Key Points
-
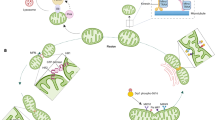
Mitochondria are dynamic organelles. They continually fuse and divide, are actively recruited to specific cellular locations and have dynamic structures.
-
Mitochondrial fusion requires three large GTPases: the outer membrane proteins MFN1 and MFN2, and the inner membrane protein OPA1.
-
Mitochondrial fission requires the dynamin GTPase DRP1 and the outer membrane protein FIS1.
-
The fusion and fission of mitochondria have several important functions. These processes control the morphology of mitochondria, allow content exchange between mitochondria, control mitochondrial distribution and facilitate the release of intermembrane space proteins during apoptosis.
-
Several structural changes in mitochondria are important for rapid and efficient apoptosis: the mitochondria must be fragmented, their outer membranes must become permeable and the cristae junctions must be widened.
-
Mitochondrial dynamics is particularly important to neurons, and defects result in neurodegenerative disease.
Abstract
Recent findings have sparked renewed appreciation for the remarkably dynamic nature of mitochondria. These organelles constantly fuse and divide, and are actively transported to specific subcellular locations. These dynamic processes are essential for mammalian development, and defects lead to neurodegenerative disease. But what are the molecular mechanisms that control mitochondrial dynamics, and why are they important for mitochondrial function? We review these issues and explore how defects in mitochondrial dynamics might cause neuronal disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Chan, D. C. Mitochondrial fusion and fission in mammals. Annu. Rev. Cell Dev. Biol. 22, 79–99 (2006).
Okamoto, K. & Shaw, J. M. Mitochondrial morphology and dynamics in yeast and multicellular eukaryotes. Annu. Rev. Genet. 39, 503–36 (2005).
Hollenbeck, P. J. & Saxton, W. M. The axonal transport of mitochondria. J. Cell Sci. 118, 5411–5419 (2005).
Nunnari, J. et al. Mitochondrial transmission during mating in Saccharomyces cerevisiae is determined by mitochondrial fusion and fission and the intramitochondrial segregation of mitochondrial DNA. Mol. Biol. Cell 8, 1233–1242 (1997).
Bleazard, W. et al. The dynamin-related GTPase Dnm1 regulates mitochondrial fission in yeast. Nature Cell Biol. 1, 298–304 (1999).
Chen, H. et al. Mitofusins Mfn1 and Mfn2 coordinately regulate mitochondrial fusion and are essential for embryonic development. J. Cell Biol. 160, 189–200 (2003).
Sesaki, H. & Jensen, R. E. Division versus fusion: Dnm1p and Fzo1p antagonistically regulate mitochondrial shape. J. Cell Biol. 147, 699–706 (1999).
Smirnova, E., Griparic, L., Shurland, D. L. & van der Bliek, A. M. Dynamin-related protein Drp1 is required for mitochondrial division in mammalian cells. Mol. Biol. Cell 12, 2245–2256 (2001).
Hales, K. G. & Fuller, M. T. Developmentally regulated mitochondrial fusion mediated by a conserved, novel, predicted GTPase. Cell 90, 121–129 (1997). This study identified the first component of the mitochondrial fusion machinery, thereby providing an avenue to identify new components in further yeast genetic screens.
Fehrenbacher, K. L., Yang, H. C., Gay, A. C., Huckaba, T. M. & Pon, L. A. Live cell imaging of mitochondrial movement along actin cables in budding yeast. Curr. Biol. 14, 1996–2004 (2004).
Morris, R. L. & Hollenbeck, P. J. Axonal transport of mitochondria along microtubules and F-actin in living vertebrate neurons. J. Cell Biol. 131, 1315–1326 (1995).
Ligon, L. A. & Steward, O. Role of microtubules and actin filaments in the movement of mitochondria in the axons and dendrites of cultured hippocampal neurons. J. Comp. Neurol. 427, 351–361 (2000).
Li, Z., Okamoto, K., Hayashi, Y. & Sheng, M. The importance of dendritic mitochondria in the morphogenesis and plasticity of spines and synapses. Cell 119, 873–887 (2004).
