Abstract
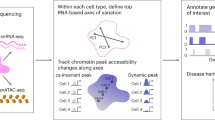
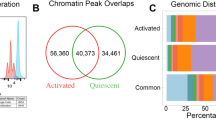
Cis-regulomes underlying immune-cell-specific genomic states have been extensively analyzed by structure-based chromatin profiling. By coupling such approaches with a high-throughput enhancer screen (self-transcribing active regulatory region sequencing (STARR–seq)), we assembled a functional cis-regulome for lipopolysaccharide-activated B cells. Functional enhancers, in contrast with accessible chromatin regions that lack enhancer activity, were enriched for enhancer RNAs (eRNAs) and preferentially interacted in vivo with B cell lineage-determining transcription factors. Interestingly, preferential combinatorial binding by these transcription factors was not associated with differential enrichment of their sites. Instead, active enhancers were resolved by principal component analysis (PCA) from all accessible regions by co-varying transcription factor motif scores involving a distinct set of signaling-induced transcription factors. High-resolution chromosome conformation capture (Hi-C) analysis revealed multiplex, activated enhancer–promoter configurations encompassing numerous multi-enhancer genes and multi-genic enhancers engaged in the control of divergent molecular pathways. Motif analysis of pathway-specific enhancers provides a catalog of diverse transcription factor codes for biological processes encompassing B cell activation, cycling and differentiation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data and accession
Data is submitted to GEO and can be accessed using the accession number GSE121753.
References
Singh, H., Medina, K. L. & Pongubala, J. M. Contingent gene regulatory networks and B cell fate specification. Proc. Natl Acad. Sci. USA 102, 4949–4953 (2005).
Cobaleda, C., Schebesta, A., Delogu, A. & Busslinger, M. Pax5: the guardian of B cell identity and function. Nat. Immunol. 8, 463–470 (2007).
Mansson, R. et al. Positive intergenic feedback circuitry, involving EBF1 and FOXO1, orchestrates B-cell fate. Proc. Natl Acad. Sci. USA 109, 21028–21033 (2012).
Grosschedl, R. Establishment and maintenance of B cell identity. Cold Spring Harb. Symp. Quant. Biol. 78, 23–30 (2013).
Kieffer-Kwon, K. R. et al. Interactome maps of mouse gene regulatory domains reveal basic principles of transcriptional regulation. Cell 155, 1507–1520 (2013).
Xu, H. et al. Regulation of bifurcating B cell trajectories by mutual antagonism between transcription factors IRF4 and IRF8. Nat. Immunol. 16, 1274–1281 (2015).
Yui, M. A. & Rothenberg, E. V. Developmental gene networks: a triathlon on the course to T cell identity. Nat. Rev. Immunol. 14, 529–545 (2014).
Li, P. & Leonard, W. J. Chromatin accessibility and interactions in the transcriptional regulation of T cells. Front. Immunol. 9, 2738 (2018).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Ho, J. W. et al. Comparative analysis of metazoan chromatin organization. Nature 512, 449–452 (2014).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Dailey, L. High throughput technologies for the functional discovery of mammalian enhancers: new approaches for understanding transcriptional regulatory network dynamics. Genomics 106, 151–158 (2015).
Tewhey, R. et al. Direct identification of hundreds of expression-modulating variants using a multiplexed reporter assay. Cell 165, 1519–1529 (2016).
Arnold, C. D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013).
Vockley, C. M. et al. Direct GR binding sites potentiate clusters of TF binding across the human genome. Cell 166, 1269–1281.e19 (2016).
Babbitt, C. C., Markstein, M. & Gray, J. M. Recent advances in functional assays of transcriptional enhancers. Genomics 106, 137–139 (2015).
Gaulton, K. J. et al. A map of open chromatin in human pancreatic islets. Nat. Genet. 42, 255–259 (2010).
Visel, A. et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009).
Ren, G. et al. CTCF-Mediated Enhancer-Promoter Interaction Is a Critical Regulator of Cell-to-Cell Variation of Gene Expression. Mol. Cell 67, 1049–1058.e6 (2017).
