Abstract
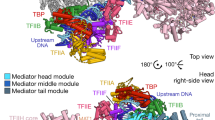
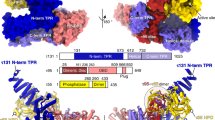
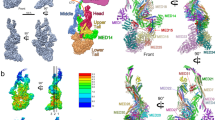
The Mediator head module stimulates basal RNA polymerase II (Pol II) transcription and enables transcriptional regulation. Here we show that the head subunits Med8, Med18 and Med20 form a subcomplex (Med8/18/20) with two submodules. The highly conserved N-terminal domain of Med8 forms one submodule that binds the TATA box–binding protein (TBP) in vitro and is essential in vivo. The second submodule consists of the C-terminal region of Med8 (Med8C), Med18 and Med20. X-ray analysis of this submodule reveals that Med18 and Med20 form related β-barrel folds. A conserved putative protein-interaction face on the Med8C/18/20 submodule includes sites altered by srb mutations, which counteract defects resulting from Pol II truncation. Our results and published data support a positive role of the Med8/18/20 subcomplex in initiation-complex formation and suggest that the Mediator head contains a multipartite TBP-binding site that can be modulated by transcriptional activators.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Malik, S. & Roeder, R.G. Transcriptional regulation through Mediator-like coactivators in yeast and metazoan cells. Trends Biochem. Sci. 25, 277–283 (2000).
Bjorklund, S. & Gustafsson, C.M. The mediator complex. Adv. Protein Chem. 67, 43–65 (2004).
Naar, A.M., Lemon, B.D. & Tjian, R. Transcriptional coactivator complexes. Annu. Rev. Biochem. 70, 475–501 (2001).
Kornberg, R.D. Mediator and the mechanism of transcriptional activation. Trends Biochem. Sci. 30, 235–239 (2005).
Bourbon, H.M. et al. A unified nomenclature for protein subunits of mediator complexes linking transcriptional regulators to RNA polymerase II. Mol. Cell 14, 553–557 (2004).
Boube, M., Joulia, L., Cribbs, D.L. & Bourbon, H.M. Evidence for a mediator of RNA polymerase II transcriptional regulation conserved from yeast to man. Cell 110, 143–151 (2002).
Nonet, M.L. & Young, R.A. Intragenic and extragenic suppressors of mutations in the heptapeptide repeat domain of Saccharomyces cerevisiae RNA polymerase II. Genetics 123, 715–725 (1989).
Thompson, C.M., Koleske, A.J., Chao, D.M. & Young, R.A. A multisubunit complex associated with the RNA polymerase II CTD and TATA-binding protein in yeast. Cell 73, 1361–1375 (1993).
Guglielmi, B. et al. A high resolution protein interaction map of the yeast Mediator complex. Nucleic Acids Res. 32, 5379–5391 (2004).
Kang, J.S. et al. The structural and functional organization of the yeast mediator complex. J. Biol. Chem. 276, 42003–42010 (2001).
Dotson, M.R. et al. Structural organization of yeast and mammalian mediator complexes. Proc. Natl. Acad. Sci. USA 97, 14307–14310 (2000).
Cantin, G.T., Stevens, J.L. & Berk, A.J. Activation domain-mediator interactions promote transcription preinitiation complex assembly on promoter DNA. Proc. Natl. Acad. Sci. USA 100, 12003–12008 (2003).
Ranish, J.A., Yudkovsky, N. & Hahn, S. Intermediates in formation and activity of the RNA polymerase II preinitiation complex: holoenzyme recruitment and a postrecruitment role for the TATA box and TFIIB. Genes Dev. 13, 49–63 (1999).
Holstege, F.C. et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95, 717–728 (1998).
Takagi, Y. & Kornberg, R.D. Mediator as a general transcription factor. J. Biol. Chem. 281, 80–89 (2006).
Lee, Y.C., Park, J.M., Min, S., Han, S.J. & Kim, Y.J. An activator binding module of yeast RNA polymerase II holoenzyme. Mol. Cell. Biol. 19, 2967–2976 (1999).