Miller, K. E. & Sheetz, M. P. Axonal mitochondrial transport and potential are correlated. J. Cell Sci. 117, 2791–2804 (2004).
Pilling, A. D., Horiuchi, D., Lively, C. M. & Saxton, W. M. Kinesin-1 and dynein are the primary motors for fast transport of mitochondria in Drosophila motor axons. Mol. Biol. Cell 17, 2057–2068 (2006).
Morris, R. L. & Hollenbeck, P. J. The regulation of bidirectional mitochondrial transport is coordinated with axonal outgrowth. J. Cell Sci. 104, 917–927 (1993).
Chang, D. T., Honick, A. S. & Reynolds, I. J. Mitochondrial trafficking to synapses in cultured primary cortical neurons. J. Neurosci. 26, 7035–7045 (2006).
Chada, S. R. & Hollenbeck, P. J. Nerve growth factor signaling regulates motility and docking of axonal mitochondria. Curr. Biol. 14, 1272–1276 (2004).
Mannella, C. A. Structure and dynamics of the mitochondrial inner membrane cristae. Biochim. Biophys. Acta 1763, 542–548 (2006).
Scorrano, L. et al. A distinct pathway remodels mitochondrial cristae and mobilizes cytochrome c during apoptosis. Dev. Cell 2, 55–67 (2002).
Gilkerson, R. W., Selker, J. M. & Capaldi, R. A. The cristal membrane of mitochondria is the principal site of oxidative phosphorylation. FEBS Lett. 546, 355–358 (2003).
Vogel, F., Bornhovd, C., Neupert, W. & Reichert, A. S. Dynamic subcompartmentalization of the mitochondrial inner membrane. J. Cell Biol. 175, 237–247 (2006).
Wurm, C. A. & Jakobs, S. Differential protein distributions define two sub-compartments of the mitochondrial inner membrane in yeast. FEBS Lett. 580, 5628–5634 (2006).
Hermann, G. J. et al. Mitochondrial fusion in yeast requires the transmembrane GTPase Fzo1p. J. Cell Biol. 143, 359–373 (1998).
Shaw, J. M. & Nunnari, J. Mitochondrial dynamics and division in budding yeast. Trends Cell Biol. 12, 178–184 (2002).
Meeusen, S., McCaffery, J. M. & Nunnari, J. Mitochondrial fusion intermediates revealed in vitro. Science 305, 1747–1752 (2004). This study described an in vitro fusion assay that identifies mitochondrial fusion intermediates.
Detmer, S. A. & Chan, D. C. Complementation between mouse Mfn1 and Mfn2 protects mitochondrial fusion defects caused by CMT2A disease mutations. J. Cell Biol. 176, 405–414 (2007).
Chen, H., Chomyn, A. & Chan, D. C. Disruption of fusion results in mitochondrial heterogeneity and dysfunction. J. Biol. Chem. 280, 26185–26192 (2005).
Koshiba, T. et al. Structural basis of mitochondrial tethering by mitofusin complexes. Science 305, 858–862 (2004).
Meeusen, S. et al. Mitochondrial inner-membrane fusion and crista maintenance requires the dynamin-related GTPase Mgm1. Cell 127, 383–395 (2006).
Cipolat, S., Martins de Brito, O., Dal Zilio, B. & Scorrano, L. OPA1 requires mitofusin 1 to promote mitochondrial fusion. Proc. Natl Acad. Sci. USA 101, 15927–15932 (2004).
Legros, F., Lombes, A., Frachon, P. & Rojo, M. Mitochondrial fusion in human cells is efficient, requires the inner membrane potential, and is mediated by mitofusins. Mol. Biol. Cell 13, 4343–4354 (2002).
Malka, F. et al. Separate fusion of outer and inner mitochondrial membranes. EMBO Rep. 6, 853–859 (2005).
