Abstract
The restriction (R)-point decision is fundamental to normal differentiation and the G1–S transition, and the decision-making machinery is perturbed in nearly all cancer cells. The mechanisms underlying the cellular context–dependent R-point decision remain poorly understood. We found that the R-point was dysregulated in Runx3−/−mouse embryonic fibroblasts (MEFs), which formed tumors in nude mice. Ectopic expression of Runx3 restored the R-point and abolished the tumorigenicity of Runx3−/−MEFs and K-Ras–activated Runx3−/−MEFs (Runx3−/−;K-RasG12D/+). During the R-point, Runx3 transiently formed a complex with pRb and Brd2 and induced Cdkn1a (p21Waf1/Cip1/Sdi1; p21), a key regulator of the R-point transition. Cyclin D–CDK4/6 promoted dissociation of the pRb–Runx3–Brd2 complex, thus turning off p21 expression. However, cells harboring oncogenic K-Ras maintained the pRb–Runx3–Brd2 complex and p21 expression even after introduction of Cyclin D1. Thus, Runx3 plays a critical role in R-point regulation and defense against cellular transformation.
Similar content being viewed by others
Introduction
The restriction (R)-point is a critical event in which a mammalian cell makes the decision in response to mitogen stimulation. After this decision, depending on its differentiation stage, the cell either remains in early G1, retreats from the active cycle into G0, or advances into late G1.1, 2, 3 The R-point transition is governed by the R-point–associated proteins (R-proteins), which include c-Myc, Cyclins, CDKs, p21, p27, E2F, and pRB,4 with pRB serving as the primary molecular regulator 2. In early and mid-G1 phase, Cyclin D–CDK4/6 phosphorylates pRB prior to the R-point gate. Although p21 was originally identified as a CDK inhibitor,5 subsequent studies revealed that p21 family members activate Cyclin D–CDK4/6 by stimulating the association of its component proteins, whereas they inhibit Cyclin E–CDK2, Cyclin A–CDC2, and Cyclin B–CDC2.6, 7 Therefore, p21 promotes entry into the R-point at early/mid-G1 phase by activating Cyclin D–CDK4/6 but prevents further progression through the R-point by inhibiting Cyclin E–CDK2.2, 8
The postmitotic interval of G1, which lasts from mitosis to the R-point, is remarkably constant (3–4 h) in all tested cell lines.9, 10, 11, 12 p21 is induced 1 h after mitogenic stimulation and reduced to basal level 4 h after in mouse embryonic fibroblast cells (MEFs),13 consistent with the idea that p21 plays critical roles in R-point regulation. Moreover, p21 also has an important function in the G1 checkpoint.14 Accumulating evidence indicates that deregulation of the R-point is associated with the formation of most cancer cells.4 To date, however, it remains unclear how the R-point decision is made in a cell context–dependent manner, as well as how the R-point is disturbed in cancer cells.
MEFs are a classic model system for studying cell immortalization and transformation; this model exhibits clear parallels to key genetic alterations that occur during human tumorigenesis.15 In general, the p53 pathway is inactivated during MEF immortalization, but the immortalized MEFs are not tumorigenic, which requires activation of oncogenes.15, 16 Although overexpression of oncogenic Ras in primary MEFs induces senescence through the p53 pathway,17, 18 oncogenic K-Ras expressed at endogenous levels in primary MEFs can overcome p53 pathway–mediated defenses.19, 20 However, the immortalized K-Ras activated MEFs are not tumorigenic in immuno-compromised mice.19, 20 Therefore, it is likely that additional barriers exist against endogenous oncogenic K-Ras–induced tumor development, although these mechanisms remain to be elucidated.
Runx3, which plays pivotal roles in lineage determination and functions as a tumor suppressor, is frequently inactivated in multiple types of human tumors.21 Runx3 regulates the cell cycle and apoptosis by inducing p21 and Arf in a context-dependent manner, and deletion of Runx3 from mouse lungs results in development of lung adenoma.22 These observations raise questions regarding how Runx3 deletion induces cancer without oncogene activation. In this study, we found that the R-point is disturbed in Runx3−/− MEFs, which develop into tumors in nude mice. Expression of Runx3 restored the R-point and abolished the tumorigenicity of both Runx3−/− and K-Ras–activated Runx3−/− MEFs, demonstrating that Runx3 is an essential component of a R-point–associated barrier to tumorigenesis. We also found that the pRb–Runx3–Brd2 complex formed early after mitogenic stimulation, and dissociated after CDK4/6 activation to allow transient p21 induction. Together, our results indicate that cells can monitor their own status via the pRb–Runx3–Brd2 complex, allowing them to make context-dependent R-point decisions, and further demonstrate that disruption of this monitoring system is associated with tumorigenesis.
Results
Immortalized Runx3−/− MEFs are tumorigenic in nude mice
Floxed alleles of Runx3f/+, Runx3f/f, Rx3f/+;K-RasLSL-G12D/+, and Rx3f/f;K-RasLSL-G12D/+ MEFs were targeted by an adenovirus carrying Cre recombinase (Ad-Cre) (Figure 1a). The targeted MEFs were named Rx3+/−, Rx3−/−, Rx3+/−;K-Ras*, and Rx3−/−;K-Ras*, respectively. Expression of WT and Runx3-targeted alleles was confirmed by RT-PCR; the Runx3-targeted allele expresses a short Runx3 mRNA lacking exon 4 (Figure 1b, Supplementary Figures S1A and S1B).
Runx3−/− MEFs bearing p53 mutation develop into tumors in nude mice. (a) Targeting of Runx3flox and K-RasLSL-G12D alleles by Ad-Cre in MEFs was verified by genomic PCR. PCR primers were described previously.22 (b) Expression of Runx3 mRNA. Total mRNA was obtained from Rx3+/+ (WT), Rx3+/− (Runx3f/+ treated with Ad-Cre), and Rx3−/− (Runx3f/f treated with Ad-Cre) MEFs, and expression of Runx3 was measured by RT-PCR using the primer pairs shown in Supplementary Figures 1A and B. Arrows indicate Runx3 mRNAs from WT alleles, and arrowheads indicate those from exon 4–deleted alleles. (c) Growth curves of Rx3+/+, Rx3+/−, and Rx3−/− MEFs cultivated according to a 3T3 protocol. The Y-axis indicates the estimated number of accumulated cells. Rx3−/−#1 and Rx3−/−#2 MEFs were derived from two independent embryos. (d) IB analysis of p53 and its phosphorylation at Ser-15 in immortalized or early passage (p3 = passage 3) MEFs, either not treated (−) or treated (+) with 1 μM doxorubicin (Doxo) for 6 h. (e) Immortalized Rx3+/− and Rx3−/− (#1 and #2) MEFs were injected into the left and right sides, respectively, of the back of each nude mouse. The mice were photographed 36 days later, with tumors indicated by circles. (f) IB analysis of ERT-Runx3 (ERT-Rx3) expression in Rx3−/−;ERT-Rx3 (#1 and #2) MEFs in the absence of inducer. Rx3−/− MEFs stably transfected with empty vector (Rx3−/−;Vec) were used as negative controls. (g) Subcellular localization of ERT-Rx3 in the presence or absence of inducer (4-OHT = 4-hydroxy tamoxifen), as determined by subcellular fractionation followed by IB. Very low levels of inducer-independent nuclear ERT-Rx3 are indicated by arrows. (h) Rx3−/−;Vec (#1 and #2) and Rx3−/−;ERT-Rx3 (#1 and #2) MEFs were injected into the left and right sides, respectively, of the back of each nude mouse. The mice were photographed 36 days later, with tumors indicated by circles. The same mice were viewed from the left, middle and right sides.