Nakahashi, H. et al. A genome-wide map of CTCF multivalency redefines the CTCF code. Cell Rep. 3, 1678–1689 (2013).
Palozola, K. C. et al. Mitotic transcription and waves of gene reactivation during mitotic exit. Science 358, 119–122 (2017).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Weirauch, M. T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Lara-Astiaso, D. et al. Immunogenetics. Chromatin state dynamics during blood formation. Science 345, 943–949 (2014).
Waters, L. R., Ahsan, F. M., Wolf, D. M., Shirihai, O. & Teitell, M. A. Initial B cell activation induces metabolic reprogramming and mitochondrial remodeling. iScience 5, 99–109 (2018).
Sen, R. & Grosschedl, R. Memories of lost enhancers. Genes Dev. 24, 973–979 (2010).
Levine, M., Cattoglio, C. & Tjian, R. Looping back to leap forward: transcription enters a new era. Cell 157, 13–25 (2014).
Maricque, B. B., Dougherty, J. D. & Cohen, B. A. A genome-integrated massively parallel reporter assay reveals DNA sequence determinants of cis-regulatory activity in neural cells. Nucleic Acids Res. 45, e16 (2017).
Zabidi, M. A. et al. Enhancer-core-promoter specificity separates developmental and housekeeping gene regulation. Nature 518, 556–559 (2015).
Muerdter, F. et al. Resolving systematic errors in widely used enhancer activity assays in human cells. Nat. Methods 15, 141–149 (2018).
Kieffer-Kwon, K. R. et al. Myc Regulates Chromatin Decompaction and Nuclear Architecture during B Cell Activation. Mol. Cell 67, 566–578 e10 (2017).
Johanson, T. M. et al. Transcription-factor-mediated supervision of global genome architecture maintains B cell identity. Nat. Immunol. 19, 1257–1264 (2018).
Zheng, Y. et al. Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature 463, 808–812 (2010).
Vaidyanathan, B. et al. The aryl hydrocarbon receptor controls cell-fate decisions in B cells. J. Exp. Med. 214, 197–208 (2017).
Fukaya, T., Lim, B. & Levine, M. Enhancer control of transcriptional bursting. Cell 166, 358–368 (2016).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Roy, A. L., Sen, R. & Roeder, R. G. Enhancer-promoter communication and transcriptional regulation of Igh. Trends Immunol. 32, 532–539 (2011).
Ansel, K. M., Lee, D. U. & Rao, A. An epigenetic view of helper T cell differentiation. Nat. Immunol. 4, 616–623 (2003).
Rochman, Y. et al. TSLP signaling in CD4+ T cells programs a pathogenic T helper 2 cell state. Sci. Signal 11, eaam8858 (2018).
Vanhille, L. et al. High-throughput and quantitative assessment of enhancer activity in mammals by CapStarr–seq. Nat. Commun. 6, 6905 (2015).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Thevenot, E. A., Roux, A., Xu, Y., Ezan, E. & Junot, C. Analysis of the human adult urinary metabolome variations with age, body mass index, and gender by implementing a comprehensive workflow for univariate and OPLS statistical analyses. J. Proteome Res. 14, 3322–3335 (2015).
Phanstiel, D. H., Boyle, A. P., Araya, C. L. & Snyder, M. P. Sushi.R: flexible, quantitative and integrative genomic visualizations for publication-quality multi-panel figures. Bioinformatics 30, 2808–2810 (2014).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
R Development Core Team. R: a language and environment for statistical computing. 3.3.0 edn. https://www.R-project.org (2018).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Lawrence, M., Gentleman, R. & Carey, V. rtracklayer: an R package for interfacing with genome browsers. Bioinformatics 25, 1841–1842 (2009).
Langdon, W. B. Performance of genetic programming optimised Bowtie2 on genome comparison and analytic testing (GCAT) benchmarks. BioData Min. 8, 1 (2015).