Ishihara, N., Fujita, Y., Oka, T. & Mihara, K. Regulation of mitochondrial morphology through proteolytic cleavage of OPA1. EMBO J. 25, 2966–2977 (2006).
Altmann, K. & Westermann, B. Role of essential genes in mitochondrial morphogenesis in Saccharomyces cerevisiae. Mol. Biol. Cell 16, 5410–5417 (2005).
Dimmer, K. S. et al. Genetic basis of mitochondrial function and morphology in Saccharomyces cerevisiae. Mol. Biol. Cell 13, 847–853 (2002).
Choi, S. Y. et al. A common lipid links Mfn-mediated mitochondrial fusion and SNARE-regulated exocytosis. Nature Cell Biol. 8, 1255–1262 (2006).
Fratti, R. A., Jun, Y., Merz, A. J., Margolis, N. & Wickner, W. Interdependent assembly of specific regulatory lipids and membrane fusion proteins into the vertex ring domain of docked vacuoles. J. Cell Biol. 167, 1087–1098 (2004).
Vitale, N. et al. Phospholipase D1: a key factor for the exocytotic machinery in neuroendocrine cells. EMBO J. 20, 2424–2434 (2001).
Ingerman, E. et al. Dnm1 forms spirals that are structurally tailored to fit mitochondria. J. Cell Biol. 170, 1021–1027 (2005).
Bhar, D., Karren, M. A., Babst, M. & Shaw, J. M. Dimeric Dnm1-G385D interacts with Mdv1 on mitochondria and can be stimulated to assemble into fission complexes containing Mdv1 and Fis1. J. Biol. Chem. 281, 17312–17320 (2006).
Fekkes, P., Shepard, K. A. & Yaffe, M. P. Gag3p, an outer membrane protein required for fission of mitochondrial tubules. J. Cell Biol. 151, 333–340 (2000).
Mozdy, A. D., McCaffery, J. M. & Shaw, J. M. Dnm1p GTPase-mediated mitochondrial fission is a multi-step process requiring the novel integral membrane component Fis1p. J. Cell Biol. 151, 367–380 (2000).
Tieu, Q. & Nunnari, J. Mdv1p is a WD repeat protein that interacts with the dynamin-related GTPase, Dnm1p, to trigger mitochondrial division. J. Cell Biol. 151, 353–366 (2000). References 42–44 showed the power of yeast genetic screens for identifying essential components of the mitochondrial fission machinery.
Griffin, E. E., Graumann, J. & Chan, D. C. The WD40 protein Caf4p is a component of the mitochondrial fission machinery and recruits Dnm1p to mitochondria. J. Cell Biol. 170, 237–248 (2005).
Lee, Y. J., Jeong, S. Y., Karbowski, M., Smith, C. L. & Youle, R. J. Roles of the mammalian mitochondrial fission and fusion mediators Fis1, Drp1, and Opa1 in apoptosis. Mol. Biol. Cell 15, 5001–5011 (2004).
Koch, A. et al. Dynamin-like protein 1 is involved in peroxisomal fission. J. Biol. Chem. 278, 8597–8605 (2003).
Koch, A., Yoon, Y., Bonekamp, N. A., McNiven, M. A. & Schrader, M. A role for Fis1 in both mitochondrial and peroxisomal fission in mammalian cells. Mol. Biol. Cell 16, 5077–5086 (2005).
Escobar-Henriques, M., Westermann, B. & Langer, T. Regulation of mitochondrial fusion by the F-box protein Mdm30 involves proteasome-independent turnover of Fzo1. J. Cell Biol. 173, 645–650 (2006).
Neutzner, A. & Youle, R. J. Instability of the mitofusin Fzo1 regulates mitochondrial morphology during the mating response of the yeast Saccharomyces cerevisiae. J. Biol. Chem. 280, 18598–18603 (2005).
Karbowski, M., Neutzner, A. & Youle, R. J. The mitochondrial E3 ubiquitin ligase MARCH5 is required for Drp1 dependent mitochondrial division. J. Cell Biol. 178, 71–84 (2007).
