Abstract
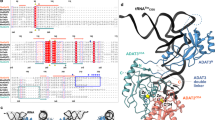
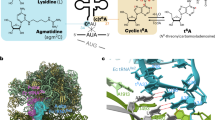
Bacterial tRNA adenosine deaminases (TadAs) catalyze the hydrolytic deamination of adenosine to inosine at the wobble position of tRNAArg2, a process that enables this single tRNA to recognize three different arginine codons in mRNA. In addition, inosine is also introduced at the wobble position of multiple eukaryotic tRNAs. The genes encoding these deaminases are essential in bacteria and yeast, demonstrating the importance of their biological activity. Here we report the crystallization and structure determination to 2.0 Å of Staphylococcus aureus TadA bound to the anticodon stem-loop of tRNAArg2 bearing nebularine, a non-hydrolyzable adenosine analog, at the wobble position. The cocrystal structure reveals the basis for both sequence and structure specificity in the interactions of TadA with RNA, and it additionally provides insight into the active site architecture that promotes efficient hydrolytic deamination.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Carter, C.W. Jr. The nucleoside deaminases for cytidine and adenosine: structure, transition state stabilization, mechanism, and evolution. Biochimie 77, 92–98 (1995).
Maas, S., Rich, A. & Nishikura, K. A-to-I RNA editing: recent news and residual mysteries. J. Biol. Chem. 278, 1391–1394 (2003).
Pham, P., Bransteitter, R. & Goodman, M.F. Reward versus risk: DNA cytidine deaminases triggering immunity and disease. Biochemistry 44, 2703–2715 (2005).
Wedekind, J.E., Dance, G.S., Sowden, M.P. & Smith, H.C. Messenger RNA editing in mammals: new members of the APOBEC family seeking roles in the family business. Trends Genet. 19, 207–216 (2003).
Johansson, E., Mejlhede, N., Neuhard, J. & Larsen, S. Crystal structure of the tetrameric cytidine deaminase from Bacillus subtilis at 2.0 Å resolution. Biochemistry 41, 2563–2570 (2002).
Wilson, D.K., Rudolph, F.B. & Quiocho, F.A. Atomic structure of adenosine deaminase complexed with a transition-state analog: understanding catalysis and immunodeficiency mutations. Science 252, 1278–1284 (1991).
Bass, B.L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71, 817–846 (2002).
Petersen-Mahrt, S.K., Harris, R.S. & Neuberger, M.S. AID mutates E. coli suggesting a DNA deamination mechanism for antibody diversification. Nature 418, 99–103 (2002).
Sheehy, A.M., Gaddis, N.C., Choi, J.D. & Malim, M.H. Isolation of a human gene that inhibits HIV-1 infection and is suppressed by the viral Vif protein. Nature 418, 646–650 (2002).
Teng, B., Burant, C.F. & Davidson, N.O. Molecular cloning of an apolipoprotein B messenger RNA editing protein. Science 260, 1816–1819 (1993).
Wolf, J., Gerber, A.P. & Keller, W. TadA, an essential tRNA-specific adenosine deaminase from Escherichia coli. EMBO J. 21, 3841–3851 (2002).
Sprinzl, M., Horn, C., Brown, M., Ioudovitch, A. & Steinberg, S. Compilation of tRNA sequences and sequences of tRNA genes. Nucleic Acids Res. 26, 148–153 (1998).
Gerber, A.P. & Keller, W. An adenosine deaminase that generates inosine at the wobble position of tRNAs. Science 286, 1146–1149 (1999).
Gerber, A., Grosjean, H., Melcher, T. & Keller, W. Tad1p, a yeast tRNA-specific adenosine deaminase, is related to the mammalian pre-mRNA editing enzymes ADAR1 and ADAR2. EMBO J. 17, 4780–4789 (1998).
Grosjean, H. et al. Enzymatic conversion of adenosine to inosine and to N1-methylinosine in transfer RNAs: a review. Biochimie 78, 488–501 (1996).
Betts, L., Xiang, S., Short, S.A., Wolfenden, R. & Carter, C.W. Jr. Cytidine deaminase. The 2.3 Å crystal structure of an enzyme: transition-state analog complex. J. Mol. Biol. 235, 635–656 (1994).
Chung, S.J., Fromme, J.C. & Verdine, G.L. Structure of human cytidine deaminase bound to a potent inhibitor. J. Med. Chem. 48, 658–660 (2005).
Xie, K. et al. The structure of a yeast RNA-editing deaminase provides insight into the fold and function of activation-induced deaminase and APOBEC-1. Proc. Natl. Acad. Sci. USA 101, 8114–8119 (2004).
Kinoshita, T., Nishio, N., Nakanishi, I., Sato, A. & Fujii, T. Structure of bovine adenosine deaminase complexed with 6-hydroxy-1,6-dihydropurine riboside. Acta Crystallogr. D Biol. Crystallogr. 59, 299–303 (2003).
Elias, Y. & Huang, R.H. Biochemical and structural studies of A-to-I editing by tRNA:A34 deaminases at the wobble position of transfer RNA. Biochemistry 44, 12057–65 (2005).
Kuratani, M. et al. Crystal structure of tRNA adenosine deaminase (TadA) from Aquifex aeolicus. J. Biol. Chem. 280, 16002–16008 (2005).