MEFs were cultivated according to a 3T3 protocol; the growth curves of Rx3+/+, Rx3+/−, and two lines of Rx3−/− (#1 and #2) MEFs derived from independent embryos are shown in Figure 1c. All Rx3+/+, Rx3+/−, and Rx3−/−MEF lines entered into a characteristic cell-cycle arrest known as senescence after passage 6 (Figure 1c). Analysis of p53 status revealed that p53 was mutated during the immortalization of MEFs of all three genotypes. In early passage (p3) MEFs, p53 was phosphorylated and stabilized, in response to the DNA-damaging agent doxorubicin (Doxo); however, in immortalized MEFs, p53 was phosphorylated, but its level was unchanged (Figure 1d).
Notably, both immortalized Rx3−/− #1 and #2 MEFs induced rapidly growing tumors in nude mice (Figure 1e, right rear), whereas Rx3+/− MEFs failed to induce tumors (Figure 1e, left rear). The ERT inducible system is designed to trigger activation of ectopically expressed genes upon treatment with inducer. To determine whether Runx3 expression would abolish the tumorigenicity of Rx3−/− MEFs, Rx3−/−#1 and Rx3−/−#2 MEFs were stably transfected with pCS4-ERT1-Runx3 plasmid or empty vector, yielding the cell lines Rx3−/−;ERT-Rx3 (#1 and #2) and Rx3−/−;Vec (#1 and #2) (Figure 1f). Subcellular fractionation revealed that, in the absence of inducer, very low levels of ERT-Runx3 protein were present in the nucleus, suggesting that the ERT inducible system is slightly leaky in MEFs (Figure 1g). However, when injected into nude mice, the Rx3−/−;ERT-Rx3 #1 and #2 MEF lines did not form any tumors in the absence of inducer (Figure 1h). These results indicate that loss of Runx3 is a critical molecular event in MEF tumorigenesis, and that a very low level of inducer-independent leaky activation of ERT-Runx3 is sufficient to abolish the tumorigenicity of Rx3−/− MEFs in a p53-independent manner.
Expression of Runx3 abolishes tumorigenicity of Rx3−/−;K-RasG12D MEFs
Subsequently, Rx3+/−;K-Ras* #1, #2 and Rx3−/−;K-Ras* #1, #2 MEFs, derived from independent embryos, were immortalized using a 3T3 protocol. Growth curves of these MEFs are shown in Figure 2a. Both Rx3+/−;K-Ras* and Rx3−/−;K-Ras* MEFs continued to proliferate without replicative stress–induced crisis, as reported previously.19, 20 Rx3−/−;K-Ras* MEFs proliferated more quickly than Rx3+/−;K-Ras* MEFs until passage 20, suggesting that Runx3 decreased the proliferation rate of the K-Ras–activated MEFs (Figure 2a). However, after passage 24, the proliferation rate of Rx3+/−;K-Ras* MEFs increased and became equivalent to that of Rx3−/−;K-Ras* MEFs (Figure 2a). These results suggested that Runx3 was inactivated in Rx3+/−;K-Ras* MEFs during immortalization (see below).
Expression of Runx3 abolished the tumorigenicity of Rx3−/−;K-RasG12D/+ MEFs. (a) Growth curves of Rx3+/−;K-Ras* (#1 and #2) and Rx3−/−;K-Ras* (#1 and #2) MEFs, cultivated according to a 3T3 protocol. The Y-axis indicates the estimated number of accumulated cells. (b) Rx3+/−;K-Ras* (#1 and #2) and Rx3−/−;K-Ras* (#1 and #2) MEFs were injected into the left and right sides, respectively, of the back of each nude mouse, and photographs were taken 36 days later. (c) Hematoxylin–eosin staining of tumors generated by Rx3−/−, Rx3−/−;K-Ras*, and Rx3+/−;K-Ras* MEFs. (d) Levels of ERT-Runx3 in the indicated MEFs, measured by IB. (e) Proliferation rates of MEFs of the indicated genotypes. (f) Rx3−/−;K-Ras*;Vec (#1 and #2) and Rx3−/−;K-Ras*;ERT-Rx3 (#1 and #2) MEFs were injected into the left and right sides, respectively, of the back of each nude mouse. (g) RT-PCR analysis of Runx1, Runx2, and Runx3 mRNA levels in Rx3+/+, Rx3+/−, Rx3−/−;K-Ras* (#1 and #2), and Rx3+/−;K-Ras* (#1 and #2) MEFs. Predicted sizes of the Runx3 cDNAs from wild-type Runx3 mRNA are indicated. The four Runx3 PCR products amplified from Rx3+/− MEFs using two different primer pairs are indicated by circles. These PCR products were clearly present in Rx3+/−, but not in Rx3+/−;K-Ras* (#1 and #2) MEFs. (h) Amplification of the CpG island region of Runx3 by unmethylated (UM) or methylated (M) DNA–specific PCR from Rx3+/+, Rx3+/−, Rx3−/−;K-Ras*, and Rx3+/−;K-Ras* MEFs. Mw = molecular size markers.
Although immortalized Rx3+/+;K-Ras* MEFs (only K-RasLSL-G12Dallele was targeted by Cre recombinase) failed to develop into tumors in nude mice,19, 20 both Rx3+/−;K-Ras* and Rx3−/−;K-Ras* MEFs developed into tumors of similar size in nude mice (Figure 2b). Microscopic analysis of these tumors revealed that the Rx3−/−, Rx3+/−;K-Ras*, and Rx3−/−;K-Ras* MEFs developed into the same type of sarcoma, differing only in regard to the cell densities within the tumors: specifically, the densities of Rx3+/−;K-Ras* and Rx3−/−;K-Ras* MEFs were higher than that of Rx3−/− MEFs (Figure 2c).
To understand the tumor-suppressive effect of Runx3 against oncogenic K-Ras, pCS4-ERT1-Runx3 plasmid or empty vector was stably transfected into Rx3−/−;K-Ras* #1 and #2 MEFs, yielding Rx3−/−;K-Ras*;ERT-Rx3 (#1 and #2) and Rx3−/−;K-Ras*;Vec (#1 and #2) MEFs (Figure 2d). Because Runx3 suppresses MEF proliferation, the Rx3−/−;K-Ras*;ERT-Rx3 MEFs proliferated at a lower rate than Rx3−/−;K-Ras*;Vec MEFs (Figure 2e). Notably, both lines of Rx3−/−;K-Ras*;ERT-Rx3 MEFs failed to develop into tumors in nude mice in the absence of inducer (Figure 2f). These results demonstrate that Runx3 plays a critical role in defenses against tumorigenesis, even when K-Ras is activated.
