Abstract
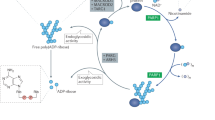
Nicotinamide phosphoribosyltransferase (NMPRTase) has a crucial role in the salvage pathway of NAD+ biosynthesis, and a potent inhibitor of NMPRTase, FK866, can reduce cellular NAD+ levels and induce apoptosis in tumors. We have determined the crystal structures at up to 2.1-Å resolution of human and murine NMPRTase, alone and in complex with the reaction product nicotinamide mononucleotide or the inhibitor FK866. The structures suggest that Asp219 is a determinant of substrate specificity of NMPRTase, which is confirmed by our mutagenesis studies. FK866 is bound in a tunnel at the interface of the NMPRTase dimer, and mutations in this binding site can abolish the inhibition by FK866. Contrary to current knowledge, the structures show that FK866 should compete directly with the nicotinamide substrate. Our structural and biochemical studies provide a starting point for the development of new anticancer agents.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ziegler, M. New functions of a long-known molecule. Emerging roles of NAD in cellular signaling. Eur. J. Biochem. 267, 1550–1564 (2000).
Guarente, L. & Picard, F. Calorie restriction-the SIR2 connection. Cell 120, 473–482 (2005).
Marmorstein, R. Structure and chemistry of the Sir2 family of NAD+-dependent histone/protein deacetylases. Biochem. Soc. Trans. 32, 904–909 (2004).
Magni, G. et al. Enzymology of NAD+ homeostasis in man. Cell. Mol. Life Sci. 61, 19–34 (2004).
Araki, T., Sasaki, Y. & Milbrandt, J. Increased nuclear NAD biosynthesis and SIRT1 activation prevent axonal degeneration. Science 305, 1010–1013 (2004).
Guse, A.H. Second messenger function and the structure-activity relationship of cyclic adenosine diphosphoribose (cADPR). FEBS J. 272, 4590–4597 (2005).
Rongvaux, A. et al. Pre-B-cell colony-enhancing factor, whose expression is up-regulated in activated lymphocytes, is a nicotinamide phorphoribosyltransferase, a cytosolic enzyme involved in NAD biosynthesis. Eur. J. Immunol. 32, 3225–3234 (2002).
Wosikowski, K. et al. WK175, a novel antitumor agent, decreases the intracellular nicotinamide adenine dinucleotide concentration and induces the apoptotic cascade in human leukemia cells. Cancer Res. 62, 1057–1062 (2002).
Hasmann, M. & Schemainda, I. FK866, a highly specific noncompetitive inhibitor of nicotinamide phosphoribosyltransferase, represents a novel mechanism for induction of tumor cell apoptosis. Cancer Res. 63, 7436–7442 (2003).
Hufton, S.E. et al. A profile of differentially expressed genes in primary colorectal cancer using suppression substractive hybridization. FEBS Lett. 463, 77–82 (1999).
van Beijnum, J.R. et al. Target validation for genomics using peptide-specific phage antibodies: a study of five gene products overexpressed in colorectal cancer. Int. J. Cancer 101, 118–127 (2002).
Muruganandham, M. et al. Metabolic signatures associated with a NAD synthesis inhibitor-induced tumor apoptosis identified by 1H-decoupled-31P magnetic resonance spectroscopy. Clin. Cancer Res. 11, 3503–3513 (2005).
Drevs, J., Loser, R., Rattel, B. & Esser, N. Antiangiogenic potency of FK866/K22.175, a new inhibitor of intracellular NAD biosynthesis, in murine renal cell carcinoma. Anticancer Res. 23, 4853–4858 (2003).
Samal, B. et al. Cloning and characterization of the cDNA encoding a novel human pre-B-cell colony-enhancing factor. Mol. Cell. Biol. 14, 1431–1437 (1994).
Fukuhara, A. et al. Visfatin, a protein secreted by visceral fat that mimics the effects of insulin. Science 307, 426–430 (2005).
Hug, C. & Lodish, H.F. Visfatin: a new adipokine. Science 307, 366–367 (2005).
Sethi, J.K. & Vidal-Puig, A. Visfatin: the missing link between intra-abdominal obesity and diabetes? Trends Mol. Med. 11, 344–347 (2005).
Stephens, J.M. & Vidal-Puig, A.J. An update on visfatin/pre-B cell colony-enhancing factor, an ubiquitously expressed, illusive cytokine that is regulated in obesity. Curr. Opin. Lipidol. 17, 128–131 (2006).
Kitani, T., Okuno, S. & Fujisawa, H. Growth phase-dependent changes in the subcellular localization of pre-B-cell colony-enhancing factor. FEBS Lett. 544, 74–78 (2003).
Eads, J.C., Ozturk, D., Wexler, T.B., Grubmeyer, C. & Sacchettini, J.C. A new function for a common fold: the crystal structure of quinolinic acid phosphoribosyltransferase. Structure 5, 47–58 (1997).
