Abstract
The BBSome is a coat-like ciliary trafficking complex composed of proteins mutated in Bardet-Biedl syndrome (BBS). A critical step in BBSome-mediated sorting is recruitment of the BBSome to membranes by the GTP-bound Arf-like GTPase ARL6. We have determined crystal structures of Chlamydomonas reinhardtii ARL6–GDP, ARL6–GTP and the ARL6–GTP–BBS1 complex. The structures demonstrate how ARL6–GTP binds the BBS1 β-propeller at blades 1 and 7 and explain why GTP- but not GDP-bound ARL6 can recruit the BBSome to membranes. Single point mutations in the ARL6-GTP-BBS1 interface abolish the interaction of ARL6 with the BBSome and prevent the import of BBSomes into cilia. Furthermore, we show that BBS1 with the M390R mutation, responsible for 30% of all reported BBS disease cases, fails to interact with ARL6–GTP, thus providing a molecular rationale for patient pathologies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Veland, I.R., Awan, A., Pedersen, L.B., Yoder, B.K. & Christensen, S.O.R.T. Primary cilia and signaling pathways in mammalian development, health and disease. Nephron. Physiol. 111, 39–53 (2009).
Sung, C.-H. & Leroux, M.R. The roles of evolutionarily conserved functional modules in cilia-related trafficking. Nat. Cell Biol. 15, 1387–1397 (2013).
Scholey, J.M. Intraflagellar transport. Annu. Rev. Cell Dev. Biol. 19, 423–443 (2003).
Rosenbaum, J.L. & Witman, G.B. Intraflagellar transport. Nat. Rev. Mol. Cell Biol. 3, 813–825 (2002).
Ishikawa, H. & Marshall, W.F. Ciliogenesis: building the cell's antenna. Nat. Rev. Mol. Cell Biol. 12, 222–234 (2011).
Bhogaraju, S., Engel, B.D. & Lorentzen, E. Intraflagellar transport complex structure and cargo interactions. Cilia 2, 10 (2013).
Taschner, M., Bhogaraju, S. & Lorentzen, E. Architecture and function of IFT complex proteins in ciliogenesis. Differentiation 83, S12–S22 (2012).
Hou, Y. et al. Functional analysis of an individual IFT protein: IFT46 is required for transport of outer dynein arms into flagella. J. Cell Biol. 176, 653–665 (2007).
Ishikawa, H. et al. TTC26/DYF13 is an intraflagellar transport protein required for transport of motility-related proteins into flagella. Elife 3, e01566 (2014).
Bhogaraju, S. et al. Molecular basis of tubulin transport within the cilium by IFT74 and IFT81. Science 341, 1009–1012 (2013).
Domire, J.S. et al. Dopamine receptor 1 localizes to neuronal cilia in a dynamic process that requires the Bardet-Biedl syndrome proteins. Cell. Mol. Life Sci. 68, 2951–2960 (2011).
Seo, S. et al. A novel protein LZTFL1 regulates ciliary trafficking of the BBSome and Smoothened. PLoS Genet. 7, e1002358 (2011).
Nachury, M.V. et al. A core complex of BBS proteins cooperates with the GTPase Rab8 to promote ciliary membrane biogenesis. Cell 129, 1201–1213 (2007).
Berbari, N.F., Lewis, J.S., Bishop, G.A., Askwith, C.C. & Mykytyn, K. Bardet-Biedl syndrome proteins are required for the localization of G protein-coupled receptors to primary cilia. Proc. Natl. Acad. Sci. USA 105, 4242–4246 (2008).
Zhang, Q. et al. Bardet-Biedl syndrome 3 (Bbs3) knockout mouse model reveals common BBS-associated phenotypes and Bbs3 unique phenotypes. Proc. Natl. Acad. Sci. USA 108, 20678–20683 (2011).
Loktev, A.V. et al. A BBSome subunit links ciliogenesis, microtubule stability, and acetylation. Dev. Cell 15, 854–865 (2008).
Katsanis, N., Lupski, J.R. & Beales, P.L. Exploring the molecular basis of Bardet-Biedl syndrome. Hum. Mol. Genet. 10, 2293–2299 (2001).
Tobin, J.L. & Beales, P.L. Bardet-Biedl syndrome: beyond the cilium. Pediatr. Nephrol. 22, 926–936 (2007).
Sheffield, V.C. The blind leading the obese: the molecular pathophysiology of a human obesity syndrome. Trans. Am. Clin. Climatol. Assoc. 121, 172–181 discussion 181–182 (2010).
