Abstract
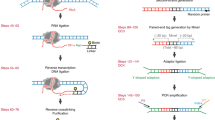
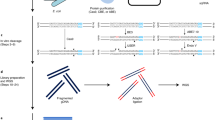
R-loops are prevalent three-stranded non-B DNA structures composed of an RNA–DNA hybrid and a single strand of DNA. R-loops are implicated in various basic nuclear processes, such as class-switch recombination, transcription termination and chromatin patterning. Perturbations in R-loop metabolism have been linked to genomic instability and have been implicated in human disorders, including cancer. As a consequence, the accurate mapping of these structures has been of increasing interest in recent years. Here, we describe two related immunoprecipitation-based methods for mapping R-loop structures: basic DRIP-seq (DNA–RNA immunoprecipitation followed by high-throughput DNA sequencing), an easy, robust, but resolution-limited technique; and DRIPc-seq (DNA–RNA immunoprecipitation followed by cDNA conversion coupled to high-throughput sequencing), a high-resolution and strand-specific iteration of the method that permits accurate R-loop mapping genome wide. Briefly, after gentle DNA extraction and restriction digestion with a cocktail of enzymes, R-loop structures are immunoprecipitated with the anti-RNA–DNA hybrid S9.6 antibody. Compared with DRIP-seq, in which the immunoprecipitated DNA is directly sequenced, DRIPc-seq permits the recovery of the RNA moiety of R-loops, and these RNA strands are subjected to strand-specific RNA sequencing (RNA-seq) analysis. DRIPc-seq can be performed in 5 d and can be applied to any cell type, provided sufficient starting material can be collected. Accurately mapping R-loop distribution in various cell lines and under varied conditions is essential to understanding the formation, roles and dynamic resolution of these important structures.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Santos-Pereira, J. M. & Aguilera, A. R loops: new modulators of genome dynamics and function. Nat. Rev. Genet. 6, 583–597 (2015).
Kreuzer, K. N. & Brister, J. R. Initiation of bacteriophage T4 DNA replication and replication fork dynamics: a review in the Virology Journal series on bacteriophage T4 and its relatives. Virol. J. 7, 358 (2010).
Carles-Kinch, K. & Kreuzer, K. N. RNA-DNA hybrid formation at a bacteriophage T4 replication origin. J. Mol. Biol. 266, 915–926 (1997).
Masukata, H. & Tomizawa, J. A mechanism of formation of a persistent hybrid between elongating RNA and template DNA. Cell 62, 331–338 (1990).
Itoh, T. & Tomizawa, J. Formation of an RNA primer for initiation of replication of ColE1 DNA by ribonuclease H. Proc. Natl. Acad. Sci USA 77, 2450–2454 (1980).
Akman, G. et al. Pathological ribonuclease H1 causes R-loop depletion and aberrant DNA segregation in mitochondria. Proc. Natl. Acad. Sci USA 113, E4276–E4285 (2016).
Lee, D. Y. & Clayton, D. A. Initiation of mitochondrial DNA replication by transcription and R-loop processing. J. Biol. Chem. 273, 30614–30621 (1998).
Xu, B. & Clayton, D. A. A persistent RNA-DNA hybrid is formed during transcription at a phylogenetically conserved mitochondrial DNA sequence. Mol. Cell. Biol. 15, 580–589 (1995).
Daniels, G. A. & Lieber, M. R. RNA:DNA complex formation upon transcription of immunoglobulin switch regions: implications for the mechanism and regulation of class switch recombination. Nucleic Acids Res. 23, 5006–5011 (1995).
Huang, F. T., Yu, K., Hsieh, C. L. & Lieber, M. R. Downstream boundary of chromosomal R-loops at murine switch regions: implications for the mechanism of class switch recombination. Proc. Natl. Acad. Sci. USA 103, 5030–5035 (2006).
Reaban, M. E. & Griffin, J. A. Induction of RNA-stabilized DNA conformers by transcription of an immunoglobulin switch region. Nature 348, 342–344 (1990).
Yu, K., Chedin, F., Hsieh, C. L., Wilson, T. E. & Lieber, M. R. R-loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells. Nat. Immunol. 4, 442–451 (2003).
Ratmeyer, L., Vinayak, R., Zhong, Y. Y., Zon, G. & Wilson, W. D. Sequence specific thermodynamic and structural properties for DNA.RNA duplexes. Biochemistry 33, 5298–5304 (1994).
Roberts, R. W. & Crothers, D. M. Stability and properties of double and triple helices: dramatic effects of RNA or DNA backbone composition. Science 258, 1463–1466 (1992).
Ginno, P. A., Lott, P. L., Christensen, H. C., Korf, I. & Chedin, F. R-loop formation is a distinctive characteristic of unmethylated human CpG island promoters. Mol. Cell 45, 814–825 (2012).
