Abstract
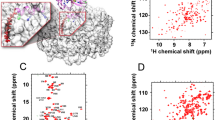
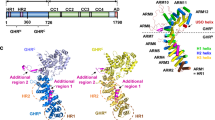
ADP ribosylation factors (Arfs) are N-myristoylated GTP/GDP switch proteins that have key regulatory roles in vesicle transport in eukaryotic cells. ARFs execute their roles by anchoring to membrane surfaces, where they interact with other proteins to initiate budding and maturation of transport vesicles. However, existing structures of Arf•GTP are limited to nonmyristoylated and truncated forms with impaired membrane binding. We report a high-resolution NMR structure for full-length myristoylated yeast (Saccharomyces cerevisiae) Arf1 in complex with a membrane mimic. The two-domain structure, in which the myristoylated N-terminal helix is separated from the C-terminal domain by a flexible linker, suggests a level of adaptability in binding modes for the myriad of proteins with which Arf interacts and allows predictions of specific lipid binding sites on some of these proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kahn, R.A. Toward a model for Arf GTPases as regulators of traffic at the Golgi. FEBS Lett. 583, 3872–3879 (2009).
Luo, R., Ha, V.L., Hayashi, R. & Randazzo, P.A. Arf GAP2 is positively regulated by coatomer and cargo. Cell. Signal. 21, 1169–1179 (2009).
Pucadyil, T.J. & Schmid, S.L. Conserved functions of membrane active GTPases in coated vesicle formation. Science 325, 1217–1220 (2009).
D'Souza-Schorey, C. & Chavrier, P. ARF proteins: roles in membrane traffic and beyond. Nat. Rev. Mol. Cell Biol. 7, 347–358 (2006).
Casanova, J.E. Regulation of arf activation: the sec7 family of guanine nucleotide exchange factors. Traffic 8, 1476–1485 (2007).
Gillingham, A.K. & Munro, S. The small G proteins of the arf family and their regulators. Annu. Rev. Cell Dev. Biol. 23, 579–611 (2007).
Inoue, H. & Randazzo, P.A. Arf GAPs and their interacting proteins. Traffic 8, 1465–1475 (2007).
Renault, L., Guibert, B. & Cherfils, J. Structural snapshots of the mechanism and inhibition of a guanine nucleotide exchange factor. Nature 426, 525–530 (2003).
Shiba, T. et al. Molecular mechanism of membrane recruitment of GGA by ARF in lysosomal protein transport. Nat. Struct. Biol. 10, 386–393 (2003).
Menetrey, J. et al. Structural basis for ARF1-mediated recruitment of ARHGAP21 to Golgi membranes. EMBO J. 26, 1953–1962 (2007).
Amor, J.C., Harrison, D.H., Kahn, R.A. & Ringe, D. Structure of the human ADP-ribosylation factor 1 complexed with GDP. Nature 372, 704–708 (1994).
Goldberg, J. Structural basis for activation of ARF GTPase: mechanisms of guanine nucleotide exchange and GTP-myristoyl switching. Cell 95, 237–248 (1998).
Liu, Y., Kahn, R.A. & Prestegard, J.H. Structure and membrane interaction of myristoylated ARF1. Structure 17, 79–87 (2009).
Amor, J.C. et al. Structures of yeast ARF2 and ARL1: distinct roles for the N terminus in the structure and function of ARF family GTPases. J. Biol. Chem. 276, 42477–42484 (2001).
Liu, Y. & Prestegard, J.H. Direct measurement of dipole-dipole/CSA cross-correlated relaxation by a constant-time experiment. J. Magn. Reson. 193, 23–31 (2008).
Losonczi, J.A. & Prestegard, J.H. Nuclear magnetic resonance characterization of the myristoylated, N-terminal fragment of ADP-ribosylation factor 1 in a magnetically oriented membrane array. Biochemistry 37, 706–716 (1998).
Losonczi, J.A., Tian, F. & Prestegard, J.H. Nuclear magnetic resonance studies of the N-terminal fragment of adenosine diphosphate ribosylation factor 1 in micelles and bicelles: influence of N-myristoylation. Biochemistry 39, 3804–3816 (2000).
