Abstract
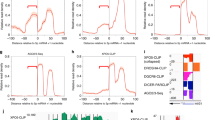
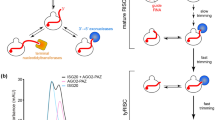
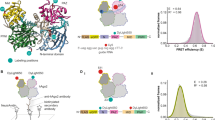
MicroRNAs (miRNAs) regulate expression of their target mRNAs through the RNA-induced silencing complex (RISC), which contains an Argonaute (Ago) family protein as a core component. In Drosophila melanogaster, miRNAs are generally sorted into Ago1-containing RISC (Ago1-RISC). We established a native gel system that can biochemically dissect the Ago1-RISC assembly pathway. We found that miRNA-miRNA* duplexes are loaded into Ago1 as double-stranded RNAs in an ATP-dependent fashion. In contrast, unexpectedly, unwinding of miRNA-miRNA* duplexes is a passive process that does not require ATP or slicer activity of Ago1. Central mismatches direct miRNA-miRNA* duplexes into pre-Ago1-RISC, whereas mismatches in the seed or guide strand positions 12–15 promote conversion of pre-Ago1-RISC into mature Ago1-RISC. Our findings show that unwinding of miRNAs is a precise mirror-image process of target recognition, and both processes reflect the unique geometry of RNAs in Ago proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Tomari, Y. & Zamore, P.D. Perspective: machines for RNAi. Genes Dev. 19, 517–529 (2005).
Filipowicz, W. RNAi: the nuts and bolts of the RISC machine. Cell 122, 17–20 (2005).
Okamura, K., Ishizuka, A., Siomi, H. & Siomi, M.C. Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev. 18, 1655–1666 (2004).
Förstemann, K., Horwich, M.D., Wee, L., Tomari, Y. & Zamore, P.D. Drosophila microRNAs are sorted into functionally distinct Argonaute complexes after production by Dicer-1. Cell 130, 287–297 (2007).
Tomari, Y., Du, T. & Zamore, P.D. Sorting of Drosophila small silencing RNAs. Cell 130, 299–308 (2007).
Kawamura, Y. et al. Drosophila endogenous small RNAs bind to Argonaute2 in somatic cells. Nature 453, 793–797 (2008).
Ghildiyal, M. et al. Endogenous siRNAs derived from transposons and mRNAs in Drosophila somatic cells. Science 320, 1077–1081 (2008).
Azuma-Mukai, A. et al. Characterization of endogenous human Argonautes and their miRNA partners in RNA silencing. Proc. Natl. Acad. Sci. USA 105, 7964–7969 (2008).
Su, H., Trombly, M.I., Chen, J. & Wang, X. Essential and overlapping functions for mammalian Argonautes in microRNA silencing. Genes Dev. 23, 304–317 (2009).
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004).
Meister, G. et al. Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol. Cell 15, 185–197 (2004).
Liu, Q. et al. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 301, 1921–1925 (2003).
Tomari, Y. et al. RISC assembly defects in the Drosophila RNAi mutant armitage. Cell 116, 831–841 (2004).
Pham, J.W., Pellino, J.L., Lee, Y.S., Carthew, R.W. & Sontheimer, E.J.A. Dicer-2-dependent 80S complex cleaves targeted mRNAs during RNAi in Drosophila. Cell 117, 83–94 (2004).
Förstemann, K. et al. Normal microRNA maturation and germ-line stem cell maintenance requires Loquacious, a double-stranded RNA-binding domain protein. PLoS Biol. 3, e236 (2005).
Saito, K., Ishizuka, A., Siomi, H. & Siomi, M.C. Processing of pre-microRNAs by the Dicer-1-Loquacious complex in Drosophila cells. PLoS Biol. 3, e235 (2005).
Liu, X. et al. Dicer-1, but not Loquacious, is critical for assembly of miRNA-induced silencing complexes. RNA 13, 2324–2329 (2007).
Jin, Z. & Xie, T. Dcr-1 maintains Drosophila ovarian stem cells. Curr. Biol. 17, 539–544 (2007).
Nykänen, A., Haley, B. & Zamore, P.D. ATP requirements and small interfering RNA structure in the RNA interference pathway. Cell 107, 309–321 (2001).