Hartono, S. R., Korf, I. F. & Chedin, F. GC skew is a conserved property of unmethylated CpG island promoters across vertebrates. Nucleic Acids Res. 43, 9729–9741 (2015).
Aguilera, A. & Garcia-Muse, T. R loops: from transcription byproducts to threats to genome stability. Mol. Cell 46, 115–124 (2012).
Sollier, J. & Cimprich, K. A. Breaking bad: R-loops and genome integrity. Trends Cell Biol. 25, 514–522 (2015).
Costantino, L. & Koshland, D. The Yin and Yang of R-loop biology. Curr. Opin. Cell Biol. 34, 39–45 (2015).
Skourti-Stathaki, K. & Proudfoot, N. J. A double-edged sword: R loops as threats to genome integrity and powerful regulators of gene expression. Genes Dev. 28, 1384–1396 (2014).
Richard, P. & Manley, J. L. R loops and links to human disease. J. Mol. Biol. 429, 3168–3180 (2017).
Phillips, D. D. et al. The sub-nanomolar binding of DNA-RNA hybrids by the single-chain Fv fragment of antibody S9.6. J. Mol. Recognit. 26, 376–381 (2013).
Sanz, L. A. et al. Prevalent, dynamic, and conserved R-loop structures associate with specific epigenomic signatures in mammals. Mol. Cell 63, 167–178 (2016).
Skourti-Stathaki, K., Proudfoot, N. J. & Gromak, N. Human senataxin resolves RNA/DNA hybrids formed at transcriptional pause sites to promote Xrn2-dependent termination. Mol. Cell 42, 794–805 (2011).
El Hage, A. & Tollervey, D. Immunoprecipitation of RNA:DNA hybrids from budding yeast. Methods Mol. Biol. 1703, 109–129 (2018).
Garcia-Rubio, M., Barroso, S. I. & Aguilera, A. Detection of DNA-RNA hybrids in vivo. Methods Mol. Biol. 1672, 347–361 (2018).
Stork, C. T. et al. Co-transcriptional R-loops are the main cause of estrogen-induced DNA damage. eLlife 5, e17548 (2016).
Manzo, S. G. et al. DNA Topoisomerase I differentially modulates R-loops across the human genome. Genome Biol. 19, 100 (2018).
Hartono, S. R. et al. The affinity of the S9.6 antibody for double-stranded RNAs impacts the accurate mapping of R-loops in fission yeast. J. Mol. Biol. 430, 272–284 (2018).
Halasz, L. et al. RNA-DNA hybrid (R-loop) immunoprecipitation mapping: an analytical workflow to evaluate inherent biases. Genome Res. 27, 1063–1073 (2017).
Ginno, P. A., Lim, Y. W., Lott, P. L., Korf, I. & Chedin, F. GC skew at the 5′ and 3′ ends of human genes links R-loop formation to epigenetic regulation and transcription termination. Genome Res.. 23, 1590–1600 (2013).
Konig, F., Schubert, T. & Langst, G. The monoclonal S9.6 antibody exhibits highly variable binding affinities towards different R-loop sequences. PLoS ONE 12, e0178875 (2017).
Cerritelli, S. M. & Crouch, R. J. Ribonuclease H: the enzymes in eukaryotes. FEBS J. 276, 1494–1505 (2009).
Vanoosthuyse, V. Strengths and weaknesses of the current strategies to map and characterize R-loops. Noncoding RNA 4, 9 (2018).
Zhang, Z. Z., Pannunzio, N. R., Hsieh, C. L., Yu, K. & Lieber, M. R. Complexities due to single-stranded RNA during antibody detection of genomic rna:dna hybrids. BMC Res. Notes 8, 127 (2015).
Ausubel, F. et al. Curr. Protoc. Mol. Biol. Suppl. 8, p. 3.13.1 (1995).
Thomas, M., White, R. L. & Davis, R. W. Hybridization of RNA to double-stranded DNA: formation of R-loops. Proc. Natl. Acad. Sci. USA 73, 2294–2498 (1976).
White, R. L. & Hogness, D. S. R loop mapping of the 18S and 28S sequences in the long and short repeating units of Drosophila melanogaster rDNA. Cell 10, 177–192 (1977).
Drolet, M., Bi, X. & Liu, L. F. Hypernegative supercoiling of the DNA template during transcription elongation in vitro. J. Biol. Chem. 269, 2068–2074 (1994).
Duquette, M. L., Handa, P., Vincent, J. A., Taylor, A. F. & Maizels, N. Intracellular transcription of G-rich DNAs induces formation of G-loops, novel structures containing G4 DNA. Genes Dev. 18, 1618–1629 (2004).