Koleske, A.J., Buratowski, S., Nonet, M. & Young, R.A. A novel transcription factor reveals a functional link between the RNA polymerase II CTD and TFIID. Cell 69, 883–894 (1992).
Koh, S.S., Ansari, A.Z., Ptashne, M. & Young, R.A. An activator target in the RNA polymerase II holoenzyme. Mol. Cell 1, 895–904 (1998).
Lee, T.I. et al. Interplay of positive and negative regulators in transcription initiation by RNA polymerase II holoenzyme. Mol. Cell. Biol. 18, 4455–4462 (1998).
Holm, L. & Sander, C. Dali: a network tool for protein structure comparison. Trends Biochem. Sci. 20, 478–480 (1995).
Lima, C.D., Wang, L.K. & Shuman, S. Structure and mechanism of yeast RNA triphosphatase: an essential component of the mRNA capping apparatus. Cell 99, 533–543 (1999).
Sismeiro, O., Trotot, P., Biville, F., Vivares, C. & Danchin, A. Aeromonas hydrophila adenylyl cyclase 2: a new class of adenylyl cyclases with thermophilic properties and sequence similarities to proteins from hyperthermophilic archaebacteria. J. Bacteriol. 180, 3339–3344 (1998).
Iyer, L.M. & Aravind, L. The catalytic domains of thiamine triphosphatase and CyaB-like adenylyl cyclase define a novel superfamily of domains that bind organic phosphates. BMC Genomics 3, 33 (2002).
Brower, C.S. et al. Mammalian mediator subunit mMED8 is an Elongin BC-interacting protein that can assemble with Cul2 and Rbx1 to reconstitute a ubiquitin ligase. Proc. Natl. Acad. Sci. USA 99, 10353–10358 (2002).
Baumli, S., Hoeppner, S. & Cramer, P. A conserved mediator hinge revealed in the structure of the MED7/MED21 (Med7/Srb7) heterodimer. J. Biol. Chem. 280, 18171–18178 (2005).
Hoeppner, S., Baumli, S. & Cramer, P. Structure of the mediator subunit cyclin C and its implications for CDK8 function. J. Mol. Biol. 350, 833–842 (2005).
van de Peppel, J. et al. Mediator expression profiling epistasis reveals a signal transduction pathway with antagonistic submodules and highly specific downstream targets. Mol. Cell 19, 511–522 (2005).
Yudkovsky, N., Ranish, J.A. & Hahn, S. A transcription reinitiation intermediate that is stabilized by activator. Nature 408, 225–229 (2000).
Zhu, X. et al. Genome-wide occupancy profile of mediator and the Srb8–11 module reveals interactions with coding regions. Mol. Cell 22, 169–178 (2006).
Andrau, J.C. et al. Genome-wide location of the coactivator mediator: Binding without activation and transient Cdk8 interaction on DNA. Mol. Cell 22, 179–192 (2006).
Johnson, K.M. & Carey, M. Assembly of a mediator/TFIID/TFIIA complex bypasses the need for an activator. Curr. Biol. 13, 772–777 (2003).
Wu, S.Y., Zhou, T. & Chiang, C.M. Human mediator enhances activator-facilitated recruitment of RNA polymerase II and promoter recognition by TATA-binding protein (TBP) independently of TBP-associated factors. Mol. Cell. Biol. 23, 6229–6242 (2003).
Taatjes, D.J., Naar, A.M., Andel, F., III, Nogales, E. & Tjian, R. Structure, function, and activator-induced conformations of the CRSP coactivator. Science 295, 1058–1062 (2002).
Liu, Y., Ranish, J.A., Aebersold, R. & Hahn, S. Yeast nuclear extract contains two major forms of RNA polymerase II mediator complexes. J. Biol. Chem. 276, 7169–7175 (2001).
Budisa, N. et al. High-level biosynthetic substitution of methionine in proteins by its analogs 2-aminohexanoic acid, selenomethionine, telluromethionine and ethionine in Escherichia coli. Eur. J. Biochem. 230, 788–796 (1995).
