Abstract
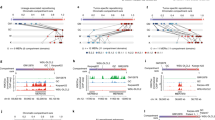
H2A.Z association with specific genomic loci is thought to contribute to a chromatin structure that promotes transcription activation. Acetylation of H2A.Z at promoters of oncogenes has been linked to tumorigenesis. The mechanism is unknown. Here, we show that in triple negative breast cancer cells, H2A.Z bound to the promoter of the constitutively, weakly expressed cyclin D1 oncogene (CCND1), a key regulator of cellular proliferation. Depleting the pool of H2A.Z stimulated transcription of CCND1 in the absence of its cognate transcription factor, the estrogen receptor (ER). During activation of CCND1, H2A.Z was released from the transcription start site (TSS) and downstream enhancer (enh2) sequences. Concurrently, acetylation of H2A.Z, H3 and H4 at the TSS was increased but only H2A.Z was acetylated at enh2. Acetylation of H2A.Z required the Tip60 acetyltransferase to be associated with the activated CCND1 on both TSS and enh2 sites. Depletion of Tip60 prevented CCND1 activation. Chromosome conformation capture experiments (3C) revealed specific contacts between the TSS and enh2 chromatin regions. These results suggest that release of a histone H2A.Z-mediated repression loop activates CCND1 for transcription. Our findings open new avenues for controlling and understanding aberrant gene expression associated with tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Frasor J, Danes JM, Komm B, Chang KC, Lyttle CR, Katzenellenbogen BS . Profiling of estrogen up- and down-regulated gene expression in human breast cancer cells: insights into gene networks and pathways underlying estrogenic control of proliferation and cell phenotype. Endocrinology 2003; 144: 4562–4574.
Soulez M, Parker MG . Identification of novel oestrogen receptor target genes in human ZR75-1 breast cancer cells by expression profiling. J Mol Endocrinol 2001; 27: 259–274.
Jensen EV, Desombre ER, Hurst DJ, Kawashima T, Jungblut PW . Estrogen-receptor interactions in target tissues. Arch Anat Microsc Morphol Exp 1967; 56: 547–569.
Chu S, Fuller PJ . Identification of a splice variant of the rat estrogen receptor beta gene. Mol Cell Endocrinol 1997; 132: 195–199.
Metivier R, Stark A, Flouriot G, Hubner MR, Brand H, Penot G et al. A dynamic structural model for estrogen receptor-alpha activation by ligands, emphasizing the role of interactions between distant A and E domains. Mol Cell 2002; 10: 1019–1032.
Ogawa S, Inoue S, Watanabe T, Orimo A, Hosoi T, Ouchi Y et al. Molecular cloning and characterization of human estrogen receptor betacx: a potential inhibitor ofestrogen action in human. Nucleic Acids Res 1998; 26: 3505–3512.
Thompson EW, Paik S, Brunner N, Sommers CL, Zugmaier G, Clarke R et al. Association of increased basement membrane invasiveness with absence of estrogen receptor and expression of vimentin in human breast c3ancer cell lines. J Cell Physiol 1992; 150: 534–544.
Ottaviano YL, Issa JP, Parl FF, Smith HS, Baylin SB, Davidson NE . Methylation of the estrogen receptor gene CpG island marks loss of estrogen receptor expression in human breast cancer cells. Cancer Res 1994; 54: 2552–2555.
Giamarchi C, Solanas M, Chailleux C, Augereau P, Vignon F, Rochefort H et al. Chromatin structure of the regulatory regions of pS2 and cathepsin D genes in hormone-dependent and -independent breast cancer cell lines. Oncogene 1999; 18: 533–541.
Touitou I, Vignon F, Cavailles V, Rochefort H . Hormonal regulation of cathepsin D following transfection of the estrogen or progesterone receptor into three sex steroid hormone resistant cancer cell lines. J Steroid Biochem Mol Biol 1991; 40: 231–237.