Vold, R.R. & Prosser, R.S. Magnetically oriented phospholipid bilayered micelles for structural studies of polypeptides. Does the ideal bicelle exist? J. Magn. Reson. 113, 267–271 (1996).
Lee, D. et al. Bilayer in small bicelles revealed by lipid-protein interactions using NMR spectroscopy. J. Am. Chem. Soc. 130, 13822–13823 (2008).
Fischer, M.W.F., Losonczi, J.A., Weaver, J.L. & Prestegard, J.H. Domain orientation and dynamics in multidomain proteins from residual dipolar couplings. Biochemistry 38, 9013–9022 (1999).
Tolman, J.R. & Ruan, K. NMR residual dipolar couplings as probes of biomolecular dynamics. Chem. Rev. 106, 1720–1736 (2006).
Tolman, J.R., Al-Hashimi, H.M., Kay, L.E. & Prestegard, J.H. Structural and dynamic analysis of residual dipolar coupling data for proteins. J. Am. Chem. Soc. 123, 1416–1424 (2001).
Hubbell, W.L. & Altenbach, C. Investigation of structure and dynamics in membrane-proteins using site-directed spin-labeling. Curr. Opin. Struct. Biol. 4, 566–573 (1994).
Clore, G.M., Tang, C. & Iwahara, J. Elucidating transient macromolecular interactions using paramagnetic relaxation enhancement. Curr. Opin. Struct. Biol. 17, 603–616 (2007).
Tang, C., Iwahara, J. & Clore, G.M. Visualization of transient encounter complexes in protein-protein association. Nature 444, 383–386 (2006).
Iwahara, J., Schwieters, C.D. & Clore, G.M. Ensemble approach for NMR structure refinement against 1H paramagnetic relaxation enhancement data arising from a flexible paramagnetic group attached to a macromolecule. J. Am. Chem. Soc. 126, 5879–5896 (2004).
Solomon, I. & Bloembergen, N. Nuclear magnetic interactions in the Hf molecule. J. Chem. Phys. 25, 261–266 (1956).
Bruschweiler, R. et al. Influence of rapid intramolecular motion on NMR cross-relaxation rates - a molecular-dynamics study of antamanide in solution. J. Am. Chem. Soc. 114, 2289–2302 (1992).
Lipari, G. & Szabo, A. Model-free approach to the interpretation of nuclear magnetic-resonance relaxation in macromolecules. 1. Theory and range of validity. J. Am. Chem. Soc. 104, 4546–4559 (1982).
Schwieters, C.D. & Clore, G.M. Reweighted atomic densities to represent ensembles of NMR structures. J. Biomol. NMR 23, 221–225 (2002).
Ambroggio, E. et al. ArfGAP1 generates an Arf1 gradient on continuous lipid membranes displaying flat and curved regions. EMBO J. 29, 292–303 (2010).
Prestegard, J.H. & Obrien, M.P. Membrane and vesicle fusion. Annu. Rev. Phys. Chem. 38, 383–411 (1987).
Cevc, G. & Richardsen, H. Lipid vesicles and membrane fusion. Adv. Drug Deliv. Rev. 38, 207–232 (1999).
Cukierman, E., Huber, I., Rotman, M. & Cassel, D. The ARF1 GTPase-activating protein: zinc finger motif and Golgi complex localization. Science 270, 1999–2002 (1995).
Godi, A. et al. FAPPs control Golgi-to-cell-surface membrane traffic by binding to ARF and PtdIns(4)P. Nat. Cell Biol. 6, 393–404 (2004).
D'Angelo, G. et al. Glycosphingolipid synthesis requires FAPP2 transfer of glucosylceramide. Nature 449, 62–67 (2007).
Baraldi, E. et al. Structure of the PH domain from Bruton's tyrosine kinase in complex with inositol 1,3,4,5-tetrakisphosphate. Structure 7, 449–460 (1999).
Seidel, R.D., Amor, J.C., Kahn, R.A. & Prestegard, J.H. Structural perturbations in human ADP ribosylation factor-1 accompanying the binding of phosphatidylinositides. Biochemistry 43, 15393–15403 (2004).
Randazzo, P.A. Functional interaction of ADP-ribosylation factor 1 with phosphatidylinositol 4,5-bisphosphate. J. Biol. Chem. 272, 7688–7692 (1997).