Nakamura, N., Kimura, Y., Tokuda, M., Honda, S. & Hirose, S. MARCH-V is a novel mitofusin 2- and Drp1-binding protein able to change mitochondrial morphology. EMBO Rep. 7, 1019–1022 (2006).
Eura, Y., Ishihara, N., Oka, T. & Mihara, K. Identification of a novel protein that regulates mitochondrial fusion by modulating mitofusin (Mfn) protein function. J. Cell Sci. 119, 4913–4925 (2006).
Wasiak, S., Zunino, R. & McBride, H. M. Bax/Bak promote sumoylation of DRP1 and its stable association with mitochondria during apoptotic cell death. J. Cell Biol. 177, 439–450 (2007).
Taguchi, N., Ishihara, N., Jofuku, A., Oka, T. & Mihara, K. Mitotic phosphorylation of dynamin-related GTPase Drp1 participates in mitochondrial fission. J. Biol. Chem. 282, 11521–11529 (2007).
Meisinger, C. et al. The morphology proteins Mdm12/Mmm1 function in the major β-barrel assembly pathway of mitochondria. EMBO J. 26, 2229–2239 (2007).
Guo, X. et al. The GTPase dMiro is required for axonal transport of mitochondria to Drosophila synapses. Neuron 47, 379–393 (2005).
Stowers, R. S., Megeath, L. J., Gorska-Andrzejak, J., Meinertzhagen, I. A. & Schwarz, T. L. Axonal transport of mitochondria to synapses depends on milton, a novel Drosophila protein. Neuron 36, 1063–1077 (2002).
Glater, E. E., Megeath, L. J., Stowers, R. S. & Schwarz, T. L. Axonal transport of mitochondria requires milton to recruit kinesin heavy chain and is light chain independent. J. Cell Biol. 173, 545–557 (2006).
Frederick, R. L., McCaffery, J. M., Cunningham, K. W., Okamoto, K. & Shaw, J. M. Yeast Miro GTPase, Gem1p, regulates mitochondrial morphology via a novel pathway. J. Cell Biol. 167, 87–98 (2004).
Sesaki, H., Southard, S. M., Yaffe, M. P. & Jensen, R. E. Mgm1p, a dynamin-related GTPase, is essential for fusion of the mitochondrial outer membrane. Mol. Biol. Cell 14, 2342–2356 (2003).
Olichon, A. et al. Loss of OPA1 perturbates the mitochondrial inner membrane structure and integrity, leading to cytochrome c release and apoptosis. J. Biol. Chem. 278, 7743–7746 (2003).
Amutha, B., Gordon, D. M., Gu, Y. & Pain, D. A novel role of Mgm1p, a dynamin-related GTPase, in ATP synthase assembly and cristae formation/maintenance. Biochem. J. 381, 19–23 (2004).
Frezza, C. et al. OPA1 controls apoptotic cristae remodeling independently from mitochondrial fusion. Cell 126, 177–189 (2006).
Paumard, P. et al. The ATP synthase is involved in generating mitochondrial cristae morphology. EMBO J. 21, 221–230 (2002).
Minauro-Sanmiguel, F., Wilkens, S. & Garcia, J. J. Structure of dimeric mitochondrial ATP synthase: novel F0 bridging features and the structural basis of mitochondrial cristae biogenesis. Proc. Natl Acad. Sci. USA 102, 12356–12358 (2005).
Dudkina, N. V., Heinemeyer, J., Keegstra, W., Boekema, E. J. & Braun, H. P. Structure of dimeric ATP synthase from mitochondria: an angular association of monomers induces the strong curvature of the inner membrane. FEBS Lett. 579, 5769–5772 (2005).
Messerschmitt, M. et al. The inner membrane protein Mdm33 controls mitochondrial morphology in yeast. J. Cell Biol. 160, 553–564 (2003).
John, G. B. et al. The mitochondrial inner membrane protein mitofilin controls cristae morphology. Mol. Biol. Cell 16, 1543–1554 (2005).