Acknowledgements
This study was supported by funds from the Cincinnati Children’s Research Foundation (CCRF) to H.S. We thank the core facilities of the Cincinnati Children’s Hospital Medical Center for assistance with mouse colony management, flow cytometry and high-throughput DNA sequencing. We thank M. Haas (CCHMC) for advice with FAIRE–seq and D. Fletcher (CCHMC) for technical support in sequencing of STARR–seq and eRNA libraries. The genomic analyses were made possible through use of the CCHMC BMI computing cluster and the Center for Research Computing at the University of Pittsburgh (PA, USA). We are particularly grateful to the following colleagues at CCHMC: L. Grimes, M.Weirauch, E. Miraldi and S. Pasare for useful scientific discussions and critical input on the manuscript.
Author information
Authors and Affiliations
Contributions
V.K.C. and H.S. conceptualized the study and designed the experiments. V.K.C. performed FAIRE–seq, STARR–seq, Hi-C, nascent RNA–seq and all computational genomic analyses. K.D-S. performed RNA–seq and Z.W. performed histone ChIP–seq. M.S. and V.K.C. pursued the transfection experiments with luciferase reporters in CH12 cells. V.K.C. and H.S. interpreted the results and wrote the manuscript. H.S. supervised the project and is responsible for funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Reproducibility of RNA-seq, chromatin profiling and STARR-seq in LPS activated B-cells.
Analyzed data is from biological replicates. a, RNA-seq data is plotted for all known genes as log transform of FPKM values. b, FAIRE-seq peaks are plotted as quantile normalized tag counts for 55,133 regions c,d. H3K27ac peaks (n=23,938) and H3K4me3 peaks (n=16,871) plotted for quantile normalized tag counts, respectively. e,f. A pool of size-selected FAIRE-DNA fragments were used to construct two libraries, each of which was transfected into LPS activated splenic B cells at either 48h or 60h. RNA was isolated at 72h post-activation for sequencing. Quantile normalized tag counts of STARR-seq output for the 55,130 FAIRE-seq peaks are plotted. In all panels, a value of 1 was added to the counts before transforming to log2. Indicated PCC values are Pearson correlation coefficients for biological replicates. g, Levels of nascent transcripts from genes whose promoters are situated within 2 Mb intervals around STARR-pos or STARR-neg regions used in luciferase assay. Indicated p-value is based on a two-sided t-test involving genes in STARR-pos (n=762) and STARR-neg (n=377) sets.
Extended Data Fig. 2 Combinatorial binding of B cell determining TFs across genome.
a, Combinatorial binding of B-cell TFs was analyzed using the ChIP-seq peaks for the 6 TFs, Ebf1, Pax5, E2A, PU.1, Blimp-1 and Ets1. Peaks were binned based on how many represented singular binding events versus those that evidenced binding by 2 or more of the six TFs. b, Assessment of TF ChIP-seq bins analyzed in panel a for the co-occurrence of TF binding motifs of the same TFs. Motif co-occurrence frequency (horizontal axis) across each TF ChIP-seq bin (vertical axis) is plotted as a heatmap. c, FAIRE-seq peak tag counts for each of the bins analyzed. FAIRE-seq peak counts in each bin were as follows; bin1=10,500, bin2=7,224, bin3=5,997, bin4=5,602, bin5=3,795 and bin6=1,014. Boxplots display median values (notches) along with 95% confidence intervals. d, STARR-seq peak tag counts for each of the indicated ChIP-seq bins. STARR-seq peak counts in each bin were as follows; bin1=1,615, bin2=1,596, bin3=1,899, bin4=2,317, bin5=1,952 and bin6=628. e. STARR-positive and STARR-negative regions were analyzed for TF binding peaks for indicated B lineage determining TFs and scanned for their motifs, shown in Fig. 2a. The ratios of fractions of STARR-pos (n=11,809) and STARR-neg (11,942) sequences overlapping the TF binding peaks (violet bars) or containing the indicated TF motifs (turquoise bars) are plotted. Paired two-sided t-test comparing TF binding versus TF motif occurrence, P=0.0009. f. The regions in panel S3A were further analyzed for motif combinations. Percentages of STARR-pos or STARR-neg sequences with 1-6 TF motifs are plotted. Boxplots in panels c, d display median values along with the inter-quartile distributions.