Runx3 is selectively silenced in Rx3+/−;K-RasG12D MEFs
In light of these results, why did the Rx3+/−;K-Ras* MEFs, which retain a WT Runx3 allele, become tumorigenic (Figure 2b)? To address this question, we assessed Runx3 mRNA levels by RT-PCR in various MEFs. In Rx3+/−;K-Ras* MEFs, Runx3 mRNA was barely detectable, in contrast to the Runx1 and Runx2 mRNAs, which were readily detected (Figure 2g, Supplementary Figure S2A). Since RUNX3 is frequently silenced by DNA methylation in various types of cancers,21 we measured CpG island methylation of the Runx3 p2 promoter region. Methylation-specific PCR of the CpG island of the Runx3 p2 promoter region revealed that Runx3 was silenced by DNA methylation only in Rx3+/−;K-Ras* MEFs (Figure 2h, Supplementary Figure S2B). However, the targeted allele of Runx3 in Rx3−/−;K-Ras* MEFs was not silenced (Figures 2g and h), suggesting that Runx3 is not actively silenced by oncogenic K-Ras; instead, Runx3-silenced cells are preferentially selected during culture of K-Ras–activated cells. These results suggest that Rx3+/−;K-Ras* MEFs are equivalent to Rx3−/−;K-Ras* MEFs in terms of loss of Runx3 activity, and further confirm the critical role of Runx3 in defense against tumorigenesis.
The R-point is disturbed in Runx3 deleted MEFs
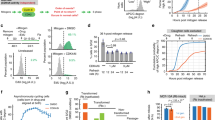
The concept of the R-point was established based on the observation that cells remain in G0/G1 if serum is removed before the R-point, whereas serum removal no longer affects cell-cycle progression if the cells have already passed through this point.3 To determine whether Runx3 plays a role in the R-point decision, we measured the minimum serum exposure time required for progression into S phase. For this purpose, we treated serum-starved MEFs with 10% serum for a limited time, and then further cultured them under serum-free conditions. FACS analysis 18 h after initial serum treatment revealed that Rx3−/− MEFs required only 1–2 h of exposure to serum, whereas Rx3+/+ MEFs required at least 4 h of exposure for entry into S phase (Figure 3a). This abnormally short serum exposure time for S-phase entry of Rx3−/− MEFs is essentially the same as that of Rb−/− MEFs,23 indicating that the R-point is disturbed in Rx3−/− MEFs. Notably, the R-point was completely restored by leaky activation of ERT-Runx3 (Figure 3a, Rx3−/−;ERT-Rx3 #1 and #2). These results, along with the critical role of R-point in tumorigenesis,1 suggest that the tumorigenicity of Rx3−/− MEFs is closely associated with the disruption of the R-point.
Deletion of Runx3 disturbs R-point in MEFs. (a) Immortalized MEFs were synchronized by serum deprivation for 24 h, stimulated with 10% serum for the indicated times, and then cultured for additional periods of time under serum starvation conditions. Eighteen hours after serum treatment, cells were harvested, and the proportion of S-phase cells was measured by FACS analysis. (b) Rx3+/+and Rx3−/− MEFs (three independent lines each) were synchronized by serum deprivation and treated with 10% serum for 2 h. RNA was extracted from the MEFs, and the gene expression pattern was analyzed by mRNA sequencing. Expression 2 h after serum stimulation was quantified as the log2 of the fold change relative to the average of control reactions (i.e., before serum stimulation) of Rx3+/+and Rx3−/− MEFs. Differential expression changes were analyzed by plotting log2(ERx3+2h/ERx3+0h) and log2(ERx3-2h/ERx3-0h). ERx3+0h, ERx3+2h, ERx3-2h, and ERx3-0h are the average expression levels of genes in Rx3+/+ or Rx3−/− MEFs 0 or 2 h after serum stimulation. Group A and group B indicate genes induced or suppressed by serum stimulation, respectively. Gray and red spots indicate genes regulated in a Runx3-independent and Runx3-dependent manner, respectively (FDR<0.001, P<0.05). (c) Runx3-dependent genes involved in major signaling pathways. (d) Expression levels of p21 and p27 before and 2 h after serum stimulation in Rx3+/+and Rx3−/− MEFs, obtained from RNA sequencing data. Relative expression levels of p21 and p27 at the indicated times are depicted by the bar graph. (e) Rx3+/+ and Rx3−/− MEFs were synchronized by serum deprivation and treated with 10% serum for the indicated times. The time course of p21 expression levels was determined by Northern blotting (NB). Gapdh mRNA was used as a loading control. (f) Levels or phosphorylation status of early G1–associated proteins in similarly treated Rx3+/+ and Rx3−/− MEFs, measured by IB. (g) Rx3−/− and Rx3−/−;ERT-Rx3 MEFs were synchronized by serum deprivation and treated with 10% serum for the indicated times. The time course of p21 expression levels was determined by Northern blotting.
To understand how Runx3 contributes to R-point commitment, we performed mRNA sequencing (RNA-seq) to identify genes differentially expressed at the R-point. For this purpose, Rx3+/+ and Rx3−/− MEFs were synchronized by serum deprivation, and total RNA was extracted from the MEFs before (0 h) and 2 h after serum treatment. Analysis of gene expression patterns revealed that 989 and 1442 genes were induced and suppressed, respectively (|log2(fold change)| ⩾1), in Rx3+/+ MEFs 2 h after serum stimulation. Detailed RNA-seq results are provided in the Excel file Rx3+;Rx3-.xlsx (Supplementary dataset 1 of Supplementary information). Two-dimensional plotting of the expression changes in response to serum stimulation in Rx3+/+ MEFs and Rx3−/− MEFs revealed that most of the differentially expressed genes were either induced (group A in Figure 3b) or suppressed (group B in Figure 3b) commonly in both strains of MEFs. However, expression changes in 65 genes differed significantly (P<0.05) between Rx3+/+ and Rx3−/− MEFs (red spots in Figure 3b), indicating that about 2.7% of early G1–associated genes were regulated in a Runx3-dependent manner. The Runx3-dependent genes involved in major signaling pathways are listed in Figures 3c, and a comparison of up-/downregulation of the 65 genes in Rx3+/+ and Rx3−/− MEFs is provided in Supplementary Figure S3.
p21 (Cdkn1a) and p27 (Cdkn1b) play key roles in facilitating R-point entry by activating Cyclin D–Cdk4/6. Our RNA-seq analysis revealed that p21 was induced 8.3-fold 2 h after serum stimulation in Rx3+/+ MEFs, but not in Rx3−/− MEFs (Figure 3d). By contrast, p27 was suppressed 2 h after serum stimulation in both Rx3+/+ and Rx3−/− MEFs (Figure 3d). Northern blotting (NB) analysis confirmed that p21 expression was induced 1–2 h after serum stimulation in Rx3+/+ MEFs, but not in Rx3−/− MEFs (Figure 3e). However, the expression patterns of p27, Cyclin D1, p-Erk1/2, and p-Akt, which play roles in R-point entry, were not affected by the presence or absence of Runx3 (Figure 3f). Consistent with the observation that expression of ERT-Rx3 in Rx3−/− MEFs restored the R-point, ERT-Rx3 expression also restored serum-stimulated p21 induction (Figure 3g). These results demonstrate that p21, a key regulator of the R-point, is a target of Runx3.
