Abstract
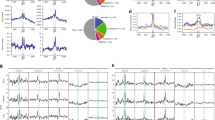
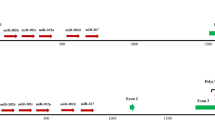
MicroRNAs (miRNAs) are 19–22-nucleotide noncoding RNAs that post-transcriptionally regulate mRNA targets. We have identified endogenous miRNA binding sites in mouse embryonic stem cells (mESCs), by performing photo-cross-linking immunoprecipitation using antibodies to Argonaute (Ago2) followed by deep sequencing of RNAs (CLIP-seq). We also performed CLIP-seq in Dicer−/− mESCs that lack mature miRNAs, allowing us to define whether the association of Ago2 with the identified sites was miRNA dependent. A significantly enriched motif, GCACUU, was identified only in wild-type mESCs in 3′ untranslated and coding regions. This motif matches the seed of a miRNA family that constitutes ~68% of the mESC miRNA population. Unexpectedly, a G-rich motif was enriched in sequences cross-linked to Ago2 in both the presence and absence of miRNAs. Expression analysis and reporter assays confirmed that the seed-related motif confers miRNA-directed regulation on host mRNAs and that the G-rich motif can modulate this regulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
12 May 2011
In the version of this article initially published, the blue curve in Figure 2c was mistakenly replaced with a duplicate of that in Figure 2a. The error has been corrected in the HTML and PDF versions of the article.
References
Ambros, V. The functions of animal microRNAs. Nature 431, 350–355 (2004).
Leung, A.K. & Sharp, P.A. MicroRNA functions in stress responses. Mol. Cell 40, 205–215 (2010).
Stefani, G. & Slack, F.J. Small non-coding RNAs in animal development. Nat. Rev. Mol. Cell Biol. 9, 219–230 (2008).
Bartel, D.P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Carthew, R.W. & Sontheimer, E.J. Origins and Mechanisms of miRNAs and siRNAs. Cell 136, 642–655 (2009).
Brennecke, J., Stark, A., Russell, R.B. & Cohen, S.M. Principles of microRNA-target recognition. PLoS Biol. 3, e85 (2005).
Doench, J.G. & Sharp, P.A. Specificity of microRNA target selection in translational repression. Genes Dev. 18, 504–511 (2004).
Grimson, A. et al. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol. Cell 27, 91–105 (2007).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Nielsen, C.B. et al. Determinants of targeting by endogenous and exogenous microRNAs and siRNAs. RNA 13, 1894–1910 (2007).
Friedman, R.C., Farh, K.K., Burge, C.B. & Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Baek, D. et al. The impact of microRNAs on protein output. Nature 455, 64–71 (2008).
Tay, Y., Zhang, J., Thomson, A.M., Lim, B. & Rigoutsos, I. MicroRNAs to Nanog, Oct4 and Sox2 coding regions modulate embryonic stem cell differentiation. Nature 455, 1124–1128 (2008).
Hafner, M. et al. Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP. Cell 141, 129–141 (2010).
Chi, S.W., Zang, J.B., Mele, A. & Darnell, R.B. Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature 460, 479–486 (2009).
Zisoulis, D.G. et al. Comprehensive discovery of endogenous Argonaute binding sites in Caenorhabditis elegans. Nat. Struct. Mol. Biol. 17, 173–179 (2010).
Babiarz, J.E., Ruby, J.G., Wang, Y., Bartel, D.P. & Blelloch, R. Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes Dev. 22, 2773–2785 (2008).
Calabrese, J.M., Seila, A.C., Yeo, G.W. & Sharp, P.A. RNA sequence analysis defines Dicer's role in mouse embryonic stem cells. Proc. Natl. Acad. Sci. USA 104, 18097–18102 (2007).
Houbaviy, H.B., Murray, M.F. & Sharp, P.A. Embryonic stem cell-specific MicroRNAs. Dev. Cell 5, 351–358 (2003).
Licatalosi, D.D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
Yeo, G.W. et al. An RNA code for the FOX2 splicing regulator revealed by mapping RNA-protein interactions in stem cells. Nat. Struct. Mol. Biol. 16, 130–137 (2009).
