Abstract
The brain's structural organization is so complex that 2,500 years of analysis leaves pervasive uncertainty about (i) the identity of its basic parts (regions with their neuronal cell types and pathways interconnecting them), (ii) nomenclature, (iii) systematic classification of the parts with respect to topographic relationships and functional systems and (iv) the reliability of the connectional data itself. Here we present a prototype knowledge management system (http://brancusi.usc.edu/bkms/) for analyzing the architecture of brain networks in a systematic, interactive and extendable way. It supports alternative interpretations and models, is based on fully referenced and annotated data and can interact with genomic and functional knowledge management systems through web services protocols.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Watson, J.D. & Crick, F.H.C. Molecular structure of nucleic acids: a structure for deoxyribose nucleic acid. Nature 171, 737–738 (1953).
Davidson, E.H. Genomic Regulatory Systems: Development and Evolution (Academic, San Diego, 2001).
Hasty, J., McMillen, D. & Collins, J.J. Engineered gene circuits. Nature 420, 224–230 (2002).
Swanson, L.W. Brain Architecture: Understanding the Basic Plan (Oxford Univ. Press, Oxford, 2003).
Meyer, A. Historical Aspects of Cerebral Anatomy (Oxford Univ. Press, London, 1971).
Swanson, L.W. A history of neuroanatomical mapping. in Brain Mapping: the Systems (eds. Toga, A.W. & Mazziotta, J.C.) 77–109 (Academic, San Diego, 2000).
Swanson, L.W. Cerebral hemisphere regulation of motivated behavior. Brain Res. 886, 113–164 (2000).
Nauta, W.J.H. & Ryan, L.F. Selective silver impregnation of degenerating axons in the central nervous system. Stain Technol. 27, 175–179 (1952).
Swanson, L.W. The neuroanatomy revolution of the 1970s and the hypothalamus. Brain Res. Bull. 50, 397 (1999).
Kandel, E.R., Schwartz, J.H. & Jessell, T.M. Principles of Neural Science 4th edn. (McGraw-Hill, New York, 2000).
Brodal, A. Neurological Anatomy (Oxford Univ. Press, London, 1948).
Steno, N. Lecture on the Anatomy of the Brain (facsimile and translation of the 1669 original, published by A. Busck, Copenhagen, 1965).
Golgi, C. Sulla struttura della sostanza grigia dell cervello. Gazz. Med. Ital. 33, 244–246 (1873).
Swanson, L.W. Mapping the human brain: past, present, and future. Trends Neurosci. 18, 471–474 (1995).
Swanson, L.W. What is the brain? Trends Neurosci. 23, 519–527 (2000).
Bota, M. & Arbib, M.A. Integrating databases and expert systems for the analysis of brain structures, connections, and homologies. Neuroinformatics (in press).
Finger, S. Origins of Neuroscience: a History of Explorations into Brain Function (Oxford Univ. Press, New York, 1994).
Clarke, E. & O'Malley, C.D. The Human Brain and Spinal Cord 2nd edn. (Norman, San Francisco, 1996).
Swanson, L.W. Brain Maps: Structure of the Rat Brain 2nd edn. (Elsevier, Amsterdam, 1999).
Wilder, B.G. Neural terms, international and national. J. Comp. Neurol. 6, 216–352 (1896).
Rasmussen, A.T. Some Trends in Neuroanatomy (Brown, Dubuque, 1947).
Dashti, A.E., Ghandeharizadeh, S., Stone, J., Swanson, L.W. & Thompson, R.H. Database challenges and solutions in neuroscientific applications. Neuroimage 5, 97–115 (1997).
Swanson, L.W. Interactive brain maps and atlases. in Computing the Brain: a Guide to Neuroinformatics (eds. Arbib, M.A. & Grethe, J.G.) 167–177 (Academic, San Diego, 2001).
Stephan, K.E., Zilles, K. & Kötter, R. Coordinate-independent mapping of structural and functional data by objective relational transformation (ORT). Phil. Trans. R. Soc. Lond. B Biol. Sci. 335, 37–54 (2000).
Stephan, K.E. et al. Advanced database methodology for the collation of connectivity data on the Macaque brain (CoCoMac). Phil. Trans. R. Soc. Lond. B Biol. Sci. 356, 1159–1186 (2001).
Burns, G.A.P.C. Knowledge management of the neuroscientific literature: the data model and underlying strategy of the NeuroScholar system. Phil. Trans. R. Soc. Lond. B Biol. Sci. 356, 1187–1208 (2001).
Egenhoffer, M. & Franzosa, R. Point-set topological spatial relations. Int. J. Geogr. Inf. Sci. 5, 161–174 (1991).
Burns, G.A.P.C. Neural Connectivity of the Rat: Theory, Methods and Applications. Thesis, Oxford Univ. (1997).
Burns, G.A.P.C. & Young, M.P. Analysis of the connectional organisation of neural systems associated with the hippocampus in rats. Phil. Trans. R. Soc. Lond. B Biol. Sci. 355, 55–70 (2000).
Csete, M.E. & Doyle, J.C. Reverse engineering of biological complexity. Science 295, 1664–1669 (2002).
Clark, W.E.L. Morphological aspects of the hypothalamus. in The Hypothalamus: Morphological, Functional, Clinical and Surgical Aspects (eds. Clark, W.E.L., Beattie, J., Riddoch, G. & Dott, N.M.) 2–68 (Oliver and Boyd, Edinburgh, 1938).
Ingram, W.R. Nuclear organization and chief connections of the primate hypothalamus. Res. Publ.Assoc. Res. Nerv. Ment. Dis. 20, 195–244 (1940).
Raisman, G. Neural connexions of the hypothalamus. Brit. Med. Bull. 22, 197–201 (1966).