Bartkova J, Lukas J, Strauss M, Bartek J . Cell cycle-related variation and tissue-restricted expression of human cyclin D1 protein. J Pathol 1994; 172: 237–245.
Boulon S, Dantonel JC, Binet V, Vie A, Blanchard JM, Hipskind RA et al. Oct-1 potentiates CREB-driven cyclin D1 promoter activation via a phospho-CREB- and CREB binding protein-independent mechanism. Mol Cell Biol 2002; 22: 7769–7779.
Tetsu O, McCormick F . Beta-catenin regulates expression of cyclin D1 in colon carcinoma cells. Nature 1999; 398: 422–426.
Fu M, Wang C, Li Z, Sakamaki T, Pestell RG . Minireview: Cyclin D1: normal and abnormal functions. Endocrinology 2004; 145: 5439–5447.
Dalvai M, Bystricky K . Cell cycle and anti-estrogen effects synergize to regulate cell proliferation and ER target gene expression. PloS one 2010; 5: e11011.
Eeckhoute J, Carroll JS, Geistlinger TR, Torres-Arzayus MI, Brown M . A cell-type-specific transcriptional network required for estrogen regulation of cyclin D1 and cell cycle progression in breast cancer. Genes Dev 2006; 20: 2513–2526.
Lehn S, Tobin NP, Berglund P, Nilsson K, Sims AH, Jirstrom K et al. Down-regulation of the oncogene cyclin D1 increases migratory capacity in breast cancer and is linked to unfavorable prognostic features. Am J Pathol 2010; 177: 2886–2897.
Tobin NP, Sims AH, Lundgren KL, Lehn S, Landberg G . Cyclin D1, Id1 and EMT in breast cancer. BMC cancer 2011; 11: 417.
Carroll JS, Meyer CA, Song J, Li W, Geistlinger TR, Eeckhoute J et al. Genome-wide analysis of estrogen receptor binding sites. Nat Genet 2006; 38: 1289–1297.
Planas-Silva MD, Donaher JL, Weinberg RA . Functional activity of ectopically expressed estrogen receptor is not sufficient for estrogen-mediated cyclin D1 expression. Cancer Res 1999; 59: 4788–4792.
Lupien M, Eeckhoute J, Meyer CA, Wang Q, Zhang Y, Li W et al. FoxA1 translates epigenetic signatures into enhancer-driven lineage-specific transcription. Cell 2008; 132: 958–970.
Fleury L, Gerus M, Lavigne AC, Richard-Foy H, Bystricky K . Eliminating epigenetic barriers induces transient hormone-regulated gene expression in estrogen receptor negative breast cancer cells. Oncogene 2008; 27: 4075–4085.
Bhaumik SR, Smith E, Shilatifard A . Covalent modifications of histones during development and disease pathogenesis. Nat Struct Mol Biol 2007; 14: 1008–1016.
Dalvai M, Bystricky K . The role of histone modifications and variants in regulating gene expression in breast cancer. J Mammary Gland Biol Neoplasia 2010; 15: 19–33.
Hajkova P, Ancelin K, Waldmann T, Lacoste N, Lange UC, Cesari F et al. Chromatin dynamics during epigenetic reprogramming in the mouse germ line. Nature 2008; 452: 877–881.
Henikoff S, Ahmad K . Assembly of variant histones into chromatin. Annu Rev Cell Dev Biol 2005; 21: 133–153.
Metivier R, Penot G, Hubner MR, Reid G, Brand H, Kos M et al. Estrogen receptor-alpha directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter. Cell 2003; 115: 751–763.
Strahl BD, Allis CD . The language of covalent histone modifications. Nature 2000; 403: 41–45.
Suganuma T, Workman JL . Crosstalk among Histone Modifications. Cell 2008; 135: 604–607.
Gevry N, Hardy S, Jacques PE, Laflamme L, Svotelis A, Robert F et al. Histone H2A.Z is essential for estrogen receptor signaling. Genes Dev 2009; 23: 1522–1533.