Extended Data Fig. 3 PC1 and PC2 loadings of FAIRE-seq regions using mscore matrix.
Key TF motifs that contribute substantially to the variance explained by PC1 and PC2 are displayed on the plot. Highlighted TF motifs with their mscore distributions in the STARR-pos (n=11,809) versus STARR-neg (11,942) regions are displayed in boxplots with indicated p-values for two-sided Welch 2-sample test. Boxplots for selected motifs display median values of mscores along with the inter-quartile distributions.
Extended Data Fig. 4 Hi-C analysis and connectivity of functional B cell cis-regulome.
Hi-C replicates of LPS activated B cells (72h). Biologically independent experiments were analyzed for reproducibility by bin-wise genome wide correlation for the indicated bin-sizes at ~300 million read depths for each replicate. The Homer function getHiCcorrDiff.pl was used to generate correlation matrices. Correlation coefficients of each bin at a given resolution are plotted. Violin plots show kernel probability density of the data at the indicated levels of Hi-C resolution. Number of genomic bins at a given level of resolution were as follows; 10kb n= 258,640, 20kb n= 129,308, 5kb n=295,287 and 50kb n=51,678.
Extended Data Fig. 5 Histone profiles of multi-genic enhancers.
Genome browser plots of H3K27ac and H3K4me3 peaks at indicated multi-genic enhancers in resting (rB) and activated (aB) B cells. P10 and P11 refer to the numbers of interacting promoters for indicated enhancer.
Extended Data Fig. 6 Genes and TF motif enrichment analysis in activated B cells.
a, KEGG pathway enrichment using GSEA with RNA-seq data from resting (rB) and LPS activated B cells (aB), n=2 biological replicates. Enrichment plots for indicated KEGG terms are displayed b, TF motif enrichment analysis (HOMER) was performed using STARR-seq tags for peaks that were uniquely present in the 48h (n=1,851) or 60h (n=2,869) transfection datasets. TF motif enrichment values (log2 odds ratios) were determined for each transfection time point using the shared peaks for both time points as the background set. In the plotted data, TF motifs that were enriched with logP values < -20 (HOMER output) are indicated as colored circles. c, TF motif enrichment analysis (HOMER) was performed using STARR-seq tags for peaks that were uniquely present in the 48h transfection dataset using the 60h dataset as background or vice versa (see panel b). TFs whose motifs (simple or composite elements) are highly enriched (logP < -20) are displayed in the bar plot with their log2 odds ratios.
Extended Data Fig. 7 PCA-DA analysis of monogenic PC and GC enhancers.
PC1 and PC2 loadings for all TF motifs constituting the mscore matrix are displayed corresponding to the PCA plot in Fig. 5d. Motif names are displayed instead of dots. Where a single motif represents the binding site of 2 or more related TFs, TF names are separated by “:” to indicate all TFs that recognize that motif. The dashed line delineates TF motifs whose mscore variance distinguishes the PC (n=247) and GC (n=228) enhancers.
Supplementary information
Supplementary Tables
The tables are an extensive set of genomic resource information that was generated by the experimental and computational analyses in this study
Rights and permissions
About this article
Cite this article
Chaudhri, V.K., Dienger-Stambaugh, K., Wu, Z. et al. Charting the cis-regulome of activated B cells by coupling structural and functional genomics. Nat Immunol 21, 210–220 (2020). https://doi.org/10.1038/s41590-019-0565-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0565-0
This article is cited by
-
Cis-regulatory atlas of primary human CD4+ T cells
BMC Genomics (2023)
-
Three-dimensional genome organization in immune cell fate and function
Nature Reviews Immunology (2023)
-
Pre-mitotic genome re-organisation bookends the B cell differentiation process
Nature Communications (2021)
-
Gene regulatory networks STARR-ing B cells
Nature Immunology (2020)