Abstract
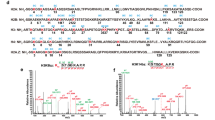
Methylation of histone 3 lysine 4 (H3K4) by yeast Set1-COMPASS requires prior monoubiquitination of histone H2B. To define whether other residues within the histones are also required for H3K4 methylation, we systematically generated a complete library of the alanine substitutions of all of the residues of the four core histones in Saccharomyces cerevisiae. From this study we discovered that 18 residues within the four histones are essential for viability on complete growth media. We also identified several cis-regulatory residues on the histone H3 N-terminal tail, including histone H3 lysine 14 (H3K14), which are required for normal levels of H3K4 trimethylation. Several previously uncharacterized trans-regulatory residues on histones H2A and H2B form a patch on nucleosomes and are required for methylation mediated by COMPASS. This library will be a valuable tool for defining the role of histone residues in processes requiring chromatin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Kornberg, R.D. & Lorch, Y. Twenty-five years of the nucleosome, fundamental particle of the eukaryote chromosome. Cell 98, 285–294 (1999).
Shilatifard, A. Chromatin modifications by methylation and ubiquitination: implications in the regulation of gene expression. Annu. Rev. Biochem. 75, 243–269 (2006).
Jenuwein, T. & Allis, C.D. Translating the histone code. Science 293, 1074–1080 (2001).
Luger, K., Mader, A.W., Richmond, R.K., Sargent, D.F. & Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 resolution. Nature 389, 251–260 (1997).
Mito, Y., Henikoff, J.G. & Henikoff, S. Histone replacement marks the boundaries of cis-regulatory domains. Science 315, 1408–1411 (2007).
Polo, S.E., Roche, D. & Almouzni, G. New histone incorporation marks sites of UV repair in human cells. Cell 127, 481–493 (2006).
Lacoste, N. & Almouzni, G. Epigenetic memory: H3.3 steps in the groove. Nat. Cell Biol. 10, 7–9 (2008).
Bhaumik, S.R., Smith, E. & Shilatifard, A. Covalent modifications of histones during development and disease pathogenesis. Nat. Struct. Mol. Biol. 14, 1008–1016 (2007).
Berger, S.L. Histone modifications in transcriptional regulation. Curr. Opin. Genet. Dev. 12, 142–148 (2002).
van Leeuwen, F., Gafken, P.R. & Gottschling, D.E. Dot1p modulates silencing in yeast by methylation of the nucleosome core. Cell 109, 745–756 (2002).
Ng, H.H. et al. Lysine methylation within the globular domain of histone H3 by Dot1 is important for telomeric silencing and Sir protein association. Genes Dev. 16, 1518–1527 (2002).
Schneider, J., Bajwa, P., Johnson, F.C., Bhaumik, S.R. & Shilatifard, A. Rtt109 is required for proper H3K56 acetylation: a chromatin mark associated with the elongating RNA polymerase II. J. Biol. Chem. 281, 37270–37274 (2006).
Masumoto, H., Hawke, D., Kobayashi, R. & Verreault, A. A role for cell-cycle-regulated histone H3 lysine 56 acetylation in the DNA damage response. Nature 436, 294–298 (2005).
Xu, F., Zhang, K. & Grunstein, M. Acetylation in histone H3 globular domain regulates gene expression in yeast. Cell 121, 375–385 (2005).
Nagy, P.L., Griesenbeck, J., Kornberg, R.D. & Cleary, M.L. A trithorax-group complex purified from Saccharomyces cerevisiae is required for methylation of histone H3. Proc. Natl. Acad. Sci. USA 99, 90–94 (2002).
Krogan, N.J. et al. The Paf1 complex is required for histone H3 methylation by COMPASS and Dot1p: linking transcriptional elongation to histone methylation. Mol. Cell 11, 721–729 (2003).
Schneider, J. et al. Molecular regulation of histone H3 trimethylation by COMPASS and the regulation of gene expression. Mol. Cell 19, 849–856 (2005).
Miller, T. et al. COMPASS: a complex of proteins associated with a trithorax-related SET domain protein. Proc. Natl. Acad. Sci. USA 98, 12902–12907 (2001).
Roguev, A. et al. The Saccharomyces cerevisiae Set1 complex includes an Ash2 homologue and methylates histone 3 lysine 4. EMBO J. 20, 7137–7148 (2001).
Bernstein, B.E. et al. Methylation of histone H3 Lys 4 in coding regions of active genes. Proc. Natl. Acad. Sci. USA 99, 8695–8700 (2002).