Guillemette B, Bataille AR, Gevry N, Adam M, Blanchette M, Robert F et al. Variant histone H2A.Z is globally localized to the promoters of inactive yeast genes and regulates nucleosome positioning. PLoS Biol 2005; 3: e384.
Updike DL, Mango SE . Temporal regulation of foregut development by HTZ-1/H2A.Z and PHA-4/FoxA. Plos Genet 2006; 2: e161.
March-Diaz R, Garcia-Dominguez M, Lozano-Juste J, Leon J, Florencio FJ, Reyes JC . Histone H2A.Z and homologues of components of the SWR1 complex are required to control immunity in Arabidopsis. Plant J 2008; 53: 475–487.
Adam M, Robert F, Larochelle M, Gaudreau L . H2A.Z is required for global chromatin integrity and for recruitment of RNA polymerase II under specific conditions. Mol Cell Biol 2001; 21: 6270–6279.
Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z et al. High-resolution profiling of histone methylations in the human genome. Cell 2007; 129: 823–837.
Farris SD, Rubio ED, Moon JJ, Gombert WM, Nelson BH, Krumm A . Transcription-induced chromatin remodeling at the c-myc gene involves the local exchange of histone H2A.Z. J Biol Chem 2005; 280: 25298–25303.
Serandour AA, Avner S, Percevault F, Demay F, Bizot M, Lucchetti-Miganeh C et al. Epigenetic switch involved in activation of pioneer factor FOXA1-dependent enhancers. Genome Res 2011; 21: 555–565.
Valdes-Mora F, Song JZ, Statham AL, Strbenac D, Robinson MD, Nair SS et al. Acetylation of H2A.Z is a key epigenetic modification associated with gene deregulation and epigenetic remodeling in cancer. Genome Res 2011; 22: 307–321.
Vigushin DM, Ali S, Pace PE, Mirsaidi N, Ito K, Adcock I et al. Trichostatin A is a histone deacetylase inhibitor with potent antitumor activity against breast cancer in vivo. Clin Cancer Res 2001; 7: 971–976.
Barzily-Rokni M, Friedman N, Ron-Bigger S, Isaac S, Michlin D, Eden A . Synergism between DNA methylation and macroH2A1 occupancy in epigenetic silencing of the tumor suppressor gene p16(CDKN2A). Nucleic Acids Res 2011; 39: 1326–1335.
Bruzzese F, Leone A, Rocco M, Carbone C, Piro G, Caraglia M et al. HDAC inhibitor vorinostat enhances the antitumor effect of gefitinib in squamous cell carcinoma of head and neck by modulating ErbB receptor expression and reverting EMT. J Cell Physiol 2011; 226: 2378–2390.
Esteller M . Epigenetics in cancer. N Engl J Med 2008; 358: 1148–1159.
Ma X, Ezzeldin HH, Diasio RB . Histone deacetylase inhibitors: current status and overview of recent clinical trials. Drugs 2009; 69: 1911–1934.
Yang X, Phillips DL, Ferguson AT, Nelson WG, Herman JG, Davidson NE . Synergistic activation of functional estrogen receptor (ER)-alpha by DNA methyltransferase and histone deacetylase inhibition in human ER-alpha-negative breast cancer cells. Cancer Res 2001; 61: 7025–7029.
Shia WJ, Pattenden SG, Workman JL . Histone H4 lysine 16 acetylation breaks the genome’s silence. Genome Biol 2006; 7: 217.
Yun M, Wu J, Workman JL, Li B . Readers of histone modifications. Cell Res 2011; 21: 564–578.
Brookes E, Pombo A . Modifications of RNA polymerase II are pivotal in regulating gene expression states. EMBO Rep 2009; 10: 1213–1219.
Altaf M, Auger A, Covic M, Cote J . Connection between histone H2A variants and chromatin remodeling complexes. Biochem Cell Biol 2009; 87: 35–50.