Leung, A.K., Calabrese, J.M. & Sharp, P.A. Quantitative analysis of Argonaute protein reveals microRNA-dependent localization to stress granules. Proc. Natl. Acad. Sci. USA 103, 18125–18130 (2006).
Tan, G.S. et al. Expanded RNA-binding activities of mammalian Argonaute 2. Nucleic Acids Res. 37, 7533–7545 (2009).
Yoda, M. et al. ATP-dependent human RISC assembly pathways. Nat. Struct. Mol. Biol. 17, 17–23 (2010).
Ciaudo, C. et al. Highly dynamic and sex-specific expression of microRNAs during early ES cell differentiation. PLoS Genet. 5, e1000620 (2009).
Bailey, T.L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Sinha, S. & Tompa, M. Discovery of novel transcription factor binding sites by statistical overrepresentation. Nucleic Acids Res. 30, 5549–5560 (2002).
Behm-Ansmant, I., Rehwinkel, J. & Izaurralde, E. MicroRNAs silence gene expression by repressing protein expression and/or by promoting mRNA decay. Cold Spring Harb. Symp. Quant. Biol. 71, 523–530 (2006).
Farh, K.K. et al. The widespread impact of mammalian MicroRNAs on mRNA repression and evolution. Science 310, 1817–1821 (2005).
Lim, L.P. et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433, 769–773 (2005).
Djuranovic, S. et al. Allosteric regulation of Argonaute proteins by miRNAs. Nat. Struct. Mol. Biol. 17, 144–150 (2010).
Pillai, R.S., Artus, C.G. & Filipowicz, W. Tethering of human Ago proteins to mRNA mimics the miRNA-mediated repression of protein synthesis. RNA 10, 1518–1525 (2004).
Wang, Y. et al. Embryonic stem cell-specific microRNAs regulate the G1-S transition and promote rapid proliferation. Nat. Genet. 40, 1478–1483 (2008).
Benetti, R. et al. A mammalian microRNA cluster controls DNA methylation and telomere recombination via Rbl2-dependent regulation of DNA methyltransferases. Nat. Struct. Mol. Biol. 15, 268–279 (2008).
Sinkkonen, L. et al. MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nat. Struct. Mol. Biol. 15, 259–267 (2008).
Foshay, K.M. & Gallicano, G.I. miR-17 family miRNAs are expressed during early mammalian development and regulate stem cell differentiation. Dev. Biol. 326, 431–443 (2009).
O'Donnell, K.A., Wentzel, E.A., Zeller, K.I., Dang, C.V. & Mendell, J.T. c-Myc-regulated microRNAs modulate E2F1 expression. Nature 435, 839–843 (2005).
Xiao, C. et al. Lymphoproliferative disease and autoimmunity in mice with increased miR-17–92 expression in lymphocytes. Nat. Immunol. 9, 405–414 (2008).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–345 (2005).
Kloosterman, W.P., Wienholds, E., Ketting, R.F. & Plasterk, R.H. Substrate requirements for let-7 function in the developing zebrafish embryo. Nucleic Acids Res. 32, 6284–6291 (2004).
Gu, S., Jin, L., Zhang, F., Sarnow, P. & Kay, M.A. Biological basis for restriction of microRNA targets to the 3′ untranslated region in mammalian mRNAs. Nat. Struct. Mol. Biol. 16, 144–150 (2009).
Melton, C., Judson, R.L. & Blelloch, R. Opposing microRNA families regulate self-renewal in mouse embryonic stem cells. Nature 463, 621–626 (2010).
Landgraf, P. et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129, 1401–1414 (2007).
Judson, R.L., Babiarz, J.E., Venere, M. & Blelloch, R. Embryonic stem cell-specific microRNAs promote induced pluripotency. Nat. Biotechnol. 27, 459–461 (2009).
Choi, W.Y., Giraldez, A.J. & Schier, A.F. Target protectors reveal dampening and balancing of Nodal agonist and antagonist by miR-430. Science 318, 271–274 (2007).
Rosa, A., Spagnoli, F.M. & Brivanlou, A.H. The miR-430/427/302 family controls mesendodermal fate specification via species-specific target selection. Dev. Cell 16, 517–527 (2009).