Hobbs, A. E., Srinivasan, M., McCaffery, J. M. & Jensen, R. E. Mmm1p, a mitochondrial outer membrane protein, is connected to mitochondrial DNA (mtDNA) nucleoids and required for mtDNA stability. J. Cell Biol. 152, 401–410 (2001).
Dimmer, K. S., Jakobs, S., Vogel, F., Altmann, K. & Westermann, B. Mdm31 and Mdm32 are inner membrane proteins required for maintenance of mitochondrial shape and stability of mitochondrial DNA nucleoids in yeast. J. Cell Biol. 168, 103–115 (2005).
Chen, H., McCaffery, J. M. & Chan, D. C. Mitochondrial fusion protects against neurodegeneration in the cerebellum. Cell 130, 548–562 (2007). This study showed that mitochondrial fusion is essential for respiratory function and that it protects neurons in the cerebellum from degeneration — findings that will probably be relevant for understanding CMT2A pathogenesis.
Legros, F., Malka, F., Frachon, P., Lombes, A. & Rojo, M. Organization and dynamics of human mitochondrial DNA. J. Cell Sci. 117, 2653–2662 (2004).
Alavi, M. V. et al. A splice site mutation in the murine Opa1 gene features pathology of autosomal dominant optic atrophy. Brain 130, 1029–1042 (2007).
Davies, V. J. et al. Opa1 deficiency in a mouse model of autosomal dominant optic atrophy impairs mitochondrial morphology, optic nerve structure and visual function. Hum. Mol. Genet. 16, 1307–1318 (2007).
Labrousse, A. M., Zappaterra, M. D., Rube, D. A. & van der Bliek, A. M. C. elegans dynamin-related protein DRP-1 controls severing of the mitochondrial outer membrane. Mol. Cell 4, 815–826 (1999).
Waterham, H. R. et al. A lethal defect of mitochondrial and peroxisomal fission. N. Engl. J. Med. 356, 1736–1741 (2007). This study provided clinical evidence for the essentiality of mitochondrial fission.
Verstreken, P. et al. Synaptic mitochondria are critical for mobilization of reserve pool vesicles at Drosophila neuromuscular junctions. Neuron 47, 365–378 (2005).
Campello, S. et al. Orchestration of lymphocyte chemotaxis by mitochondrial dynamics. J. Exp. Med. 203, 2879–2886 (2006).
Arnoult, D. Mitochondrial fragmentation in apoptosis. Trends Cell Biol. 17, 6–12 (2007).
Youle, R. J. & Karbowski, M. Mitochondrial fission in apoptosis. Nature Rev. Mol. Cell Biol. 6, 657–663 (2005).
Frank, S. et al. The role of dynamin-related protein 1, a mediator of mitochondrial fission, in apoptosis. Dev. Cell 1, 515–525 (2001). This study provided the first evidence that mitochondrial fission has a significant role in apoptosis.
Parone, P. A. et al. Inhibiting the mitochondrial fission machinery does not prevent Bax/Bak-dependent apoptosis. Mol. Cell. Biol. 26, 7397–7408 (2006).
Abdelwahid, E. et al. Mitochondrial disruption in Drosophila apoptosis. Dev. Cell 12, 793–806 (2007).
Goyal, G., Fell, B., Sarin, A., Youle, R. J. & Sriram, V. Role of mitochondrial remodeling in programmed cell death in Drosophila melanogaster. Dev. Cell 12, 807–816 (2007).
Jagasia, R., Grote, P., Westermann, B. & Conradt, B. DRP-1-mediated mitochondrial fragmentation during EGL-1-induced cell death in C. elegans. Nature 433, 754–760 (2005).
Karbowski, M., Norris, K. L., Cleland, M. M., Jeong, S. Y. & Youle, R. J. Role of Bax and Bak in mitochondrial morphogenesis. Nature 443, 658–662 (2006).
Brooks, C. et al. Bak regulates mitochondrial morphology and pathology during apoptosis by interacting with mitofusins. Proc. Natl Acad. Sci. USA 104, 11649–11654 (2007).