RUNX3 and pRB form a complex in response to serum stimulation and induce p21
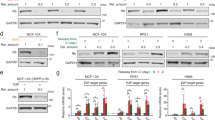
Based on the finding that deletion of either Runx3 or Rb disturbed the R-point in MEFs, we investigated the possibility that Runx3 and pRb function together by forming a complex. Immunoprecipitation (IP) with anti-Runx3 antibody followed by IB with an anti-pRB antibody revealed that the Runx3–pRb interaction occurs 1 h after serum stimulation in Rx3+/+ MEFs (Figure 4a). The Runx3–pRb interaction was maintained up until 2 h after stimulation, and decreased after 4 h (Figure 4a). The Runx3–pRb interaction and p21 induction were also observed 1–2 h after serum stimulation in HEK293 human embryonic kidney cells (Figure 4b). Specific interaction between RUNX3 and pRB was confirmed by IP with IgG or Anti-RUNX3 antibody followed by IB with anti pRB antibody in MEFs (Supplementary Figure S4A) and HEK293 cells (Supplementary Figure S4B). Exogenously expressed Myc-tagged RUNX3 (Myc-Rx3) and HA-tagged RB (HA-RB) also interacted transiently at similar time points after serum stimulation in HEK293 cells (Figure 4c). Mapping analysis of the interacting domains revealed that the A-pocket and B-pocket regions of pRB interact with RUNX3 (Supplementary Figure S4C), and that both the Runt domain and C-terminal region of RUNX3 interact with pRB (Supplementary Figure S4D).
RUNX3 and pRB form a complex in response to serum stimulation and induce p21. (a) WT MEFs were serum-starved for 40 h and treated with 10% serum. The expression levels of endogenous pRB and RUNX3, and the interaction between the two endogenous proteins, were measured by IP and IB with anti-RUNX3 antibody and anti-pRB antibody, respectively, at the indicated time points. (b) HEK293 cells were serum-starved for 24 h and treated with 10% serum. The expression levels of endogenous pRB and RUNX3, and the interaction between the two endogenous proteins, were measured as described above. (c) HEK293 cells were transfected with Myc-RX3 and HA-RB. Starting 24 h post-transfection, the cells were serum-starved for 24 h and stimulated with serum, and the expression levels of the transfected genes and the RUNX3–pRB interaction were measured by IP and IB, respectively. (d) Wild-type RUNX3 and RUNX3 deletion mutants were transfected into HEK293 cells either with or without RB. β-Gal was co-transfected as an internal control. The reporter activity of p21 promoter–driven luciferase was measured by luciferase assay. A schematic diagram of the RUNX3 deletion constructs is shown. Runt=Runt domain. (e) HEK293 cells were treated with si-RB RNA or si-control RNA for 24 h, cultured under serum-free conditions for 24 h, and then treated with serum for the indicated times. The expression levels of RB, RUNX3, Cyclin D1 (CycD1), and p21 were measured by IB. (f) HEK293 cells were treated with either control siRNA or RX3-specific siRNA for 24 h, and then serum-starved for 24 h. The cells were then treated with 10% serum, and the binding of pRB to the p21 promoter was measured by ChIP analysis at the indicated time points. One-thirtieth of the lysates were PCR amplified for input. Knockdown of RUNX3 by specific siRNA was verified by IB.
The RUNX3–pRB complex binds to and activates the p21 promoter in response to serum stimulation
Because HEK293 cells and MEFs exhibited essentially identical responses to serum stimulation with regard to the pRB–RUNX3 interaction and p21 induction, we used HEK293 cells to analyze the role of the pRB–RUNX3 complex on p21 induction, as well as the molecular mechanism underlying the association/dissociation of the complex. Previously, we reported that the p21 promoter contains three RUNX-binding sites that are critical for the RUNX3-mediated transactivation of the promoter.24 In a p21 promoter–reporter assay, RUNX3 alone increased reporter activity by about 5-fold, whereas the transactivation activity of RB alone was very weak (Figure 4d). However, co-expression of RUNX3 and RB increased reporter activity up to 11-fold, suggesting that RUNX3 and pRB function synergistically on the p21 promoter (Figure 4d). Similar experiments using a construct in which the Runt domain (DNA-binding domain) of RUNX3 was deleted revealed that this domain is required for functional cooperation between pRB and RUNX3 (Figure 4d). Consistent with this, serum-stimulated induction of p21 expression was abolished by siRNA-mediated RB knockdown (Figure 4e). These results suggest that RUNX3 and pRB function together to induce p21 expression.
The reporter assay also showed that co-expression of RB and RUNX3 had no effect on a p21 reporter containing a mutated RUNX-binding site (p21-mABC-Luc) (Figure 4d). Therefore, we performed chromatin immunoprecipitation (ChIP) to ascertain whether the RUNX3–pRB complex binds to the p21 promoter region. For this purpose, HEK293 cells cultured under serum-free conditions for 24 h were stimulated with serum. At various time points after serum stimulation, IP with an anti-RUNX3 or anti-pRB antibody followed by PCR amplification revealed that RUNX3 and pRB bound to the p21 promoter region 1–2 h after serum stimulation (Figure 4f). However, pRB–p21 promoter binding was markedly reduced after siRNA-mediated knockdown of RUNX3 (Figure 4f). Knockdown of RUNX3 by si-RUNX3 was efficient (Figure 4f). These results suggest that pRB binds to the p21 promoter by interacting with RUNX3, 1–2 h after serum stimulation.
The RUNX3–pRB complex is dissociated by Cyclin D1–CDK4/6
The dissociation of RUNX3–pRB occurred 4 h after serum stimulation, and Cyclin D1 was induced at the same time point (Figure 4b), suggesting that the RUNX–pRB complex forms before pRB is phosphorylated by CDK4/6. To confirm this observation, we mutated the nine N-terminal Ser/Thr residues (RB-M1-9), seven C-terminal Ser/Thr residues (RB-M10-16), or all Ser/Thr residues (RB-1-16) within pRB to Ala residues (Supplementary Figure S5A). The expression levels of RBs mutated at Ser/Thr residues were much lower than those of wild-type pRB (Figure 5a, Supplementary Figure S5B). However, the RB-M10-16 mutant interacted with RUNX3 more strongly than wild-type pRB (Figure 5a), supporting the idea that RUNX3 preferentially interacts with hypo-phosphorylated pRB. Consistent with this, the p21 promoter–reporter assay revealed that RUNX3 cooperated more effectively with the mutated pRBs than wild-type pRB in activating the p21 promoter (Figure 5b). These results demonstrate that RUNX3 preferentially interacts with hypo-phosphorylated pRB and induces p21 expression, and suggest that pRB phosphorylation may inhibit the RUNX3–pRB interaction.