Li, X., Cassidy, J.J., Reinke, C.A., Fischboeck, S. & Carthew, R.W. A microRNA imparts robustness against environmental fluctuation during development. Cell 137, 273–282 (2009).
Marson, A. et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134, 521–533 (2008).
Tsang, J., Zhu, J. & van Oudenaarden, A. MicroRNA-mediated feedback and feedforward loops are recurrent network motifs in mammals. Mol. Cell 26, 753–767 (2007).
Caudy, A.A., Myers, M., Hannon, G.J. & Hammond, S.M. Fragile X-related protein and VIG associate with the RNA interference machinery. Genes Dev. 16, 2491–2496 (2002).
Edbauer, D. et al. Regulation of synaptic structure and function by FMRP-associated microRNAs miR-125b and miR-132. Neuron 65, 373–384 (2010).
Höck, J. et al. Proteomic and functional analysis of Argonaute-containing mRNA-protein complexes in human cells. EMBO Rep. 8, 1052–1060 (2007).
Kertesz, M., Iovino, N., Unnerstall, U., Gaul, U. & Segal, E. The role of site accessibility in microRNA target recognition. Nat. Genet. 39, 1278–1284 (2007).
Blanchette, M. et al. Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res. 14, 708–715 (2004).
Ule, J., Jensen, K., Mele, A. & Darnell, R.B. CLIP: a method for identifying protein-RNA interaction sites in living cells. Methods 37, 376–386 (2005).
Hafner, M. et al. Identification of microRNAs and other small regulatory RNAs using cDNA library sequencing. Methods 44, 3–12 (2008).
Calabrese, J.M. & Sharp, P.A. Characterization of the short RNAs bound by the P19 suppressor of RNA silencing in mouse embryonic stem cells. RNA 12, 2092–2102 (2006).
Bailey, T.L. & Gribskov, M. Combining evidence using p-values: application to sequence homology searches. Bioinformatics 14, 48–54 (1998).
Acknowledgements
We thank C. Burge, J. Wilusz, A. Ravi and members of the Sharp laboratory for critical comments, M. Lindstrom for illustration, T. Cybulski for technical help, A. Leshinsky and R. Cook for running the Solexa sequencing samples in the KI Biopolymer & Proteomics Core Facility, M. Luo and L. Smeester for microarray technical assistance in the MIT Department of Biology Biomicrocenter and C. Whittaker for bioinformatics support in the Bioinformatics & Computing Core Facility at the Koch Institute. A.K.L.L. was supported by a special fellowship from the Leukemia and Lymphoma Society. A.G.Y. was partially supported by a David H. Koch graduate fellowship. This work was supported by United States Public Health Service grants R01-GM34277 and R01-CA133404 from the US National Institutes of Health, P01-CA42063 from the National Cancer Institute to P.A.S. and partially by Cancer Center Support (core) grant P30-CA14051 from the National Cancer Institute.
Author information
Authors and Affiliations
Contributions
A.K.L.L. and A.G.Y. designed and performed the experiments; A.K.L.L., A.G.Y. and P.A.S. wrote the paper; A.D.B. performed experiments; A.B., C.B.N. and G.X.Z. performed the bioinformatics analyses. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–7, Supplementary Notes and Supplementary Methods (PDF 9221 kb)
Rights and permissions
About this article
Cite this article
Leung, A., Young, A., Bhutkar, A. et al. Genome-wide identification of Ago2 binding sites from mouse embryonic stem cells with and without mature microRNAs. Nat Struct Mol Biol 18, 237–244 (2011). https://doi.org/10.1038/nsmb.1991
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1991
This article is cited by
-
Focal adhesion ribonucleoprotein complex proteins are major humoral cancer antigens and targets in autoimmune diseases
Communications Biology (2020)
-
Nuclear gene proximity and protein interactions shape transcript covariations in mammalian single cells
Nature Communications (2020)
-
Arsenic is more potent than cadmium or manganese in disrupting the INS-1 beta cell microRNA landscape
Archives of Toxicology (2019)
-
Inhibition of the JNK/MAPK signaling pathway by myogenesis-associated miRNAs is required for skeletal muscle development
Cell Death & Differentiation (2018)
-
Evaluation and control of miRNA-like off-target repression for RNA interference
Cellular and Molecular Life Sciences (2018)