Jian, X., Cavenagh, M., Gruschus, J.M., Randazzo, P.A. & Kahn, R.A. Modifications to the C-terminus of Arf1 alter cell functions and protein interactions. Traffic 11, 732–742 (2010).
Cornilescu, G., Marquardt, J.L., Ottiger, M. & Bax, A. Validation of protein structure from anisotropic carbonyl chemical shifts in a dilute liquid crystalline phase. J. Am. Chem. Soc. 120, 6836–6837 (1998).
Battiste, J.L. & Wagner, G. Utilization of site-directed spin labeling and high-resolution heteronuclear nuclear magnetic resonance for global fold determination of large proteins with limited nuclear Overhauser effect data. Biochemistry 39, 5355–5365 (2000).
Liang, B., Bushweller, J.H. & Tamm, L.K. Site-directed parallel spin-labeling and paramagnetic relaxation enhancement in structure determination of membrane proteins by solution NMR spectroscopy. J. Am. Chem. Soc. 128, 4389–4397 (2006).
Liu, Y. & Prestegard, J.H. Measurement of one and two bond N-C couplings in large proteins by TROSY-based J-modulation experiments. J. Magn. Reson. 200, 109–118 (2009).
Permi, P., Tossavainen, H. & Hellman, M. Efficient assignment of methyl resonances: enhanced sensitivity by gradient selection in a DE-MQ-(H)CC(m)Ht (m)-TOCSY experiment. J. Biomol. NMR 30, 275–282 (2004).
Yang, D., Zheng, Y., Liu, D. & Wyss, D.F. Sequence-specific assignments of methyl groups in high-molecular weight proteins. J. Am. Chem. Soc. 126, 3710–3711 (2004).
Cierpicki, T. & Bushweller, J.H. Charged gels as orienting media for measurement of residual dipolar couplings in soluble and integral membrane proteins. J. Am. Chem. Soc. 126, 16259–16266 (2004).
Iwahara, J., Tang, C. & Marius Clore, G. Practical aspects of 1H transverse paramagnetic relaxation enhancement measurements on macromolecules. J. Magn. Reson. 184, 185–195 (2007).
Schwieters, C.D., Kuszewski, J.J. & Clore, G.M. Using Xplor-NIH for NMR molecular structure determination. Prog. Nucl. Magn. Reson. Spectrosc. 48, 47–62 (2006).
Schwieters, C.D. & Clore, G.M. Internal coordinates for molecular dynamics and minimization in structure determination and refinement. J. Magn. Reson. 152, 288–302 (2001).
Acknowledgements
The authors thank M. Demarco (Complex Carbohydrate Research Center, Univ. of Georgia) for kindly providing the model membrane structure and C. Schwieters (Center for Information Technology, US National Institutes of Health) for kindly providing the prerelease version of Xplor-NIH that supports ambiguous RDC assignment. This work was supported by a grant from the US National Institutes of Health (GM61268).
Author information
Authors and Affiliations
Contributions
Y.L. produced the samples and collected and analyzed all data; J.H.P. and R.A.K. devised the project and jointly contributed interpretation of data and drafting of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Methods (PDF 1572 kb)
Rights and permissions
About this article
Cite this article
Liu, Y., Kahn, R. & Prestegard, J. Dynamic structure of membrane-anchored Arf•GTP. Nat Struct Mol Biol 17, 876–881 (2010). https://doi.org/10.1038/nsmb.1853
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1853
This article is cited by
-
Myristic acid as a checkpoint to regulate STING-dependent autophagy and interferon responses by promoting N-myristoylation
Nature Communications (2023)
-
Myr-Arf1 conformational flexibility at the membrane surface sheds light on the interactions with ArfGAP ASAP1
Nature Communications (2023)
-
Revealing the activation mechanism of autoinhibited RalF by integrated simulation and experimental approaches
Scientific Reports (2021)
-
Common architectures in cyanobacteria Prochlorococcus cells visualized by X-ray diffraction imaging using X-ray free electron laser
Scientific Reports (2021)
-
Three distinct regions of cRaf kinase domain interact with membrane
Scientific Reports (2019)