Rand, T.A., Petersen, S., Du, F. & Wang, X. Argonaute2 cleaves the anti-guide strand of siRNA during RISC activation. Cell 123, 621–629 (2005).
Matranga, C., Tomari, Y., Shin, C., Bartel, D.P. & Zamore, P.D. Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell 123, 607–620 (2005).
Miyoshi, K., Tsukumo, H., Nagami, T., Siomi, H. & Siomi, M.C. Slicer function of Drosophila Argonautes and its involvement in RISC formation. Genes Dev. 19, 2837–2848 (2005).
Leuschner, P.J., Ameres, S.L., Kueng, S. & Martinez, J. Cleavage of the siRNA passenger strand during RISC assembly in human cells. EMBO Rep. 7, 314–320 (2006).
Lee, Y.S. et al. Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell 117, 69–81 (2004).
Chan, S.P., Ramaswamy, G., Choi, E.Y. & Slack, F.J. Identification of specific let-7 microRNA binding complexes in Caenorhabditis elegans. RNA 14, 2104–2114 (2008).
Iwasaki, S., Kawamata, T. & Tomari, Y. Drosophila Argonaute1 and Argonaute2 employ distinct mechanisms for translational repression. Mol. Cell 34, 58–67 (2009).
Eulalio, A., Huntzinger, E. & Izaurralde, E. GW182 interaction with Argonaute is essential for miRNA-mediated translational repression and mRNA decay. Nat. Struct. Mol. Biol. 15, 346–353 (2008).
Khvorova, A., Reynolds, A. & Jayasena, S.D. Functional siRNAs and miRNAs exhibit strand bias. Cell 115, 209–216 (2003).
Schwarz, D.S. et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115, 199–208 (2003).
Grimson, A. et al. microRNA targeting specificity in mammals: determinants beyond seed pairing. Mol. Cell 27, 91–105 (2007).
Doench, J.G., Petersen, C.P. & Sharp, P.A. siRNAs can function as miRNAs. Genes Dev. 17, 438–442 (2003).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Song, J.J., Smith, S.K., Hannon, G.J. & Joshua-Tor, L. Crystal structure of Argonaute and its implications for RISC slicer activity. Science 305, 1434–1437 (2004).
Rivas, F.V. et al. Purified Argonaute2 and an siRNA form recombinant human RISC. Nat. Struct. Mol. Biol. 12, 340–349 (2005).
Ruby, J.G. et al. Evolution, biogenesis, expression, and target predictions of a substantially expanded set of Drosophila microRNAs. Genome Res. 17, 1850–1864 (2007).
Seitz, H., Ghildiyal, M. & Zamore, P.D. Argonaute loading improves the 5′ precision of both microRNAs and their miRNA* strands in flies. Curr. Biol. 18, 147–151 (2008).
Lu, J. et al. The birth and death of microRNA genes in Drosophila. Nat. Genet. 40, 351–355 (2008).
Chung, W.J., Okamura, K., Martin, R. & Lai, E.C. Endogenous RNA Interference provides a somatic defense against Drosophila transposons. Curr. Biol. 18, 795–802 (2008).
Ruby, J.G. et al. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 127, 1193–1207 (2006).
Batista, P.J. et al. PRG-1 and 21U-RNAs interact to form the piRNA complex required for fertility in C. elegans. Mol. Cell 31, 67–78 (2008).
Calabrese, J.M., Seila, A.C., Yeo, G.W. & Sharp, P.A. RNA sequence analysis defines Dicer's role in mouse embryonic stem cells. Proc. Natl. Acad. Sci. USA 104, 18097–18102 (2007).
Tam, O.H. et al. Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature 453, 534–538 (2008).
Baek, D. et al. The impact of microRNAs on protein output. Nature 455, 64–71 (2008).
Babiarz, J.E., Ruby, J.G., Wang, Y., Bartel, D.P. & Blelloch, R. Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes Dev. 22, 2773–2785 (2008).
Ishizuka, A., Siomi, M.C. & Siomi, H. A Drosophila fragile X protein interacts with components of RNAi and ribosomal proteins. Genes Dev. 16, 2497–2508 (2002).