Goldstein, J. C., Waterhouse, N. J., Juin, P., Evan, G. I. & Green, D. R. The coordinate release of cytochrome c during apoptosis is rapid, complete and kinetically invariant. Nature Cell Biol. 2, 156–162 (2000).
Cipolat, S. et al. Mitochondrial rhomboid PARL regulates cytochrome c release during apoptosis via OPA1-dependent cristae remodeling. Cell 126, 163–175 (2006).
Germain, M., Mathai, J. P., McBride, H. M. & Shore, G. C. Endoplasmic reticulum BIK initiates DRP1-regulated remodelling of mitochondrial cristae during apoptosis. EMBO J. 24, 1546–1556 (2005).
Alexander, C. et al. OPA1, encoding a dynamin-related GTPase, is mutated in autosomal dominant optic atrophy linked to chromosome 3q28. Nature Genet. 26, 211–215 (2000).
Delettre, C. et al. Nuclear gene OPA1, encoding a mitochondrial dynamin-related protein, is mutated in dominant optic atrophy. Nature Genet. 26, 207–210 (2000). References 92 and 93 showed that dominant optic atrophy is caused by mutations in OPA1.
Ferre, M., Amati-Bonneau, P., Tourmen, Y., Malthiery, Y. & Reynier, P. eOPA1: an online database for OPA1 mutations. Hum. Mutat. 25, 423–428 (2005).
Olichon, A. et al. Effects of OPA1 mutations on mitochondrial morphology and apoptosis: relevance to ADOA pathogenesis. J. Cell Physiol. 211, 423–430 (2006).
Kim, J. Y. et al. Mitochondrial DNA content is decreased in autosomal dominant optic atrophy. Neurology 64, 966–972 (2005).
Lodi, R. et al. Deficit of in vivo mitochondrial ATP production in OPA1-related dominant optic atrophy. Ann. Neurol. 56, 719–723 (2004).
Griparic, L., van der Wel, N. N., Orozco, I. J., Peters, P. J. & van der Bliek, A. M. Loss of the intermembrane space protein Mgm1/OPA1 induces swelling and localized constrictions along the lengths of mitochondria. J. Biol. Chem. 279, 18792–18798 (2004).
Zuchner, S. & Vance, J. M. Mechanisms of disease: a molecular genetic update on hereditary axonal neuropathies. Nature Clin. Pract. Neurol. 2, 45–53 (2006).
Zuchner, S. et al. Mutations in the mitochondrial GTPase mitofusin 2 cause Charcot-Marie-Tooth neuropathy type 2A. Nature Genet. 36, 449–451 (2004). This study showed that mutations in MFN2 cause the peripheral neuropathy CMT2A.
Zuchner, S. et al. Axonal neuropathy with optic atrophy is caused by mutations in mitofusin 2. Ann. Neurol. 59, 276–281 (2006).
Chung, K. W. et al. Early onset severe and late-onset mild Charcot-Marie-Tooth disease with mitofusin 2 (MFN2) mutations. Brain 129, 2103–2118 (2006).
Verhoeven, K. et al. MFN2 mutation distribution and genotype/phenotype correlation in Charcot-Marie-Tooth type 2. Brain 129, 2093–2102 (2006).
Baloh, R. H., Schmidt, R. E., Pestronk, A. & Milbrandt, J. Altered axonal mitochondrial transport in the pathogenesis of Charcot-Marie-Tooth disease from mitofusin 2 mutations. J. Neurosci. 27, 422–430 (2007).
Niemann, A., Ruegg, M., La Padula, V., Schenone, A. & Suter, U. Ganglioside-induced differentiation associated protein 1 is a regulator of the mitochondrial network: new implications for Charcot-Marie-Tooth disease. J. Cell Biol. 170, 1067–1078 (2005).
Acknowledgements
This work was supported by grants from the National Institutes of Health. D.C.C. is an Ellison Medical Foundation Senior Scholar in Aging.
Author information
Authors and Affiliations
Corresponding author
Related links
Glossary
- Cristae
-
Invaginations of the mitochondrial inner membrane.