Krogan, N.J. et al. COMPASS, a histone H3 (lysine 4) methyltransferase required for telomeric silencing of gene expression. J. Biol. Chem. 277, 10753–10755 (2002).
Krogan, N.J. et al. Methylation of histone H3 by Set2 in Saccharomyces cerevisiae is linked to transcriptional elongation by RNA polymerase II. Mol. Cell. Biol. 23, 4207–4218 (2003).
Berger, S.L. The complex language of chromatin regulation during transcription. Nature 447, 407–412 (2007).
Xiao, T. et al. Phosphorylation of RNA polymerase II CTD regulates H3 methylation in yeast. Genes Dev. 17, 654–663 (2003).
Bernstein, B.E. et al. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120, 169–181 (2005).
Liu, C.L. et al. Single-nucleosome mapping of histone modifications in S. cerevisiae. PLoS Biol. 3, e328 (2005).
Hughes, C.M. et al. Menin associates with a trithorax family histone methyltransferase complex and with the hoxc8 locus. Mol. Cell 13, 587–597 (2004).
Dover, J. et al. Methylation of histone H3 by COMPASS requires ubiquitination of histone H2B by Rad6. J. Biol. Chem. 277, 28368–28371 (2002).
Sun, Z.W. & Allis, C.D. Ubiquitination of histone H2B regulates H3 methylation and gene silencing in yeast. Nature 418, 104–108 (2002).
Kim, J., Hake, S.B. & Roeder, R.G. The human homolog of yeast BRE1 functions as a transcriptional coactivator through direct activator interactions. Mol. Cell 20, 759–770 (2005).
Zhu, B. et al. Monoubiquitination of human histone H2B: the factors involved and their roles in HOX gene regulation. Mol. Cell 20, 601–611 (2005).
Pavri, R. et al. Histone H2B monoubiquitination functions cooperatively with FACT to regulate elongation by RNA polymerase II. Cell 125, 703–717 (2006).
Kirmizis, A. et al. Arginine methylation at histone H3R2 controls deposition of H3K4 trimethylation. Nature 449, 928–932 (2007).
Guccione, E. et al. Methylation of histone H3R2 by PRMT6 and H3K4 by an MLL complex are mutually exclusive. Nature 449, 933–937 (2007).
Matsubara, K., Sano, N., Umehara, T. & Horikoshi, M. Global analysis of functional surfaces of core histones with comprehensive point mutants. Genes Cells 12, 13–33 (2007).
Schneider, J., Dover, J., Johnston, M. & Shilatifard, A. Global proteomic analysis of S. cerevisiae (GPS) to identify proteins required for histone modifications. Methods Enzymol. 377, 227–234 (2004).
Carrozza, M.J. et al. Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123, 581–592 (2005).
Kayne, P.S. et al. Extremely conserved histone H4 N terminus is dispensable for growth but essential for repressing the silent mating loci in yeast. Cell 55, 27–39 (1988).
Kruger, W. et al. Amino acid substitutions in the structured domains of histones H3 and H4 partially relieve the requirement of the yeast SWI/SNF complex for transcription. Genes Dev. 9, 2770–2779 (1995).
Luger, K. Structure and dynamic behavior of nucleosomes. Curr. Opin. Genet. Dev. 13, 127–135 (2003).
Muthurajan, U.M. et al. Crystal structures of histone Sin mutant nucleosomes reveal altered protein-DNA interactions. EMBO J. 23, 260–271 (2004).
Park, J.H., Cosgrove, M.S., Youngman, E., Wolberger, C. & Boeke, J.D. A core nucleosome surface crucial for transcriptional silencing. Nat. Genet. 32, 273–279 (2002).
Zhu, B. et al. The human PAF complex coordinates transcription with events downstream of RNA synthesis. Genes Dev. 19, 1668–1673 (2005).
Robzyk, K., Recht, J. & Osley, M.A. Rad6-dependent ubiquitination of histone H2B in yeast. Science 287, 501–504 (2000).
Wood, A. et al. Bre1, an E3 ubiquitin ligase required for recruitment and substrate selection of Rad6 at a promoter. Mol. Cell 11, 267–274 (2003).
Hwang, W.W. et al. A conserved RING finger protein required for histone H2B monoubiquitination and cell size control. Mol. Cell 11, 261–266 (2003).
Wood, A., Schneider, J., Dover, J., Johnston, M. & Shilatifard, A. The Paf1 complex is essential for histone monoubiquitination by the Rad6-Bre1 complex, which signals for histone methylation by COMPASS and Dot1p. J. Biol. Chem. 278, 34739–34742 (2003).