Cyclin D–CDK4/6 inhibits the RUNX3–pRB interaction. (a) HA-RB wild-type and HA-RB-M10-16RB (phosphorylation sites–mutated RB) (Supplementary Figure 5A) were transfected to HEK293 cells with Myc-RUNX3. pRB was immunoprecipitated with an anti-HA antibody, and RUNX3 was detected with an anti-Myc- antibody in the immunoprecipitates. (b) Wild-type RB and RB mutated at its phosphorylation sites (RB-M1-9, RB-M10-16, and RB-M1-16) (Supplementary Figure 5A) were transfected into HEK293 cells either with or without RUNX3. The reporter activities of WT p21-promoter-luciferase (p21-WT) or RUNX-binding site–mutated p21-promoter-luciferase (p21-mABC)24 were measured using a luciferase assay. A, B, and C indicate the three RUNX3-binding sites in p21 promoter. (c) HEK293 cells were transfected with Myc-RUNX3 and HA-RB. Starting 24 h post-transfection, the cells were serum-starved for 1 day, and then treated with 200 nM CDK4 inhibitor for 1 h. The cells were then stimulated with serum, and the RUNX3–pRB interaction was measured by IP and IB at the indicated time points. (d) HEK293 cells were transfected with a fixed amount of RUNX3 (0.2 μg) and RB (0.6 μg) and increasing amounts of Cyclin D1 (0, 0.2, and 0.4 μg, as indicated). Cyclin D1 (0.4 μg)–transfected cells were treated with a CDK4 inhibitor (200 nM) for 4 h, and the effect of Cyclin D1–CDK4 on RUNX3–pRB-mediated p21 promoter (WT-p21-promoter-luciferase) activation was measured using a luciferase assay.
Because Cyclin D–CDK4/6 plays an essential role in pRB phosphorylation at early G1 phase, we examined the role of Cyclin D–CDK4/6 in dissociation of RUNX3–pRB. To this end, we treated HEK293 with a CDK4 inhibitor [1,4-Dimethoxy-9-thio(10H)-acridone] and examined the RUNX3–pRB interaction by IP and IB. The results revealed that inhibition of CDK4 prolonged the RUNX3–pRB interaction (Figure 5c). Similarly, increasing amounts of Cyclin D1 inhibited the cooperative transactivation activity of RUNX3 and pRB on the p21 promoter in a dose-dependent manner (Figure 5d). This inhibition by Cyclin D1 was rescued by treatment with a CDK4 inhibitor (Figure 5d). This result suggests that Cyclin D1/CDK4-mediated pRB phosphorylation facilitates the dissociation of the RUNX3–pRB complex.
pRB–RUNX3–BRD2 ternary complex forms early after serum stimulation
Previously, we showed that RUNX3 forms a complex with BRD2 1–2 h after serum stimulation and plays a key role in surveillance against oncogenic K-RAS.22 Hence, in this study we examined whether pRB, RUNX3, and BRD2 form a ternary complex to induce p21. For this purpose, Myc-RUNX3 or Myc-RUNX3-KR-94-171 [which does not interact with BRD222] were co-expressed with Flag-BRD2 and HA-RB in HEK293 cells. Twenty-four hours after transfection, the cells were serum-starved for 24 h, and then stimulated with serum. IP and IB analysis revealed that both RUNX3 and RUNX3-KR-94-171 interacted with pRB, 1–2 h after serum stimulation (Figure 6a, RX3/pRB). pRB and BRD2 also interacted at the same time points when RUNX3 was co-expressed (Figure 6a, pRB/BRD2). However, when RUNX3-KR-94-171 was expressed in place of wild-type RUNX3, pRB did not interact with BRD2, and p21 was not induced. These results suggest that pRB interacts with BRD2 through RUNX3 (Figure 6a, pRB/BRD2) 1–2 h after serum stimulation, and that the resultant pRB–RUNX3–BRD2 ternary complex contributes to serum-stimulated early induction of p21.
Mitogenic signals trigger the interaction between RUNX3 and pRB. (a) HEK293 cells were transfected with HA-RB, Flag-BRD2, and Myc-RX3 (or Myc-RX3-KR-94-171). The cells were serum-starved for 24 h and treated with 10% serum, and then the interactions between pRB, BRD2, and RUNX3 were measured by IP and IB at the indicated time points. (b) HEK293 cells transfected with either empty vector or Myc-K-RasG12V were serum-starved for 24 h and treated with 10% serum, and then the expression levels of endogenous pRB, RUNX3, and BRD2, as well as the interactions between the three endogenous proteins, were measured by IP and IB at the indicated time points. (c) HEK293 cells were transfected with HA-RB, Flag-BRD2, and Myc-RX3 (or Myc-RX3-KR-94-171) and Myc-K-RasG12V(or MEK1-CA). Cells were serum-starved for 24 h, and then treated with 10% serum for 8 h. Expression level of p21 and interactions between pRB, BRD2, and RUNX3 were measured by IP and IB.
Oncogenic K-RAS maintains the pRB–RUNX3–BRD2 complex
Analysis of the interactions of endogenous proteins revealed that, like the exogenously expressed proteins, endogenous pRB, RUNX3, and BRD2 also formed a ternary complex 1–2 h after serum stimulation (Figure 6b). Hence, we examined the effect of oncogenic K-Ras on formation of the pRB–RUNX3–BRD2 complex. Ectopic expression of K-RasG12V triggered interactions of endogenous pRB, RUNX3, and BRD2 in the absence of serum stimulation and maintained the resulting pRB–RUNX3–BRD2 complex for a longer period of time (Figure 6b). Consistent with this, the induced level of p21 expression was also maintained (Figure 6b). Similarly, expression of constitutively active MEK1 (MEK1-CA) as well as K-RasG12V increased pRB–RUNX3–BRD2 complex formation and p21 induction in a dose-dependent manner (Figure 6c). However, Myc-RUNX3-KR-94-171, which interacts with pRB but not BRD2, failed to induce p21 in response to expression of K-RasG12V or MEK1-CA (Figure 6c). These results suggest that a complex containing all three proteins (RUNX3, pRB, and BRD2) is required for the induction of p21, and that constitutive activation of the RAS-MEK1 pathway maintains the pRB–RUNX3–BRD2 complex and p21 expression. The series of molecular events involved in the association and dissociation of the pRB–RUNX3–BRD2 complex at the R-point are summarized in Figure 7.
Schematic illustration of R-point regulation by the pRB–RUNX3–BRD2 complex. Formation of the pRB–RUNX3–BRD2 complex is triggered by the RAS-MEK pathway 1 h after serum stimulation. The complex binds to the p21 promoter through RUNX-binding sites and induces p21 expression. Four hours after stimulation, the RAS-MEK pathway activity is downregulated and Cyclin D1 is induced. The induced Cyclin D1 forms a complex with CDK4/6 with the help of the induced p21. Cyclin D1–CDK4/6 dissociates pRB by pRB phosphorylation, and Cyclin D1–HDAC4 dissociates BRD2 by RUNX3 deacetylation from RUNX3.22 As a result, p21 expression is turned off. Subsequently, the decrease in p21 expression allows activation of Cyclin E–CDK2 activation, which drives cells to pass through the R-point. However, oncogenic K-RAS inhibits destruction of the pRB–RUNX3–BRD2 complex and prolongs p21 expression, which inhibits further cell-cycle progression. This series of molecular events may enable cell to distinguish normal mitogenic signals from abnormal oncogenic K-RAS signals, and thus make context-dependent R-point commitments.