Keogh MC, Mennella TA, Sawa C, Berthelet S, Krogan NJ, Wolek A et al. The Saccharomyces cerevisiae histone H2A variant Htz1 is acetylated by NuA4. Genes Dev 2006; 20: 660–665.
Millar CB, Xu F, Zhang K, Grunstein M . Acetylation of H2AZ Lys 14 is associated with genome-wide gene activity in yeast. Genes Dev 2006; 20: 711–722.
Dekker J, Rippe K, Dekker M, Kleckner N . Capturing chromosome conformation. Science 2002; 295: 1306–1311.
Splinter E, Grosveld F, de Laat W . 3C technology: analyzing the spatial organization of genomic loci in vivo. Methods Enzymol 2004; 375: 493–507.
Jin C, Zang C, Wei G, Cui K, Peng W, Zhao K et al. H3.3/H2A.Z double variant-containing nucleosomes mark ‘nucleosome-free regions’ of active promoters and other regulatory regions. Nat Genet 2009; 41: 941–945.
Voss TC, Schiltz RL, Sung MH, Yen PM, Stamatoyannopoulos JA, Biddie SC et al. Dynamic exchange at regulatory elements during chromatin remodeling underlies assisted loading mechanism. Cell 2011; 146: 544–554.
Ptashne M, Gann A . Transcriptional activation by recruitment. Nature 1997; 386: 569–577.
Jin F, Li Y, Ren B, Natarajan R . PU.1 and C/EBP(alpha) synergistically program distinct response to NF-kappaB activation through establishing monocyte specific enhancers. Proc Natl Acad Sci USA 2011; 108: 5290–5295.
Lee S, Miller M, Shuman JD, Johnson PF . CCAAT/Enhancer-binding protein beta DNA binding is auto-inhibited by multiple elements that also mediate association with p300/CREB-binding protein (CBP). J Biol Chem 2010; 285: 21399–21410.
Kimura A, Horikoshi M . Tip60 acetylates six lysines of a specific class in core histones in vitro. Genes cells 1998; 3: 789–800.
Yamamoto T, Horikoshi M . Novel substrate specificity of the histone acetyltransferase activity of HIV-1-Tat interactive protein Tip60. J Biol Chem 1997; 272: 30595–30598.
Ikura T, Tashiro S, Kakino A, Shima H, Jacob N, Amunugama R et al. DNA damage-dependent acetylation and ubiquitination of H2AX enhances chromatin dynamics. Mol Cell Biol 2007; 27: 7028–7040.
Bray F, Sankila R, Ferlay J, Parkin DM . Estimates of cancer incidence and mortality in Europe in 1995. Eur J Cancer 2002; 38: 99–166.
Jeong KW, Kim K, Situ AJ, Ulmer TS, An W, Stallcup MR . Recognition of enhancer element-specific histone methylation by TIP60 in transcriptional activation. Nat Struct Mol Biol 2011; 18: 1358–1365.
Ong CT, Corces VG . Enhancer function: new insights into the regulation of tissue-specific gene expression. Nat Rev Genet 2011; 12: 283–293.
Stadhouders R, Thongjuea S, Andrieu-Soler C, Palstra RJ, Bryne JC, van den Heuvel A et al. Dynamic long-range chromatin interactions control Myb proto-oncogene transcription during erythroid development. EMBO J 2011; 31: 986–999.
Tolhuis B, Palstra RJ, Splinter E, Grosveld F, de Laat W . Looping and interaction between hypersensitive sites in the active beta-globin locus. Mol Cell 2002; 10: 1453–1465.
Vakoc CR, Letting DL, Gheldof N, Sawado T, Bender MA, Groudine M et al. Proximity among distant regulatory elements at the beta-globin locus requires GATA-1 and FOG-1. Mol Cell 2005; 17: 453–462.
Vogelmann J, Valeri A, Guillou E, Cuvier O, Nollmann M . Roles of chromatin insulator proteins in higher-order chromatin organization and transcription regulation. Nucleus 2011; 2: 358–369.
