effects on expression and aggregation. As a measure of the aggregation propensity, an intracellular variability index (IV) was calculated as the coefficient of variation of fluorescence values in intracellular pixels, and representative images are shown as reference. Bar, 20 μm. (c) Expression levels of inntag-GFP fusions are shown as mean values (N=30 cells) and confidence limits for the mean (α=0.05). (d) Intracellular variability for the analyzed inntag-GFP fusions. Mean values (N=30 cells) and confidence limits for the mean (α=0.05) are shown.
Supplementary Figure 2 Stability and solubility tests of selected inntags.
(a) GFP-fusions of indicated inntags were expressed in HEK293T cells and total extracts were analyzed by immunoblotting with an αGFP antibody. (b) Levels of inntag-GFP fusions in the presence of 25 µg/ml cycloheximide for 12 h were obtained by immunoblotting and made relative to the untreated control. Mean values (N=3) and confidence limits for the mean (α=0.05) are shown as bars (left axis). A soluble fraction of HEK293T cells expressing the indicated GFP fusions was obtained by centrifugation, and levels obtained by immunoblotting were made relative to those corresponding to the total cell extract. Mean values (N=3) and confidence limits for the mean (α=0.05) are shown as open circles (right axis). (c) HEK293T cells expressing the indicated GFP fusions were analyzed by time-lapse confocal microscopy under FLIP conditions. The relative cellular fluorescence recorded at different times during photobleaching is shown. (d) Total extracts of mouse NIH3T3 cells expressing human cyclin E fused to IT5, IT6, or IT10 were analyzed by immunoblotting with αHsCycE antibodies. A control extract from cells expressing untagged human cyclin E (no tag) is also shown. (e) Levels of cyclin E fusions in the presence of 25 µg/ml cycloheximide for 2 h were obtained by immunoblotting and made relative to the untreated control. Mean values (N=3) and confidence limits for the mean (α=0.05) are shown as bars.
Supplementary Figure 3 Functional characterization of inntags and other commonly used tags.
(a) GFP fusions to the different tags were overexpressed in budding yeast cells to determine growth resumption rates (bars) and chronological life span (circles). Mean values (N=4) and confidence limits (α=0.05) for the mean are shown. (b) GFP fusions to the different tags were overexpressed in budding yeast cells to determine individual critical volumes at budding. The relative variance of cells (N>400) and confidence limits (α=0.05) are plotted.
Supplementary Figure 4 Integrity analysis of selected inntags after affinity purification from E. coli cells.
(a) 6His fusions of selected inntags were expressed in E. coli, purified by metal affinity chromatography, and analyzed by SDS-PAGE. Due to their very small size, inntags IT6 and IT10 were expressed and purified as dimers. A common contaminant protein is indicated by an asterisk. (b) The corresponding fusions to 6His-GFP were expressed, purified and analyzed as in a. Purified 6His-GFP (no tag) is shown as reference.
Supplementary Figure 5 Immunoprecipitation of selected inntags and other commonly used tags as GFP fusions in budding yeast.
(a, b) Total extracts of yeast cells overexpressing GFP fused to the indicated tags were analyzed by immunoblotting with specific (a) or αGFP (b) antibodies. An αDpm1 antibody was used as loading control, and a control extract of cells overexpressing GFP (no tag) is also shown. (c) Immunoprecipitates of cell extracts with specific antibodies as indicated in each lane were analyzed by immunoblotting with αGFP antibodies.
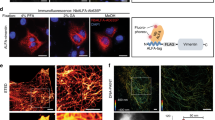
Supplementary Figure 6 Localization innocuity of inntag IT5 on proteins from different cellular compartments.
Inntag IT5 fused to βactin (cytoskeleton), STX6 (Golgi), HO1 (endoplasmic reticulum), histone H1 and HDAC2 (nucleus), or GRB2 (plasma membrane was expressed in NIH3T3 cells and analyzed by immunofluorescence (red signal) with an αIT5 antibody. GRB2 images correspond to mitotic cells, where the cytoplasmic membrane is best seen. Hoechst-stained nuclei are also shown. Bar, 10 μm.
Supplementary Figure 7 Stability and integrity of inntags and other commonly used tags fused to essential proteins in budding yeast.
(a, b) Total extracts of yeast cells expressing endogenous levels of Cdc28 (a) or Ydj1 (b) fused to IT5, IT6, HA3 or Flag6 were analyzed by immunoblotting with αCdc28 or αYdj1 antibodies, respectively. A control strain (no tag) is shown as reference.
Supplementary Figure 8 Effects on cell size by GFP as a large tag when fused to Cdc28 and Ydj1 proteins.
(a) Critical size of yeast cells expressing endogenous levels of wild-type Cdc28 (no tag) or fused to GFP (Cdc28-GFP). Mean values (N>400) and confidence limits (α=0.05) for the mean are plotted. (b) Cell volume distributions of yeast cells expressing endogenous levels of wild-type Ydj1 (no tag) or fused to GFP (Ydj1-GFP).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 949 kb)
Supplementary Table 1
List of candidates produced by the bioinformatics screen (XLS 68 kb)
Supplementary Table 2
Amino acid sequences of selected inntags (XLSX 12 kb)
Supplementary Table 3
Quantitative analysis of 6MYC and inntag IT6 interactors (XLSX 43 kb)
Rights and permissions
About this article
Cite this article
Georgieva, M., Yahya, G., Codó, L. et al. Inntags: small self-structured epitopes for innocuous protein tagging. Nat Methods 12, 955–958 (2015). https://doi.org/10.1038/nmeth.3556
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3556
This article is cited by
-
A novel epitope tag from rabies virus has versatile in vitro applications
Applied Microbiology and Biotechnology (2023)
-
The ALFA-tag is a highly versatile tool for nanobody-based bioscience applications
Nature Communications (2019)
-
Internal epitope tagging informed by relative lack of sequence conservation
Scientific Reports (2016)
-
An extra dimension in protein tagging by quantifying universal proteotypic peptides using targeted proteomics
Scientific Reports (2016)