Discussion
Runx3 constitutes a R-point–associated barrier against tumorigenesis
It is widely accepted that oncogene activation drives cellular transformation, whereas the p53 pathway plays major roles in preventing oncogene activation. However, in some types of adenomas, oncogene activation and p53 pathway inactivation occur after tumor initiation.2 In humans, for example, K-Ras activation is frequently found in lung adenocarcinomas, but rarely in lung adenomas. By contrast, RUNX3 is silenced in most human lung adenomas,25 and deletion of Runx3 in mouse lung induces lung adenomas.25 Therefore, Runx3 silencing is causally associated with the development of lung adenomas in the absence of oncogene activation. In this study, we demonstrated that the R-point is disturbed in Rx3−/− MEFs (p53-mutated), which develop into tumors in nude mice. Furthermore, we showed that re-expression of Runx3 restored the R-point and abolished the tumorigenicity of Rx3−/− MEFs. These observations suggest that Runx3 constitutes a p53-independent barrier against tumorigenesis, and that this barrier is associated with the R-point. Although the R-point is disrupted in nearly all cancer cells,1, 2 the mechanisms underlying R-point disruption remain poorly understood. Our results suggest that inactivation of RUNX3, which is frequently inactivated in multiple types of tumors,21 is associated with the deregulation of the R-point in tumor cells.
K-RasG12D knock-in mice develop lung cancer,26, 27 and loss of p53 does not have a significant impact on K-Ras–induced early lung tumorigenesis.28 Moreover, p53 affects only adenocarcinomas, and has no effect on adenomas developed in K-RasG12D knock-in mouse lung.29, 30 These results clearly demonstrate that the p53 pathway does not defend against endogenous oncogenic K-Ras.31 In this study, we found that Runx3 expression eliminated the tumorigenicity of Rx3−/−;K-Ras* MEFs. These results provide evidence for the existence of a Runx3-dependent barrier against endogenous oncogenic K-Ras. It is worth emphasizing that K-RAS is one of the most frequently mutated oncogenes in human cancers, and no effective treatment for patients carrying K-RAS mutations is currently available.32 RUNX3 is silenced in nearly all K-RAS–activated mouse and human lung adenocarcinomas, mainly by epigenetic alteration,22 and could therefore be reactivated, at least in theory.33 Thus, our results provide a theoretical basis for the development of a new therapeutic strategy against K-RAS–activated tumors.
Molecular crosstalk between differentiation and proliferation at the R-point
The R-point decision plays an essential role in cell fate determination. To achieve this goal in a context-dependent manner, the R-point decision-making machinery must be able to recognize the differentiation stage and integrity of the cell. The results of this study reveal the mechanism by which a cell recognizes its own status in regard to the R-point commitment. Cells enter the R-point via Cyclin D–CDK4/6-mediated pRB phosphorylation, and pass through the R-point via additional pRB phosphorylation mediated by Cyclin E–CDK2. Because p21 activates Cyclin D–CDK4/6 but inhibits Cyclin E–CDK2, the up- and downregulation of p21 expression must be involved in the control of entry into and passage through the R-point. We found that the pRB–RUNX3–BRD2 complex is formed 1 hour after serum stimulation, and that this complex induces p21 expression. When Cyclin D1 is induced (4 h after stimulation), the pRB–RUNX3–BRD2 complex is dissociated by Cyclin D1–CDK4/6. Subsequently, p21 expression is turned off, and the cell passes through the R-point (Figure 7). Our results reveal that RUNX3 (a lineage-determining transcription factor) and pRB (the primary regulatory of the R-point) form a complex and cooperate in regulating p21 (the CDK switch), demonstrating the existence of molecular crosstalk between differentiation and proliferation at the time of the R-point.
While RUNX3 is frequently inactivated in early stage cancers, a distinct body of work has shown that RUNX3 expression is acquired during the course of progression of some cancers. For example, RUNX3 suppresses proliferation and promotes metastasis in pancreatic ductal adenocarcinoma.34 Therefore, RUNX3 appears to function as both a tumor suppressor and a tumor promoter. Although it is unclear how RUNX3 performs opposite roles during tumorigenesis, it is possible that disruption of the R-point by deregulation of RUNX3 affects not only proliferation but also metastasis, since the R-point governs both proliferation and differentiation. Further study of the role of RUNX3 in R-point commitment may lead a fuller understanding of how RUNX3 plays dual roles in tumorigenesis.
Mammals have three RUNX family genes, RUNX1, RUNX2, and RUNX3. Their functions are linked to major developmental events and are intimately involved in tumorigenesis. RUNX1 and RUNX2 function as master regulators of hematopoiesis and osteogenesis, respectively.21, 35, 36 RUNX2 forms a complex with pRB, and the complex activates osteoblast-specific gene expression.37, 38 Because the R-point decision is dependent on differentiation stage, the pRB–RUNX2 complex might be associated with the R-point. If so, then RUNX1, RUNX2, and possibly other master regulators might also be involved in their own lineage-specific R-point commitment.
BRD2 recruits the mediator complex, SWI/SNF chromatin-remodeling complex, and RNA polymerase II, thereby contributes to transcriptional control.39 We previously reported that the RUNX3-BRD2 complex is formed 1–3 h after mitogenic stimulation and activates the p14ARF-p53 pathway.22 Because the time points of RUNX3-BRD2 complex formation overlapped with the R-point, we hypothesized that RUNX3 might be involved in the R-point. In this paper, we provide evidence that the RUNX3-BRD2 complex interacts with pRB and regulates the R-point. These observations suggest that the pRB–RUNX3–BRD2 complex provides a core for construction of a large complex that promotes the expression of target genes associated with R-point commitment. Identification of additional components of the pRB–RUNX3–BRD2 complex and the target genes of the complex will improve our understanding of the nature of R-point commitment.
Collectively, our results identify a series of molecular events that regulate R-point commitment and provide insight into the relationship between context-dependent R-point commitment and defense against tumorigenesis.
Experimental procedures
Mice
Runx3flox mice were kindly provided by Dr. I. Taniuchi.40 K-RasLSL-G12D knock-in mice were obtained from Jackson Labs (Sacramento, CA, USA). Adeno-Cre was purchased from Vector Biolabs (Philadelphia, PA, USA). Folxed Runx3 in MEFs was disrupted by infecting cells with 50 MOI Adeno-Cre. Animal studies were approved by the Institutional Animal Care Committee of Chungbuk National University.