Meinhart, A., Blobel, J. & Cramer, P. An extended winged helix domain in general transcription factor E/IIEalpha. J. Biol. Chem. 278, 48267–48274 (2003).
Juo, Z.S., Kassavetis, G.A., Wang, J., Geiduschek, E.P. & Sigler, P.B. Crystal structure of a transcription factor IIIB core interface ternary complex. Nature 422, 534–539 (2003).
Armache, K.-J., Kettenberger, H. & Cramer, P. Architecture of the initiation-competent 12-subunit RNA polymerase II. Proc. Natl. Acad. Sci. USA 100, 6964–6968 (2003).
Leslie, A. in Joint CCP4 and ESF-EACMB Newsletter on Protein Crystallography No. 26 (Daresbury Laboratory, Warrington, UK, 1992).
Terwilliger, T.C. Automated structure solution, density modification and model building. Acta Crystallogr. D Biol. Crystallogr. 58, 1937–1940 (2002).
Roussel, A. & Cambillau, C. Turbo-FRODO. in Silicon Graphics Geometry, Partners Directory 77–78 (Silicon Graphics, Mountain View, California, USA, 1989).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Meth. Enzym. 276, 307–326 (1996).
McCoy, A.J., Grosse-Kunstleve, R.W., Storoni, L.C. & Read, R.J. Likelihood-enhanced fast translation functions. Acta Crystallogr. D Biol. Crystallogr. 61, 458–464 (2005).
Collaborative Computational Project, Number 4. The CCP4 Suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr 50, 760–763 (1994).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTAL W: improving the sensibility of progressive multiple sequence alignment through sequence weighing, positions-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Gouet, P., Courcelle, E., Stuart, D.I. & Metoz, F. ESPript: analysis of multiple sequence alignments in PostScript. Bioinformatics 15, 305–308 (1999).
DeLano, W.L. The PyMOL Molecular Graphics System (DeLano Scientific, San Carlos, California, USA, 2002).
Acknowledgements
We thank S. Baumli for discussions, K. Armache for help with TFIIB preparation and other members of the Cramer laboratory for help. This work was supported by a European Molecular Biology Organization long-term fellowship to L.L., by grants of the Deutsche Forschungsgemeinschaft, the Sonderforschungsbereich SFB646 and the Fonds der chemischen Industrie to P.C. and K.S. and by the EU-grant 3D repertoire, contract no. LSHG-CT-2005-512028. Part of this work was performed at the Swiss Light Source at the Paul Scherrer Institute, Villigen, Switzerland. We thank C. Schulze-Briese and his team at the Swiss Light Source for help.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Structure of the free Med18/20 heterodimer. (PDF 1508 kb)
Supplementary Fig. 2
Conserved intersubunit contacts within the Med8C/18/20 structure. (PDF 2278 kb)
Supplementary Fig. 3
Flexibility of the Med8C/18/20 trimer. (PDF 1562 kb)
Supplementary Table 1
Med18-Med20 contacts. (PDF 35 kb)
Supplementary Table 2
Med8-Med18 contacts. (PDF 30 kb)
Rights and permissions
About this article
Cite this article
Larivière, L., Geiger, S., Hoeppner, S. et al. Structure and TBP binding of the Mediator head subcomplex Med8–Med18–Med20. Nat Struct Mol Biol 13, 895–901 (2006). https://doi.org/10.1038/nsmb1143
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1143
This article is cited by
-
Transcription regulation by the Mediator complex
Nature Reviews Molecular Cell Biology (2018)
-
Core Mediator structure at 3.4 Å extends model of transcription initiation complex
Nature (2017)
-
Architecture of the RNA polymerase II–Mediator core initiation complex
Nature (2015)
-
Redefining the modular organization of the core Mediator complex
Cell Research (2014)
-
Reconstitution of active human core Mediator complex reveals a critical role of the MED14 subunit
Nature Structural & Molecular Biology (2014)