Gamble MJ, Frizzell KM, Yang C, Krishnakumar R, Kraus WL . The histone variant macroH2A1 marks repressed autosomal chromatin, but protects a subset of its target genes from silencing. Genes Dev 2010; 24: 21–32.
Hua S, Kittler R, White KP . Genomic antagonism between retinoic acid and estrogen signaling in breast cancer. Cell 2009; 137: 1259–1271.
Fullwood MJ, Liu MH, Pan YF, Liu J, Xu H, Mohamed YB et al. An oestrogen-receptor-alpha-bound human chromatin interactome. Nature 2009; 462: 58–64.
Noordermeer D, de Wit E, Klous P, van de Werken H, Simonis M, Lopez-Jones M et al. Variegated gene expression caused by cell-specific long-range DNA interactions. Nat Cell Biol 2011; 13: 944–951.
Simonis M, Klous P, Splinter E, Moshkin Y, Willemsen R, de Wit E et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat Genet 2006; 38: 1348–1354.
Chambeyron S, Bickmore WA . Does looping and clustering in the nucleus regulate gene expression? Curr Opin Cell Biol 2004; 16: 256–262.
Kocanova S, Kerr EA, Rafique S, Boyle S, Katz E, Caze-Subra S et al. Activation of estrogen-responsive genes does not require their nuclear co-localization. Plos Genet 2010; 6: e1000922.
Mattera L, Courilleau C, Legube G, Ueda T, Fukunaga R, Chevillard-Briet M et al. The E1A-associated p400 protein modulates cell fate decisions by the regulation of ROS homeostasis. Plos Genet 2010; 6: e1000983.
Mattera L, Escaffit F, Pillaire MJ, Selves J, Tyteca S, Hoffmann JS et al. The p400/Tip60 ratio is critical for colorectal cancer cell proliferation through DNA damage response pathways. Oncogene 2009; 28: 1506–1517.
Iacovoni JS, Caron P, Lassadi I, Nicolas E, Massip L, Trouche D et al. High-resolution profiling of gammaH2AX around DNA double strand breaks in the mammalian genome. EMBO J 2010; 29: 1446–1457.
Legube G, Linares LK, Tyteca S, Caron C, Scheffner M, Chevillard-Briet M et al. Role of the histone acetyl transferase Tip60 in the p53 pathway. J Biol Chem 2004; 279: 44825–44833.
Deschenes J, Bourdeau V, White JH, Mader S . Regulation of GREB1 transcription by estrogen receptor alpha through a multipartite enhancer spread over 20 kb of upstream flanking sequences. J Biol Chem 2007; 282: 17335–17339.
Acknowledgements
We would like to thank Dr Didier Trouche for the Tip60 expressing vector and insightful discussions, J Eeckhoute, Lille for sharing unpublished work, and David Laperriere, IRIC, Montreal, for expert advice on genome database use. This work was supported by the Ligue Nationale Contre le Cancer (fellowship to LB), the Institut National du Cancer (INCa grant no. 34696) and the Fondation pour la Recherche Médicale (FRM, fellowship to MD).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Dalvai, M., Bellucci, L., Fleury, L. et al. H2A.Z-dependent crosstalk between enhancer and promoter regulates Cyclin D1 expression. Oncogene 32, 4243–4251 (2013). https://doi.org/10.1038/onc.2012.442
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2012.442
Keywords
This article is cited by
-
Loss of TIP60 (KAT5) abolishes H2AZ lysine 7 acetylation and causes p53, INK4A, and ARF-independent cell cycle arrest
Cell Death & Disease (2022)
-
Non-canonical roles of canonical telomere binding proteins in cancers
Cellular and Molecular Life Sciences (2021)
-
DNA repair complex licenses acetylation of H2A.Z.1 by KAT2A during transcription
Nature Chemical Biology (2019)
-
The H2A.Z histone variant integrates Wnt signaling in intestinal epithelial homeostasis
Nature Communications (2019)