Wood, A., Schneider, J., Dover, J., Johnston, M. & Shilatifard, A. The Bur1/Bur2 complex is required for histone H2B monoubiquitination by Rad6/Bre1 and histone methylation by COMPASS. Mol. Cell 20, 589–599 (2005).
Wood, A., Schneider, J. & Shilatifard, A. Cross-talking histones: implications for the regulation of gene expression and DNA repair. Biochem. Cell Biol. 83, 460–467 (2005).
Wood, A. et al. Ctk complex-mediated regulation of histone methylation by COMPASS. Mol. Cell. Biol. 27, 709–720 (2007).
Laribee, R.N. et al. BUR kinase selectively regulates H3 K4 trimethylation and H2B ubiquitylation through recruitment of the PAF elongation complex. Curr. Biol. 15, 1487–1493 (2005).
Xiao, T. et al. The RNA polymerase II kinase Ctk1 regulates positioning of a 5′ histone methylation boundary along genes. Mol. Cell. Biol. 27, 721–731 (2007).
Lee, J.S. et al. Histone crosstalk between H2B monoubiquitination and H3 methylation mediated by COMPASS. Cell 131, 1084–1096 (2007).
Taverna, S.D., Li, H., Ruthenburg, A.J., Allis, C.D. & Patel, D.J. How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat. Struct. Mol. Biol. 14, 1025–1040 (2007).
Howe, L. et al. Histone H3 specific acetyltransferases are essential for cell cycle progression. Genes Dev. 15, 3144–3154 (2001).
Govind, C.K., Zhang, F., Qiu, H., Hofmeyer, K. & Hinnebusch, A.G. Gcn5 promotes acetylation, eviction, and methylation of nucleosomes in transcribed coding regions. Mol. Cell 25, 31–42 (2007).
Martin, D.G. et al. The Yng1p plant homeodomain finger is a methyl-histone binding module that recognizes lysine 4-methylated histone H3. Mol. Cell. Biol. 26, 7871–7879 (2006).
Taverna, S.D. et al. Yng1 PHD finger binding to H3 trimethylated at K4 promotes NuA3 HAT activity at K14 of H3 and transcription at a subset of targeted ORFs. Mol. Cell 24, 785–796 (2006).
Kasten, M. et al. Tandem bromodomains in the chromatin remodeler RSC recognize acetylated histone H3 Lys14. EMBO J. 23, 1348–1359 (2004).
White, C.L., Suto, R.K. & Luger, K. Structure of the yeast nucleosome core particle reveals fundamental changes in internucleosome interactions. EMBO J. 20, 5207–5218 (2001).
Acknowledgements
We thank L. Shilatifard for editorial assistance and E. Smith for critical reading of the manuscript. We are grateful for M. Ruhlman and N. Dillon for large-scale preparation of polyacrylamide gels, and C. Wimberly and H. Strobietto for large-scale plasmid preparation and DNA sequencing. We are also grateful to B. Li from the Workman laboratory at the Stowers Institute for providing YBL574 and Y131 strains and shuffling plasmids. This work was supported by the US National Institute of Health grants 2R01CA89455 and 2R01GM069905 to A.S.
Author information
Authors and Affiliations
Contributions
S.N. and A.S. carried out the research and wrote the manuscript; B.W.S. and K.M.D. performed site-directed mutagenesis of histones and validation of the generated vectors; W.D.B. performed validation of the transformed yeast library; K.S.-H. and A.S. provided support and advice.
Corresponding author
Rights and permissions
About this article
Cite this article
Nakanishi, S., Sanderson, B., Delventhal, K. et al. A comprehensive library of histone mutants identifies nucleosomal residues required for H3K4 methylation. Nat Struct Mol Biol 15, 881–888 (2008). https://doi.org/10.1038/nsmb.1454
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1454
This article is cited by
-
The histone H2B Arg95 residue efficiently recruits the transcription factor Spt16 to mediate Ste5 expression of the pheromone response pathway
Scientific Reports (2023)
-
Diverse and dynamic forms of gene regulation by the S. cerevisiae histone methyltransferase Set1
Current Genetics (2023)
-
The histone H2B Arg95 residue links the pheromone response pathway to rapamycin-induced G1 arrest in yeast
Scientific Reports (2022)
-
A CRISPR-Cas9 based shuffle system for endogenous histone H3 and H4 combinatorial mutagenesis
Scientific Reports (2021)
-
Systematic genetic and proteomic screens during gametogenesis identify H2BK34 methylation as an evolutionary conserved meiotic mark
Epigenetics & Chromatin (2020)