Blacque, O.E. & Leroux, M.R. Bardet-Biedl syndrome: an emerging pathomechanism of intracellular transport. Cell. Mol. Life Sci. 63, 2145–2161 (2006).
Lechtreck, K.-F. The Chlamydomonas reinhardtii BBSome is an IFT cargo required for export of specific signaling proteins from flagella. J. Cell Biol. 187, 1117–1132 (2009).
Rosenbaum, J.L. & Witman, G.B. Intraflagellar transport. Nat. Rev. Mol. Cell Biol. 3, 813–825 (2002).
Ou, G., Blacque, O.E., Snow, J.J., Leroux, M.R. & Scholey, J.M. Functional coordination of intraflagellar transport motors. Nature 436, 583–587 (2005).
Lechtreck, K.-F. et al. Cycling of the signaling protein phospholipase D through cilia requires the BBSome only for the export phase. J. Cell Biol. 201, 249–261 (2013).
Zhang, Y. et al. BBS mutations modify phenotypic expression of CEP290-related ciliopathies. Hum. Mol. Genet. 23, 40–51 (2014).
Bujakowska, K.M. et al. Mutations in IFT172 cause isolated retinal degeneration and Bardet-Biedl syndrome. Hum. Mol. Genet. 10.1093/hmg/ddu441 (28 August 2017) (2014).
Aldahmesh, M.A. et al. IFT27, encoding a small GTPase component of IFT particles, is mutated in a consanguineous family with Bardet-Biedl syndrome. Hum. Mol. Genet. 23, 3307–3315 (2014).
Keady, B.T. et al. IFT25 links the signal-dependent movement of Hedgehog components to intraflagellar transport. Dev. Cell 22, 940–951 (2012).
Liew, G.M. et al. The intraflagellar transport protein IFT27 promotes BBSome exit from cilia through the GTPase ARL6/BBS3. Dev. Cell 10.1016/j.devcel.2014.09.004 (2014).
Eguether, T. et al. IFT27 links the BBSome to IFT for maintenance of ciliary signaling compartment. Dev. Cell 10.1016/j.devcel.2014.09.011 (2014).
Jin, H. et al. The conserved Bardet-Biedl syndrome proteins assemble a coat that traffics membrane proteins to cilia. Cell 141, 1208–1219 (2010).
Wiens, C.J. et al. Bardet-Biedl syndrome-associated small GTPase ARL6 (BBS3) functions at or near the ciliary gate and modulates Wnt signaling. J. Biol. Chem. 285, 16218–16230 (2010).
Hemsworth, G.R., Price, H.P., Smith, D.F. & Wilson, K.S. Crystal structure of the small GTPase Arl6/BBS3 from Trypanosoma brucei. Protein Sci. 22, 196–203 (2013).
Amor, J.C., Harrison, D.H., Kahn, R.A. & Ringe, D. Structure of the human ADP-ribosylation factor 1 complexed with GDP. Nature 372, 704–708 (1994).
Liu, Y., Kahn, R.A. & Prestegard, J.H. Dynamic structure of membrane-anchored ArfGTP. Nat. Struct. Mol. Biol. 17, 876–881 (2010).
Bi, X., Corpina, R.A. & Goldberg, J. Structure of the Sec23/24–Sar1 pre-budding complex of the COPII vesicle coat. Nature 419, 271–277 (2002).
Chiang, A.P. et al. Comparative genomic analysis identifies an ADP-ribosylation factor-like gene as the cause of Bardet-Biedl syndrome (BBS3). Am. J. Hum. Genet. 75, 475–484 (2004).
Fan, Y. et al. Mutations in a member of the Ras superfamily of small GTP-binding proteins causes Bardet-Biedl syndrome. Nat. Genet. 36, 989–993 (2004).
Mykytyn, K. et al. Identification of the gene (BBS1) most commonly involved in Bardet-Biedl syndrome, a complex human obesity syndrome. Nat. Genet. 31, 435–438 (2002).
Beales, P.L. et al. Genetic interaction of BBS1 mutations with alleles at other BBS loci can result in non-Mendelian Bardet-Biedl syndrome. Am. J. Hum. Genet. 72, 1187–1199 (2003).
Jékely, G. & Arendt, D. Evolution of intraflagellar transport from coated vesicles and autogenous origin of the eukaryotic cilium. BioEssays 28, 191–198 (2006).
van Dam, T.J.P. et al. Evolution of modular intraflagellar transport from a coatomer-like progenitor. Proc. Natl. Acad. Sci. USA 110, 6943–6948 (2013).
Shiba, T. et al. Molecular mechanism of membrane recruitment of GGA by ARF in lysosomal protein transport. Nat. Struct. Biol. 10, 386–393 (2003).