Cell culture, treatment, siRNA, and transfection
The HEK293 cell line was obtained from the Korean Research Institute of Bioscience and Biotechnology (KRIBB, South Korea). MEFs of various genotypes were obtained from mouse embryos at 15.5 days of gestation. HEK293 and MEFs were maintained in Dulbecco’s modified Eagle’s medium (Gibco BRL, Thermo Fisher Scientific Korea) supplemented with 10% fetal bovine serum (Gibco BRL). CDK4 inhibitor [1,4-Dimethoxy-9-thio(10H)-acridone] was obtained from Calbiochem (CA, USA). Knockdown analysis was performed by transfecting HEK293 cells with 50 nM siRNA (si-RX3; 5′- AACCUGAUGCCAUAGACUC -3′ Bioneer, Korea, si-RB RNA; sc-29468, Santa Cruz Biotechnology) using RNAi MAX (Invitrogen, Thermo Fisher Scientific Korea). Full-length RUNX3 and its deletion and lysine mutants were described previously.22 Transient transfection was carried out using the Lipofectamine Plus reagent (Invitrogen).
Xenograft analysis
MEFs were suspended at 3 × 107 cells/ml in phosphate-buffered saline, and 0.2 ml of cell suspension was injected subcutaneously into each nude mouse. Five mice (two month old males) were used for each analysis.
RNA-seq analysis
The isolated total RNA was processed for preparing mRNA sequencing library using the Illumina TruSeq Stranded mRNA Sample Preparation kit (Illumina, CA, USA) according to the manufacturer's protocol. Quality and size of libraries were assessed using Agilent 2100 bioanalyzer DNA kit (Agilent, CA, USA). All libraries were quantified by qPCR using CFX96 Real Time System (Biorad, CA, USA) and sequenced on the NextSeq500 sequencers (Illumina, CA, USA). Potentially existing sequencing adapters and raw quality bases in the raw reads were trimmed by Cutadapt software.41 The cleaned high quality reads after trimming the low quality bases and sequencing adapters were mapped to the human reference genome hg19 of UCSC genome (https://genome.ucsc.edu) by STAR software.42 To quantify the mapped reads on the human reference genome in to the gene expression values, Cufflinks software.43 The differentially expressed genes between the two selected biological conditions were analyzed by Cuffdiff software in cufflinks package.43
Antibodies
Anti-Runx3 antibody (Ab40278) and anti-p53 antibody (Ab26) were obtained from Abcam (UK). Anti-phospho-p53 (#9284) was obtained from Cell Signaling Technology. Anti-Lamin antibody (#SC-20682) was obtained from Santa Cruz Biotechnology. Polyclonal anti-c-Myc-HRP (rabbit) (Sigma-Aldrich) and epitope-tag antibodies against Myc (9E10; Santa Cruz Biotechnology, Dallas, TX, USA), HA (12CA5; Roche Applied Science, Mannheim, Germany), and Flag (M2; Sigma, St. Louis, MO, USA) were purchased from the indicated vendors. Polyclonal anti-p27 and anti-p21 antibodies (rabbit) and monoclonal anti-cyclin D1 were purchased from Santa Cruz Biotechnology. Anti-phospho Erk (Thr-202, Tyr-204), anti-MEK1, and anti-phospho Akt (Ser-473) were purchased from Cell Signaling Technology (Danvers, MA, USA). Anti-pRB was purchased from BD Bioscience (CA, USA). Tubulin antibodies were obtained from Lab Frontier (Seoul, Korea).
IP and IB
Transfected cell lysates (300 μg) were incubated with monoclonal antibodies at 4 °C for 2 h or overnight, and then incubated with protein G–Sepharose beads (Amersham Pharmacia Biotech, UK) for 1 hour. After extensive washing, the immunoprecipitates were resolved on SDS-PAGE gels and transferred to a polyvinylidene difluoride membrane (Millipore, Billerica, MA, USA). The blot was visualized using LAS-3000 (Fujifilm, Japan) after treatment with ECL solution (Amersham Pharmacia Biotech, UK).
ChIP assays
ChIP assays were performed using a ChIP assay kit (Upstate cat # 17-295; CA, USA). HEK293 cells were treated with control or RUNX3-specific siRNA for 24 h. The cells were then cultured under serum starvation conditions for 24 h, and then treated with 10% serum. Cells were harvested at the indicated time points after serum stimulation. Chromatin was immunoprecipitated with the indicated antibodies. The p21 promoter region was amplified by PCR using the following primers: p21-5'(5′-CACCAGACTTCTCTGAGCCCCAG-3′) and p21-3'(5′-GCACTGTTAGAATGAGCCCCCTTTC-3′).
Methylation-specific PCR
Genomic DNA (5–10 μg) was treated with bisulfite, which converts unmethylated cytosine to uracil, using the EZ DNA Methylation Kit (Zymo Research, Irvine, CA, USA). The Runx3 promoter region was amplified by PCR using the following primers: unmethylated DNA–specific PCR primers, UnF5′- AGGTTTGGGGTATTAATTTT and UnR5′- CAACACATCCACCATCAA ; methylated DNA–specific PCR primers, MeF 5′- AGGTTCGGGGTATTAATTTC and MeR 5′- CAACACGTCCACCATCGA .
References
Malumbres M, Barbacid M . To cycle or not to cycle: a critical decision in cancer. Nat Rev Cancer 2001; 1: 222–231.
Weinberg R editor. The Biology of Cancer. Garland Science: Garland Science, 2007.
Pardee AB . A restriction point for control of normal animal cell proliferation. Proc Natl Acad Sci USA 1974; 71: 1286–1290.
Blagosklonny MV, Campisi J, Keyomarsi K, Medrano EE . No restriction points in life and science. Cell Cycle 2002; 1: 100–102.
Harper JW, Adami GR, Wei N, Keyomarsi K, Elledge SJ . The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell 1993; 75: 805–816.
LaBaer J, Garrett MD, Stevenson LF, Slingerland JM, Sandhu C, Chou HS et al. New functional activities for the p21 family of CDK inhibitors. Genes Dev 1997; 11: 847–862.
Cheng M, Olivier P, Diehl JA, Fero M, Roussel MF, Roberts JM et al. The p21(Cip1) and p27(Kip1) CDK 'inhibitors' are essential activators of cyclin D-dependent kinases in murine fibroblasts. EMBO J 1999; 18: 1571–1583.
Sherr CJ, Roberts JM . Living with or without cyclins and cyclin-dependent kinases. Genes Dev 2004; 18: 2699–2711.
Zetterberg A, Larsson O . Kinetic analysis of regulatory events in G1 leading to proliferation or quiescence of Swiss 3T3 cells. Proc Natl Acad Sci USA 1985; 82: 5365–5369.
Zetterberg A, Larsson O . Coordination between cell growth and cell cycle transit in animal cells. Cold Spring Harb Symp Quant Biol 1991; 56: 137–147.
Zetterberg A, Larsson O, Wiman KG . What is the restriction point? Curr Opin Cell Biol 1995; 7: 835–842.
Martinsson HS, Starborg M, Erlandsson F, Zetterberg A . Single cell analysis of G1 check points-the relationship between the restriction point and phosphorylation of pRb. Exp Cell Res 2005; 305: 383–391.
Michieli P, Chedid M, Lin D, Pierce JH, Mercer WE, Givol D . Induction of WAF1/CIP1 by a p53-independent pathway. Cancer Res 1994; 54: 3391–3395.