Nauta, W.J.H. & Haymaker, W. Hypothalamic nuclei and fiber connections. in The Hypothalamus (eds. Haymaker, W., Anderson, E. & Nauta, W.J.H.) 136–209 (Thomas, Springfield, 1969).
Swanson, L.W. The hypothalamus. in Handbook of Chemical Neuroanatomy Vol. 5 (eds. Björklund, A., Hökfelt, T. & Swanson, L.W.) 1–124 (Elsevier, Amsterdam, 1987).
Raisman, G., Cowan, W.M. & Powell, T.P.S. The extrinsic afferent, commissural and association fibres of the hippocampus. Brain 88, 963–996 (1965).
Raisman, G., Cowan, W.M. & Powell, T.P.S. An experimental analysis of the efferent projection of the hippocampus. Brain 89, 83–108 (1966).
Cowan, W.M, Raisman, G. & Powell, T.P.S. The connexions of the amygdala. J. Neurol. Neurosurg. Psychiatry 28, 137–151 (1965).
Vesalius, A. De Humani corporis Fabrica Libri Septem (Oporinus, Basel, 1543).
Watson, J. The Double Helix: a Personal Account of the Discovery of the Structure of DNA (Atheneum, New York, 1968).
Brodmann, K. Vergleichende Localisationslehre der Grosshirnrinde in ihren Prinzipien dargestellt auf Grund des Zellenbaues (Barth, Leipzig, 1909) translated as Brodmann's Localization in the Cerebral Cortex (Smith-Gordon, London, 1994).
Cajal, S.R. Histologie du système nerveux de l'homme et des vertébrés, 2 vols. (Maloine, Paris, 1909–1911) translated as Histology of the Nervous System of Man and Vertebrates (Oxford Univ. Press, New York, 1995).
Paxinos, G. & Watson, C. The Rat Brain in Stereotaxic Coordinates 4th edn. (Academic, San Diego, 1998).
Armstrong, D.M., Saper, C.B., Levey, A.I., Wainer, B.H. & Terry, R.D. Distribution of cholinergic neurons in rat brain: demonstrated by the immunocytochemical localization of choline acetyltransferase. J. Comp. Neurol. 216, 53–68 (1983).
Gritti, I., Mainville, L. & Jones, B.E. Codistribution of GABA- with acetylcholine-synthesizing neurons in the basal forebrain of the rat. J. Comp. Neurol. 329, 438–457 (1993).
Acknowledgements
For invaluable discussions we especially thank M. Arbib, D. Bowden, G. Burns, S. Koslow, S. Subramaniam, and A. Toga. This work was supported by US National Institutes of Health grants PSA-99-060 and NS-16668.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Fig. 1.
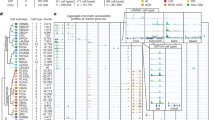
The general object-relationship (OR) schema of BAMS. Each object and relation in the figure can be captured in more than one table. The BAMS OR structure is centered on the object "Brain Part" as defined in different neuroanatomical nomenclatures (atlases). Any Brain Part is uniquely identified in BAMS with four attributes: name, species, atlas, and atlas version. The Brain Part module is constructed in an m:n relation with "Cell Types" module. It permits description of any cell type or subtype in terms of position within the associated brain region, distribution pattern, and numerical data (cell number, cell density, percentage of total cell number, and range of these parameters over multiple experiments). The "Connections" module is in an m:n relation with the Brain Part module because a brain region can send projections to, and receive inputs from, many other brain regions. BAMS can associate any major fiber tract registered in the Brain Part module with many reports of projections, and any neural connection can participate in many fiber tracts. The Connections module has a set of 40+ attributes allowing it to describe comprehensively connectivity reports collated from literature inserted by neuroanatomists. The object "Collator" stores basic information about people with permission to add, delete, or update information in BAMS. Collator is in a 1:n relationship with Brain Part because every brain part record is uniquely identified in BAMS and must therefore be inserted by a single collator. The object "Reference" stores information about sources used to insert data. The object Reference and the Brain Part module are in a 1:n relationship because atlas and atlas version are two attributes that define uniquely any brain record and refer to a single source. The Cell Types and Connections modules are m:n relationships with the objects Collator and Reference because a collator may insert data about many cell types and projections, and information associated with a cell type or projection may be provided by many collators. "Relations" is the fourth module of BAMS and consists of three parts, "Hierarchy", "Topology", and "Nomenclature". The Hierarchy part of BAMS refers to the set of tables and relationships that allow collators to construct ordered sets of brain parts according to different criteria used to organize brain nomenclatures (different taxonomies) inserted in the system. Topology refers to inserted or inferred topological relationships between brain regions defined in different brain nomenclatures. Nomenclature refers to the set of relationships established in BAMS between pairs of brain structures in different neuroanatomical nomenclatures. It includes two types of relationships that can be inserted or inferred in BAMS: identical names, and common sets of references used to identify brain parts. (GIF 11 kb)
Rights and permissions
About this article
Cite this article
Bota, M., Dong, HW. & Swanson, L. From gene networks to brain networks. Nat Neurosci 6, 795–799 (2003). https://doi.org/10.1038/nn1096
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn1096
This article is cited by
-
Effects of Antibodies to Glutamate on Cerebral Expression of the Tnfrsf1A Gene under Conditions of Spatial Amnesia Induced by Proinflammatory Protein S100A9 Fibrils in Aging Mice
Bulletin of Experimental Biology and Medicine (2021)
-
Diverse Community Structures in the Neuronal-Level Connectome of the Drosophila Brain
Neuroinformatics (2020)
-
Computational neuroscience and neuroinformatics: Recent progress and resources
Journal of Biosciences (2018)