Shi, H. & Moore, P.B. The crystal structure of yeast phenylalanine tRNA at 1.93 Å resolution: a classic structure revisited. RNA 6, 1091–1105 (2000).
Nissen, P., Thirup, S., Kjeldgaard, M. & Nyborg, J. The crystal structure of Cys-tRNACys-EF-Tu-GDPNP reveals general and specific features in the ternary complex and in tRNA. Struct. Fold. Des. 7, 143–156 (1999).
Murphy, F.V.t. & Ramakrishnan, V. Structure of a purine-purine wobble base pair in the decoding center of the ribosome. Nat. Struct. Mol. Biol. 11, 1251–1252 (2004).
Ogle, J.M., Murphy, F.V., Tarry, M.J. & Ramakrishnan, V. Selection of tRNA by the ribosome requires a transition from an open to a closed form. Cell 111, 721–732 (2002).
Nagaswamy, U., Voss, N., Zhang, Z. & Fox, G.E. Database of non-canonical base pairs found in known RNA structures. Nucleic Acids Res. 28, 375–376 (2000).
Ashraf, S.S. et al. The uridine in “U-turn”: contributions to tRNA-ribosomal binding. RNA 5, 503–511 (1999).
Xie, W., Liu, X. & Huang, R.H. Chemical trapping and crystal structure of a catalytic tRNA guanine transglycosylase covalent intermediate. Nat. Struct. Biol. 10, 781–788 (2003).
Rould, M.A., Perona, J.J. & Steitz, T.A. Structural basis of anticodon loop recognition by glutaminyl-tRNA synthetase. Nature 352, 213–218 (1991).
Wolfenden, R. & Kati, W.M. Testing the limits of protein-ligand binding discrimination with transition-state analogue inhibitors. Acc. Chem. Res. 24, 209–215 (1991).
Veliz, E.A., Easterwood, L.M. & Beal, P.A. Substrate analogues for an RNA-editing adenosine deaminase: mechanistic investigation and inhibitor design. J. Am. Chem. Soc. 125, 10867–10876 (2003).
Van Duyne, G.D., Standaert, R.F., Karplus, P.A., Schreiber, S.L. & Clardy, J. Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J. Mol. Biol. 229, 105–124 (1993).
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–563 (2000).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Terwilliger, T.C. SOLVE and RESOLVE: automated structure solution and density modification. Methods Enzymol. 374, 22–37 (2003).
COLLABORATIVE COMPUTATIONAL PROJECT. N. The CCP4 Suite: Programs for Protein Crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Cowtan, K. dm: An automated procedure for phase improvement by density modification. Joint CCP4 and ESF-EACBM Newsletter on Protein Crystallography 31, 34–38 (1994).
Morris, R.J. et al. Breaking good resolutions with ARP/wARP. J. Synchrotron Radiat. 11, 56–59 (2004).
Perrakis, A., Morris, R. & Lamzin, V.S. Automated protein model building combined with iterative structure refinement. Nat. Struct. Biol. 6, 458–463 (1999).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Brunger, A.T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Laskowski, R.J., Macarthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–290 (1993).
Kabsch, W. A solution for the best way to relate two sets of vectors. Acta Crystallogr. A 32, 922–923 (1976).
Acknowledgements
We are grateful to the entire beamline staff at NSLS X25A, especially M. Becker, for assistance in data collection and processing, and to members of the Verdine research group for valuable help and discussions and for critical reading of the manuscript. H.C.L. was supported by a postdoctoral fellowship from the Irvington Institute for Immunological Research, A.J.R. was supported by a National Science Foundation graduate fellowship and the work was also supported by a grant from the US National Institutes of Health to G.L.V.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Protein sequence alignment of TadA with related deaminases. (PDF 989 kb)
Supplementary Fig. 2
Conformational changes in TadA. (PDF 512 kb)
Supplementary Fig. 3
Comparison of base pairing observed between C32-A38 in the TadA–RNA structure, the tRNAPhe structure, and a normal C-A+ wobble pair. (PDF 104 kb)
Supplementary Fig. 4
Superposition of three anticodon stem-loops. (PDF 607 kb)
Supplementary Fig. 5
Activity of TadA on minimal RNA stem-loop substrates. (PDF 490 kb)
Rights and permissions
About this article
Cite this article
Losey, H., Ruthenburg, A. & Verdine, G. Crystal structure of Staphylococcus aureus tRNA adenosine deaminase TadA in complex with RNA. Nat Struct Mol Biol 13, 153–159 (2006). https://doi.org/10.1038/nsmb1047
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1047
This article is cited by
-
Functions and consequences of AID/APOBEC-mediated DNA and RNA deamination
Nature Reviews Genetics (2022)
-
Crystal structure of the yeast heterodimeric ADAT2/3 deaminase
BMC Biology (2020)
-
Substrate sequence selectivity of APOBEC3A implicates intra-DNA interactions
Scientific Reports (2018)
-
Crystal structure of the catalytic domain of HIV-1 restriction factor APOBEC3G in complex with ssDNA
Nature Communications (2018)
-
Crystal structure of APOBEC3A bound to single-stranded DNA reveals structural basis for cytidine deamination and specificity
Nature Communications (2017)