Yu, X., Breitman, M. & Goldberg, J. A structure-based mechanism for Arf1-dependent recruitment of coatomer to membranes. Cell 148, 530–542 (2012).
Ren, X., Farías, G.G., Canagarajah, B.J., Bonifacino, J.S. & Hurley, J.H. Structural basis for recruitment and activation of the AP-1 clathrin adaptor complex by Arf1. Cell 152, 755–767 (2013).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Storoni, L.C., McCoy, A.J. & Read, R.J. Likelihood-enhanced fast rotation functions. Acta Crystallogr. D Biol. Crystallogr. 60, 432–438 (2004).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Fiser, A. & Sali, A. Modeller: generation and refinement of homology-based protein structure models. Methods Enzymol. 374, 461–491 (2003).
Shaner, N.C. et al. A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum. Nat. Methods 10, 407–409 (2013).
Wolff, A. et al. Distribution of glutamylated alpha and beta-tubulin in mouse tissues using a specific monoclonal antibody, GT335. Eur. J. Cell Biol. 59, 425–432 (1992).
Acknowledgements
We thank the staff at Swiss Light Source for guidance with X-ray diffraction data collection, the biochemistry core facility and the crystallization facility of the Max Planck Institute of Biochemistry (MPI-B, Munich) for access to crystallization screening and the Bavarian NMR Center for NMR measurement time. We also thank I.B. Schaefer (MPI-B) for DNA encoding BBS subunits, S. Wachter (MPI-B) for assistance with GTPase assays and M. Taschner (MPI-B) for expert advice on protein production from insect cells and for carefully reading the manuscript. This work was funded by an Emmy Noether grant (Deutsche Forschungsgemeinschaft; LO1627/1-1), by the European Research Council (grant 310343) and by the European Molecular Biology Organization Young Investigator program. A.R.N. was supported by the Fayez Sarofim Fellowship of the Damon Runyon Cancer Research Foundation (DRG 2160-13). This work was supported by a grant to M.V.N. from the NIH/National Institute of General Medical Sciences (R01GM089933).
Author information
Authors and Affiliations
Contributions
A.M. carried out the protein biochemistry and structural biology under the supervision of E.L.; A.R.N. carried out the pulldown experiments of native BBSome with wild-type and mutant ARL6 and the cell biology experiments under the supervision of M.V.N.; A.M. and E.L. designed the experiments and wrote the paper with input from A.R.N. and M.V.N.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Purification and Kd determination for ARL6ΔN–BBS1N complexes.
Binding experiments of HsARL6Q73LΔN to (a) HsBBS1N, (b) HsBBS1NE234K, (c) HsBBS1NM390R and (d) CrBBS1N. Left panel shows size exclusion chromatography profiles, the central panel the corresponding SDS gel and the right panel the ITC titration experiments. Stable ARL6ΔN-BBS1N complexes are observed in all cases except for the HsBBS1NM390R protein. The HsBBS1NM390R variant was soluble and eluted as a broad peak with a maximal A280 absorption at ~9.5-10.0ml (void volume where aggregated proteins elute is 7.5ml on the S200 column). Reported Kd values are the average of 3 independent experiments.
Supplementary Figure 2 Comparison of ARL6 structures and electron density for CrARL6–GTP–CrBBS1N and GDP or GTP nucleotides.
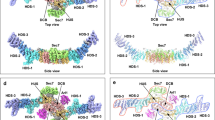
(a) Structural comparison of ARL6ΔN–GTP from different organisms, Chlamydomonas reinhardtii (green), Homo sapiens (cyan) and Trypanosoma brucei (pink) shows structural similarities between the three proteins. (b) The structures of CrARL6ΔN–GTP (orange) and CrARL6ΔN–GTP (green) from the BBS1N complex superimpose with an RMSD of 0.4Å. (c) Structures of CrARL6ΔN–GDP and Arf1–GDP (PDB code: 1HUR) superposed show a similar conformation of the interswitch region. The N-terminal amphipathic helix rests on the G-domain of Arf1. (d) Structures of CrARL6ΔN–GTP and Arf1–GTP (PDB code: 2KSQ 32) superposed show a similar conformation of the interswitch region. The N-terminal amphipathic helix is expelled from the G-domain of Arf1 and available for membrane association. (e) Structure of the CrARL6ΔN–GTP–CrBBS1N complex in transparent surface representation with the polypeptides also displayed as cartoon and the GTP molecule as sticks. CrARL6ΔN–GTP and BBS1N have been pull apart by 10Å in silico to allow visualization of the small but complementary interaction interface of 600Å2 (calculated using PISA server; Krissinel, E. & Henrick, K, J. Mol. Biol. 372, 774–797, 2007).). (f) Experimental electron density map at 1σ obtained by the MR-SAD protocol in Phaser using four molecules of CrARL6ΔN–GTP found by MR and 35 Hg sites. The final model for CrARL6 (yellow) and CrBBS1 (magenta) are displayed as Cα traces. Difference maps (Fo-Fc, 3σ) calculated using model phase before the addition of nucleotides to the model is shown in green for (g) GTP in CrARL6ΔN–GTP–CrBBS1N (h) GTP in CrARL6ΔN–GTP and (i) GDP in CrARL6ΔN–GDP. All structure figures were made using PYMOL (www.pymol.org) and electron density maps were obtained from Coot 51.