- Nebenkern structure
-
A cytosolic structure, found in some insect spermatids, that is formed by the fusion of mitochondria.
- Anterograde
-
The direction from the cell body towards the periphery.
- Retrograde
-
The direction from peripheral regions towards the cell body.
- Oxidative phosphorylation
-
A biochemical pathway for ATP production that results in oxygen consumption and is localized to the mitochondrial cristae.
- Coiled coil
-
A structural motif that is formed by polypeptide sequences that contain hydrophobic heptad repeats.
- Dynamin
-
A large GTPase that is thought to mediate vesicle scission during endocytosis.
- Mitochondrial membrane potential
-
The electrochemical gradient that exists across the mitochondrial inner membrane.
- Ergosterol
-
A steroid compound that is a component of yeast cell membranes and which might have a role similar to that of cholesterol in mammalian cell membranes.
- SNARE
-
(soluble N-ethylmaleimide-sensitive fusion protein (NSF) attachment protein (SNAP) receptor). A highly α-helical protein that mediates the specific fusion of vesicles with target membranes.
- F-box protein
-
A protein containing an F-box motif, a small domain that is used for protein interactions. The best-characterized F-box proteins are components of an E3 ubiquitin ligase, and help in ubiquitin-dependent protein degradation by recognizing specific substrates.
- β-barrel protein
-
A protein composed of a β-sheet that is rolled up into a cylinder. One such mitochondrial β-barrel protein is VDAC (voltage-dependent anion channel), which forms a pore in the outer membrane.
- Kinesin
-
A microtubule-based molecular motor protein that is most often directed towards the plus end of microtubules.
- Dynein
-
A microtubule-based molecular motor that is directed towards the minus end of microtubules.
- EF-hand domain
-
A helix-loop-helix protein motif that can bind a Ca2+ ion.
- Mitochondrial F1F0 ATP synthase
-
A large, multisubunit enzyme embedded in the mitochondrial cristae that uses the proton gradient across the inner membrane to synthesize ATP.
- Mitochondrial DNA
-
(mtDNA). A circular genome (∼16 kb in mammals) located in the mitochondrial matrix that encodes 13 polypeptides of the electron transport chain, 22 tRNAs and 2 rRNAs.
- Nucleoid
-
A compacted mass of DNA. Mitochondrial DNA is organized into nucleoids, each consisting of several mitochondrial genomes.
- Chemotaxis
-
The directed movement of cells in response to a chemical stimulus.
- Mitochondrial outer membrane permeabilization
-
(MOMP). The opening of pores in the mitochondrial outer membrane — an early event during apoptosis that releases apoptotic factors from the mitochondrial intermembrane space.
- Haploinsufficiency
-
A genetic state in diploids in which a single functional copy of a gene is insufficient to maintain a normal phenotype.
- Sural nerve
-
A sensory nerve innervating the calf and foot that is commonly investigated by biopsy for the evaluation of peripheral neuropathies.
Rights and permissions
About this article
Cite this article
Detmer, S., Chan, D. Functions and dysfunctions of mitochondrial dynamics. Nat Rev Mol Cell Biol 8, 870–879 (2007). https://doi.org/10.1038/nrm2275
Issue Date:
DOI: https://doi.org/10.1038/nrm2275
This article is cited by
-
Urolithin A Prevents Sleep-deprivation-induced Neuroinflammation and Mitochondrial Dysfunction in Young and Aged Mice
Molecular Neurobiology (2024)
-
The potential mechanism of gut microbiota-microbial metabolites-mitochondrial axis in progression of diabetic kidney disease
Molecular Medicine (2023)
-
Crosstalk between mitochondrial biogenesis and mitophagy to maintain mitochondrial homeostasis
Journal of Biomedical Science (2023)
-
SIM2s directed Parkin-mediated mitophagy promotes mammary epithelial cell differentiation
Cell Death & Differentiation (2023)
-
Ensemble heterogeneity mimics ageing for endosomal dynamics within eukaryotic cells
Scientific Reports (2023)