Deng C, Zhang P, Harper JW, Elledge SJ, Leder P . Mice lacking p21CIP1/WAF1 undergo normal development, but are defective in G1 checkpoint control. Cell 1995; 82: 675–684.
Odell A, Askham J, Whibley C, Hollstein M . How to become immortal: let MEFs count the ways. Aging (Albany NY). [Review] 2010; 2: 160–165.
Malumbres M, Barbacid M . RAS oncogenes: the first 30 years. Nat Rev Cancer 2003; 3: 459–465.
Serrano M, Lin AW, McCurrach ME, Beach D, Lowe SW . Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell 1997; 88: 593–602.
Palmero I, Pantoja C, Serrano M . p19ARF links the tumour suppressor p53 to Ras. Nature 1998; 395: 125–126.
Guerra C, Mijimolle N, Dhawahir A, Dubus P, Barradas M, Serrano M et al. Tumor induction by an endogenous K-ras oncogene is highly dependent on cellular context. Cancer Cell 2003; 4: 111–120.
Tuveson DA, Shaw AT, Willis NA, Silver DP, Jackson EL, Chang S et al. Endogenous oncogenic K-ras(G12D) stimulates proliferation and widespread neoplastic and developmental defects. Cancer Cell 2004; 5: 375–387.
Ito Y, Bae SC, Chuang LSH . The RUNX family: developmental regulators in cancer. Nature Reviews Cancer 2015; 15: 81–95.
Lee YS, Lee JW, Jang JW, Chi XZ, Kim JH, Li YH et al. Runx3 inactivation is a crucial early event in the development of lung adenocarcinoma. Cancer Cell 2013; 24: 603–616.
Herrera RE, Sah VP, Williams BO, Makela TP, Weinberg RA, Jacks T . Altered cell cycle kinetics, gene expression, and G1 restriction point regulation in Rb-deficient fibroblasts. Mol Cell Biol 1996; 16: 2402–2407.
Chi XZ, Yang JO, Lee KY, Ito K, Sakakura C, Li QL et al. RUNX3 suppresses gastric epithelial cell growth by inducing p21(WAF1/Cip1) expression in cooperation with transforming growth factor {beta}-activated SMAD. Mol Cell Biol 2005; 25: 8097–8107.
Lee KS, Lee YS, Lee JM, Ito K, Cinghu S, Kim JH et al. Runx3 is required for the differentiation of lung epithelial cells and suppression of lung cancer. Oncogene 2010; 29: 3349–3361.
Jackson EL, Willis N, Mercer K, Bronson RT, Crowley D, Montoya R et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev 2001; 15: 3243–3248.
Johnson L, Mercer K, Greenbaum D, Bronson RT, Crowley D, Tuveson DA et al. Somatic activation of the K-ras oncogene causes early onset lung cancer in mice. Nature 2001; 410: 1111–1116.
Muzumdar MD, Dorans KJ, Chung KM, Robbins R, Tammela T, Gocheva V et al. Clonal dynamics following p53 loss of heterozygosity in Kras-driven cancers. Nature communications 2016; 7: 12685.
Junttila MR, Karnezis AN, Garcia D, Madriles F, Kortlever RM, Rostker F et al. Selective activation of p53-mediated tumour suppression in high-grade tumours. Nature 2010; 468: 567–571.
Feldser DM, Kostova KK, Winslow MM, Taylor SE, Cashman C, Whittaker CA et al. Stage-specific sensitivity to p53 restoration during lung cancer progression. Nature 2010; 468: 572–575.
Berns A . Cancer: The blind spot of p53. Nature. [Comment News] 2010; 468: 519–520.
Drosten M, Barbacid M . Modeling K-Ras-driven lung adenocarcinoma in mice: preclinical validation of therapeutic targets. J Mol Med 2016; 94: 121–135.
Balmain A . Cancer: new-age tumour suppressors. Nature. [News] 2002; 417: 235–237.
Whittle MC, Izeradjene K, Rani PG, Feng L, Carlson MA, DelGiorno KE et al. RUNX3 Controls a Metastatic Switch in Pancreatic Ductal Adenocarcinoma. Cell 2015; 161: 1345–1360.
Ito Y . RUNX genes in development and cancer: regulation of viral gene expression and the discovery of RUNX family genes. Adv Cancer Res 2008; 99: 33–76.
Lund AH, van Lohuizen M . RUNX: a trilogy of cancer genes. Cancer Cell 2002; 1: 213–215.
Thomas DM, Carty SA, Piscopo DM, Lee JS, Wang WF, Forrester WC et al. The retinoblastoma protein acts as a transcriptional coactivator required for osteogenic differentiation. Mol Cell 2001; 8: 303–316.
Calo E, Quintero-Estades JA, Danielian PS, Nedelcu S, Berman SD, Lees JA . Rb regulates fate choice and lineage commitment in vivo. Nature 2010; 466: 1110–1114.
Denis GV, McComb ME, Faller DV, Sinha A, Romesser PB, Costello CE . Identification of transcription complexes that contain the double bromodomain protein Brd2 and chromatin remodeling machines. J Proteome Res 2006; 5: 502–511.
Naoe Y, Setoguchi R, Akiyama K, Muroi S, Kuroda M, Hatam F et al. Repression of interleukin-4 in T helper type 1 cells by Runx/Cbf beta binding to the Il4 silencer. J Exp Med 2007; 6 204: 1749–1755.
Martin M . Cutadapt removes adapter sequences from high-throughput sequencing reads. 2011. [next generation sequencing; small RNA; microRNA; adapter removal] 2011; 17: 1–17.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013; 29: 15–21.
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nature protocols 2012; 7: 562–578.
Acknowledgements
S-C Bae is supported by a Creative Research Grant (2014R1A3A2030690) through the National Research Foundation (NRF) of Korea. J-W Lee is supported by the Basic Science Research Program (2014R1A1A2007204) Y-S Lee is supported by Basic Science Research Program (2013R1A1A2057659) through the NRF funded by the Ministry of Education of Korea. IY Park was supported by “Cooperative Research Program for Agriculture Science and Technology Development (PJ01226101)” Rural Development Administration, Republic of Korea.
Author contributions
X-Z Chi contributed to the analysis of the tumorigenesity of the MEFs and the interaction between pRB and RUNX3. J-W Lee analyzed the interaction between BRD2 and RUNX3. Y-S Lee analyzed of Runx3 expression and methylation in MEFs. IY Park, Y Ito and S-C Bae interpreted the results and wrote the paper. All authors contributed to editing the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Chi, XZ., Lee, JW., Lee, YS. et al. Runx3 plays a critical role in restriction-point and defense against cellular transformation. Oncogene 36, 6884–6894 (2017). https://doi.org/10.1038/onc.2017.290
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.290
This article is cited by
-
RUNX3 methylation drives hypoxia-induced cell proliferation and antiapoptosis in early tumorigenesis
Cell Death & Differentiation (2021)
-
RUNX3 regulates cell cycle-dependent chromatin dynamics by functioning as a pioneer factor of the restriction-point
Nature Communications (2019)