Supplementary Figure 3 Structure-based multiple sequence alignment.
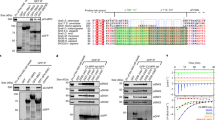
Multiple sequence alignment of (a) BBS1 and (b) ARL6 proteins from Chlamydomonas reinhardtii, Homo sapiens, Danio rerio, Xenopus tropicalis and Caenorhabditis elegans using the Clustal omega server (http://www.clustal.org/omega/). Secondary structure elements derived from the CrARL6ΔN–GTP–CrBBS1N complex structure are shown above the sequences. Conservative substitutions are shown in red font and and identical residues in white font on a red background. Interacting residues between ARL6 and BBS1 are shown with a blue asterisk (hydrophobic) or a red asterisk (hydrophilic). The alignments were generated using the ESPript server (http://espript.ibcp.fr/).
Supplementary Figure 4 SEC and GTP binding by BBS1 and ARL6 point mutants.
(a) (left) SEC elution profile of structure-based HsBBS1N point-mutations and (right) the corresponding SDS gel. (b) (left) SEC elution profile of structure-based HsARL6ΔN point-mutations and (right) the corresponding SDS gel. All HsBBS1N and HsARL6ΔN mutants elute as symmetric peaks at the same elution volume as the WT proteins and are thus folded. Superpositioning of 1D-1H-NMR spectra of (c) WT, (d) R77A, (e) L100E and (f) R108A HsARL6ΔN without nucleotide (blue) and titrated with a 1.2 excess of GTP (red). The regions of the spectra that corresponds to amide and the methyl groups are shown in the green and purple boxes, respectively. The spectra demonstrate that the mutants are folded and chemical shift perturbations (highlighted by arrows) indicate that ARL6 mutants bind GTP.
Supplementary Figure 5 GTPase assay for CrARL6ΔN and CrARL6ΔN–CrBBS1N.
(a)The GTPase activity was measured in a phosphate release assay using the EnzCheck Phosphate kit (Invitrogen). 1mM GTP was incubated with buffer (neg. control) or with 50μM of CrARL6ΔN or CrARL6ΔN-CrBBS1N and the release of inorganic phosphate was followed over the course of 20min. The experiment shows that CrBBS1N does not enhance the GTPase activity of CrArL6ΔN. In the positive control, the conversion of 100μM inorganic phosphate was followed. Curves for CrARL6ΔN and CrARL6ΔN-CrBBS1N are the average of 3 independent experiments.
Supplementary Figure 6 BBS1 disease-mutation pulldowns with GST-HsARL6.
WB using anti-His antibody for a GST pull-down experiments of GST-HsARL6 Q73L with His-tagged HsBBS1N, HsBBS1E234K or HsBBS1M390R shows that ARL6 interacts with wild-type and the E234K mutant, but not with the M390R BBS1N mutant protein (even though the His-HsBBS1NM390R was added in large excess).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 1867 kb)
Supplementary Data Set 1
Full-size gels from main figures (PDF 121 kb)
Rights and permissions
About this article
Cite this article
Mourão, A., Nager, A., Nachury, M. et al. Structural basis for membrane targeting of the BBSome by ARL6. Nat Struct Mol Biol 21, 1035–1041 (2014). https://doi.org/10.1038/nsmb.2920
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2920
This article is cited by
-
Appearing and disappearing acts of cilia
Journal of Biosciences (2023)
-
Bardet–Biedl syndrome and related disorders in Japan
Journal of Human Genetics (2020)
-
Protein interaction perturbation profiling at amino-acid resolution
Nature Methods (2017)
-
Structure of Rab11–FIP3–Rabin8 reveals simultaneous binding of FIP3 and Rabin8 effectors to Rab11
Nature Structural & Molecular Biology (2015)