Sharma, V., Grubmeyer, C. & Sacchettini, J.C. Crystal structure of quinolinic acid phosphoribosyltransferase from Mycobacterium tuberculosis: a potential TB drug target. Structure 6, 1587–1599 (1998).
Shin, D.H. et al. Crystal structure of a nicotinate phosphoribosyltransferase from Thermoplasma acidophilum. J. Biol. Chem. 280, 18326–18335 (2005).
Chappie, J.S. et al. The structure of a eukaryotic nicotinic acid phosphoribosyltransferase reveals structural heterogeneity among type II PRTases. Structure 13, 1385–1396 (2005).
Hendrickson, W.A. Determination of macromolecular structures from anomalous diffraction of synchrotron radiation. Science 254, 51–58 (1991).
Revollo, J.R., Grimm, A.A. & Imai, S.-I. The NAD biosynthesis pathway mediated by nicotinamide phosphoribosyltransferase regulates Sir2 activity in mammalian cells. J. Biol. Chem. 279, 50754–50763 (2004).
Gross, J., Rajavel, M., Segura, E. & Grubmeyer, C. Energy coupling in Salmonella typhimurium nicotinic acid phosphoribosyltransferase: identification of His-219 as site of phosphorylation. Biochemistry 35, 3917–3924 (1996).
Gross, J.W., Rajavel, M. & Grubmeyer, C. Kinetic mechanism of nicotinic acid phosphoribosyltransferase: implications for energy coupling. Biochemistry 37, 4189–4199 (1998).
Hendrickson, W.A., Horton, J.R. & LeMaster, D.M. Selenomethionyl proteins produced for analysis by multiwavelength anomalous diffraction (MAD): a vehicle for direct determination of three-dimensional structure. EMBO J. 9, 1665–1672 (1990).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Weeks, C.M. & Miller, R. The design and implementation of SnB v2.0. J. Appl. Crystallogr. 32, 120–124 (1999).
Terwilliger, T.C. SOLVE and RESOLVE: automated structure solution and density modification. Methods Enzymol. 374, 22–37 (2003).
Collaborative Compuational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brunger, A.T. et al. Crystallography & NMR System: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Jogl, G., Tao, X., Xu, Y. & Tong, L. COMO: a program for combined molecular replacement. Acta Crystallogr. D Biol. Crystallogr. 57, 1127–1134 (2001).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Zhou, T. et al. Structure of human nicotinamide/nicotinic acid mononucleotide adenylyltransferase. Basis for the dual substrate specificity and activation of the oncolytic agent tiazofurin. J. Biol. Chem. 277, 13148–13154 (2002).
Jauch, R., Humm, A., Huber, R. & Wahl, M.C. Structures of Escherichia coli NAD synthetase with substrates and products reveal mechanistic rearrangements. J. Biol. Chem. 280, 15131–15140 (2005).
Kang, G.B. et al. Crystal structure of NH3-dependent NAD+ synthetase from Helicobacter pylori. Proteins 58, 985–988 (2005).
Carson, M. Ribbon models of macromolecules. J. Mol. Graph. 5, 103–106 (1987).
Evans, S.V. SETOR: hardware lighted three-dimensional solid model representations of macromolecules. J. Mol. Graph. 11, 134–138 (1993).
Acknowledgements
We thank J. Schwanof and R. Abramowitz for setting up the X4A and X4C beamlines at the National Synchrotron Light Source, Y. Shen and S. Xiang for help with data collection at the synchrotron source, M. Matsumoto, J. Buteau and D. Accili for help with the insulin assay, H. Zhang (University of Texas Southwestern Medical Center), M. Wahl (Max-Planck-Institute for Biophysical Chemistry) and S.H. Eom (Gwangju Institute of Science & Technology) for respectively providing the bacterial expression plasmids for human NMN/NAMN adenylyltransferase, E. coli NADS and Helicobacter pylori NADS, and W.W. Cleland for helpful discussions. This work was supported in part by a grant from the US National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Structures of NAPRTase and QAPRTase dimers (PDF 760 kb)
Supplementary Fig. 2
Structural changes upon NMN and FK866 binding (PDF 144 kb)
Rights and permissions
About this article
Cite this article
Khan, J., Tao, X. & Tong, L. Molecular basis for the inhibition of human NMPRTase, a novel target for anticancer agents. Nat Struct Mol Biol 13, 582–588 (2006). https://doi.org/10.1038/nsmb1105
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1105
This article is cited by
-
Chemoproteomic mapping of the glycolytic targetome in cancer cells
Nature Chemical Biology (2023)
-
Synthesis, and biological evaluation of EGFR/HER2-NAMPT conjugates for tumor treatment
Molecular Diversity (2023)
-
Discovery of small-molecule activators of nicotinamide phosphoribosyltransferase (NAMPT) and their preclinical neuroprotective activity
Cell Research (2022)
-
Glioma-Targeted Therapeutics: Computer-Aided Drug Design Prospective
The Protein Journal (2021)
-
Extracellular NAD+ enhances PARP-dependent DNA repair capacity independently of CD73 activity
Scientific Reports (2020)