Abstract
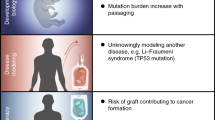
The genomic instability of stem cells in culture, caused by their routine in vitro propagation or by their genetic manipulation, is deleterious both for their clinical application and for their use in basic research. Frequent evaluation of the genomic integrity of stem cells is thus required, and it is usually performed using cytogenetic or DNA-based methods at variable sensitivities, resolutions and costs. Here we present a detailed protocol for determining the genomic integrity of pluripotent stem cells (PSCs) using their global gene expression profiles. This expression-based karyotyping (e-karyotyping) protocol uses gene expression microarray data (either originally generated or derived from the literature) and describes how to organize it properly, subject it to two complementary bioinformatic analyses and conservatively interpret the results in order to generate an accurate estimation of the chromosomal aberrations in the autosomal genome of examined stem cell lines. The experimental steps of e-karyotyping can be carried out in ∼20–30 h.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Ronen, D. & Benvenisty, N. Genomic stability in reprogramming. Curr. Opin. Genet. Dev. 22, 444–449 (2012).
Goldring, C.E. et al. Assessing the safety of stem cell therapeutics. Cell Stem Cell 8, 618–628 (2011).
Lund, R.J., Narva, E. & Lahesmaa, R. Genetic and epigenetic stability of human pluripotent stem cells. Nat. Rev. Genet. 13, 732–744 (2012).
Ben-David, U. & Benvenisty, N. Analyzing the gnomic integrity of stem cells. in StemBook (ed. The Stem Cell Research Community). <http://www.stembook.org/node/719> (2012).
Ben-David, U., Benvenisty, N. & Mayshar, Y. Genetic instability in human induced pluripotent stem cells: classification of causes and possible safeguards. Cell Cycle 9, 4603–4604 (2010).
Barrilleaux, B. & Knoepfler, P.S. Inducing iPSCs to escape the dish. Cell Stem Cell 9, 103–111 (2011).
Liu, X. et al. Trisomy eight in ES cells is a common potential problem in gene targeting and interferes with germ line transmission. Dev. Dyn. 209, 85–91 (1997).
Mayshar, Y. et al. Identification and classification of chromosomal aberrations in human induced pluripotent stem cells. Cell Stem Cell 7, 521–531 (2010).
Ben-David, U., Mayshar, Y. & Benvenisty, N. Large-scale analysis reveals acquisition of lineage-specific chromosomal aberrations in human adult stem cells. Cell Stem Cell 9, 97–102 (2011).
Ben-David, U. & Benvenisty, N. High prevalence of evolutionarily conserved and species-specific genomic aberrations in mouse pluripotent stem cells. Stem Cells 30, 612–622 (2012).
Meisner, L.F. & Johnson, J.A. Protocols for cytogenetic studies of human embryonic stem cells. Methods 45, 133–141 (2008).
Speicher, M.R. & Carter, N.P. The new cytogenetics: blurring the boundaries with molecular biology. Nat. Rev. Genet. 6, 782–792 (2005).
Baker, D.E. et al. Adaptation to culture of human embryonic stem cells and oncogenesis in vivo. Nat. Biotechnol. 25, 207–215 (2007).
Martins-Taylor, K. et al. Recurrent copy number variations in human induced pluripotent stem cells. Nat. Biotechnol. 29, 488–491 (2011).
Spits, C. et al. Recurrent chromosomal abnormalities in human embryonic stem cells. Nat. Biotechnol. 26, 1361–1363 (2008).
Laurent, L.C. et al. Dynamic changes in the copy number of pluripotency and cell proliferation genes in human ESCs and iPSCs during reprogramming and time in culture. Cell Stem Cell 8, 106–118 (2011).
Lefort, N., Perrier, A.L., Laabi, Y., Varela, C. & Peschanski, M. Human embryonic stem cells and genomic instability. Regen. Med. 4, 899–909 (2009).
Quinlan, A.R. et al. Genome sequencing of mouse induced pluripotent stem cells reveals retroelement stability and infrequent DNA rearrangement during reprogramming. Cell Stem Cell 9, 366–373 (2011).
Hussein, S.M. et al. Copy number variation and selection during reprogramming to pluripotency. Nature 471, 58–62 (2011).
Abyzov, A. et al. Somatic copy number mosaicism in human skin revealed by induced pluripotent stem cells. Nature 492, 438–442 (2012).
Bruck, T. & Benvenisty, N. Meta-analysis of the heterogeneity of X chromosome inactivation in human pluripotent stem cells. Stem Cell Res. 6, 187–193 (2011).
Stelzer, Y., Yanuka, O. & Benvenisty, N. Global analysis of parental imprinting in human parthenogenetic induced pluripotent stem cells. Nat. Struct. Mol. Biol. 18, 735–741 (2011).
Lingjaerde, O.C., Baumbusch, L.O., Liestol, K., Glad, I.K. & Borresen-Dale, A.L. CGH-Explorer: a program for analysis of array-CGH data. Bioinformatics 21, 821–822 (2005).
Acknowledgements
We thank T. Golan-Lev for her assistance with graphic design. N.B. is the Herbert Cohn Chair in Cancer Research. U.B.-D. is a Clore Fellow. Y.M. is an EMBO fellow. The research was partially funded by the Israel Science Foundation (grant no. 269/12) and by the Centers of Excellence Legacy Heritage Biomedical Science Partnership (grant no. 1801/10).
Author information
Authors and Affiliations
Contributions
All authors contributed to the development of the methodology. U.B.-D. and N.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Lists of probe sets expressed in human pluripotent stem cells for selected Affymetrix platforms.
These probe sets are expressed in undifferentiated hPSCs, and can be used for the analysis of the genomic integrity of this cell type, when using Affymetrix microarray platforms HG_U133Plus2.0 and HG_ST1.0. (XLSX 281 kb)
Rights and permissions
About this article
Cite this article
Ben-David, U., Mayshar, Y. & Benvenisty, N. Virtual karyotyping of pluripotent stem cells on the basis of their global gene expression profiles. Nat Protoc 8, 989–997 (2013). https://doi.org/10.1038/nprot.2013.051
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.051
This article is cited by
-
EpiTyping: analysis of epigenetic aberrations in parental imprinting and X-chromosome inactivation using RNA-seq
Nature Protocols (2023)
-
Functional in vivo and in vitro effects of 20q11.21 genetic aberrations on hPSC differentiation
Scientific Reports (2020)
-
Genomic evolution of cancer models: perils and opportunities
Nature Reviews Cancer (2019)
-
Genetic and transcriptional evolution alters cancer cell line drug response
Nature (2018)
-
Patient-derived xenografts undergo mouse-specific tumor evolution
Nature Genetics (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.