Robb, G.B. & Rana, T.M. RNA helicase A interacts with RISC in human cells and functions in RISC loading. Mol. Cell 26, 523–537 (2007).
Meister, G. et al. Identification of novel Argonaute-associated proteins. Curr. Biol. 15, 2149–2155 (2005).
Höck, J. et al. Proteomic and functional analysis of Argonaute-containing mRNA-protein complexes in human cells. EMBO Rep. 8, 1052–1060 (2007).
Landthaler, M. et al. Molecular characterization of human Argonaute-containing ribonucleoprotein complexes and their bound target mRNAs. RNA 14, 2580–2596 (2008).
Zhou, R. et al. Comparative analysis of Argonaute-dependent small RNA pathways in Drosophila. Mol. Cell 32, 592–599 (2008).
Wang, Y., Sheng, G., Juranek, S., Tuschl, T. & Patel, D.J. Structure of the guide-strand-containing Argonaute silencing complex. Nature 456, 209–213 (2008).
Wang, Y. et al. Structure of an Argonaute silencing complex with a seed-containing guide DNA and target RNA duplex. Nature 456, 921–926 (2008).
Elbashir, S.M., Martinez, J., Patkaniowska, A., Lendeckel, W. & Tuschl, T. Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. EMBO J. 20, 6877–6888 (2001).
Elbashir, S.M., Lendeckel, W. & Tuschl, T. RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev. 15, 188–200 (2001).
Haley, B. & Zamore, P.D. Kinetic analysis of the RNAi enzyme complex. Nat. Struct. Mol. Biol. 11, 599–606 (2004).
Doench, J.G. & Sharp, P.A. Specificity of microRNA target selection in translational repression. Genes Dev. 18, 504–511 (2004).
Brennecke, J., Stark, A., Russell, R.B. & Cohen, S.M. Principles of microRNA-target recognition. PLoS Biol. 3, e85 (2005).
Mallory, A.C. et al. microRNA control of PHABULOSA in leaf development: importance of pairing to the microRNA 5′ region. EMBO J. 23, 3356–3364 (2004).
Haley, B., Tang, G. & Zamore, P.D. In vitro analysis of RNA interference in Drosophila melanogaster. Methods 30, 330–336 (2003).
Acknowledgements
We thank M. Siomi and H. Siomi (Keio University) for the antibody to Ago1, E. Izaurralde (Max Planck Institute) for the antibody to GW182, R. Carthew (Northwestern University) for the dcr-2L811fsX flies, and M. Horwich and P. Zamore (University of Massachusetts) for the pENTR-Ago1 construct. We also thank M. Mitomi and A. Maruyama for technical assistance, S. Iwasaki and members of the Tomari laboratory for constant and helpful discussions, and H. Sasaki and P. Zamore for stimulating discussions. This work was supported in part by a Grant-in-Aid for Young Scientists (A) from the Japan Ministry of Education, Culture, Sports, Science and Technology, and a Carrier Development Award from The International Human Frontier Science Program Organization. Y.T. is a Japan Science and Technology Agency PRESTO researcher.
Author information
Authors and Affiliations
Contributions
T.K. and Y.T. conducted biochemical experiments, H.S. conducted bioinformatic analyses, Y.T. supervised the study, and all authors discussed the results and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Table 1 (PDF 2300 kb)
Rights and permissions
About this article
Cite this article
Kawamata, T., Seitz, H. & Tomari, Y. Structural determinants of miRNAs for RISC loading and slicer-independent unwinding. Nat Struct Mol Biol 16, 953–960 (2009). https://doi.org/10.1038/nsmb.1630
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1630
This article is cited by
-
Unveiling the world of bee microRNAs: computational identification and characterization of pathway genes, conserved microRNAs, and their targets
International Journal of Tropical Insect Science (2024)
-
GKLOMLI: a link prediction model for inferring miRNA–lncRNA interactions by using Gaussian kernel-based method on network profile and linear optimization algorithm
BMC Bioinformatics (2023)
-
Role of miRNA-21 in radiation-induced heart disease
Oncology and Translational Medicine (2023)
-
Unravelling the developmental and functional significance of an ancient Argonaute duplication
Nature Communications (2020)
-
MicroRNAs and Regeneration in Animal Models of CNS Disorders
Neurochemical Research (2020)