Wahba, L., Costantino, L., Tan, F. J., Zimmer, A. & Koshland, D. S1-DRIP-seq identifies high expression and polyA tracts as major contributors to R-loop formation. Genes Dev. 30, 1327–1338 (2016).
Xu, W. et al. The R-loop is a common chromatin feature of the Arabidopsis genome. Nat. Plants 3, 704–714 (2017).
Dumelie, J. G. & Jaffrey, S. R. Defining the location of promoter-associated R-loops at near-nucleotide resolution using bisDRIP-seq. Elife 6, e28306 (2017).
Chen, L. et al. R-ChIP using inactive RNase H reveals dynamic coupling of R-loops with transcriptional pausing at gene promoters. Mol. Cell 68, 745–757.e5 (2017).
Loomis, E. W., Sanz, L. A., Chedin, F. & Hagerman, P. J. Transcription-associated R-loop formation across the human FMR1 CGG-repeat region. PLoS Genet. 10, e1004294 (2014).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Lis, J. T. & Schleif, R. Size fractionation of double-stranded DNA by precipitation with polyethylene glycol. Nucleic Acids Res. 2, 383–389 (1975).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Xu, S., Grullon, S., Ge, K. & Peng, W. Spatial clustering for identification of ChIP-enriched regions (SICER) to map regions of histone methylation patterns in embryonic stem cells. Methods Mol. Biol. 1150, 97–111 (2014).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Rabani, M. et al. Metabolic labeling of RNA uncovers principles of RNA production and degradation dynamics in mammalian cells. Nat. Biotechnol. 29, 436–442 (2011).
Aronesty, E. ea-utils: Command-line tools for processing biological sequencing data. https://expressionanalysis.github.io/ea-utils/ (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Acknowledgements
We thank S.R. Hartono, J. Smolka and M. Malig for constructive comments on the manuscript. This work was supported by a grant from the National Institutes of Health (GM120607).
Author information
Authors and Affiliations
Contributions
F.C. conceived the study. L.A.S. performed the experiments. F.C. and L.A.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information: Nature Protocols thanks Francesca Storici and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Sanz, L. A. et al. Mol. Cell 63, 167–178 (2016): https://doi.org/10.1016/j.molcel.2016.05.032
Manzo, S. G. et al. Genome Biol. 19, 100 (2018): https://doi.org/10.1186/s13059-018-1478-1
Hartono, S. R. et al. J. Mol. Biol. 430, 272–284 (2018): https://doi.org/10.1016/j.jmb.2017.12.016
Integrated supplementary information
Supplementary Figure 1. Sequences captured by DRIPc-seq show no correlation with S9.6 intrinsic binding preferences.
6-mers found to be poorly or tightly bound by S9.6 were curated from Konig et al. (2017) and grouped as low and high binding. We evaluated each 6-mer frequency in the R-loop forming sequence space identified by DRIPc-seq (Sanz et al., 2016), resulting in observed frequencies. As a comparison, we retrieved non-R-loop forming genic regions derived from loci that were matched for expression, length and location and measured6-mer frequencies over this control set. For each R-loop peak, 25 random, matched peaks were extracted and the average frequency determined for each 6-mer. This resulted in expected frequencies. A. The graph shows the log2 fold ratio of observed (R-loop forming) over expected (matched non-R-loop forming) frequencies for each 6-mer.Some 6-mers are clearly more or less represented than others in DRIPc-seq data compared to expectations from control non-R-loop loci. This could reflect the intrinsic sequence preference of R-loop formation and/or the intrinsic preference of S9.6 antibody. If the latter is true, we expected S9.6-highly bound epitopes (red) to be over-represented and S9.6-poorly bound epitopes (blue) to be under-represented. This was not observed, however. Instead, S9.6 tightly or poorly bound 6-mers were equally likely to be under- or over-represented. This suggests that DRIPc-seq data does not suffer from systematic biases caused by S9.6 sequence preference. B. To account for what could be driving the over- or under-representation of certain 6-mers, we simply calculated the GA content of the motifs. As shown, depleted motifs tend to be GA-poor (CT-rich), while enriched motifs tend to be GA-rich irrespective of whether they are tightly or poorly bound by S9.6 (the dashed grey line represents 50% GA content). Given that GA-rich regions are favorable for R-loop formation, the observed trends are most likely to reflect the intrinsic sequence biases underlying R-loop formation, not S9.6 binding. Similar results were observed when 8-mers were considered.
Supplementary Figure 2 Genomic DNA digestion profiles.
DNA digestion profiles after Step 10 were visualized after agarose gel electrophoresis through a 0.8% agarose gel run in 1x TAE buffer. DNA was extracted from human NTERA-2 cells and digested with restriction enzyme cocktail indicated in Step 10. Lanes 1 and 2 show an example of incomplete digestion, as evidenced by the high molecular weight bands above 20 kilobases. Lanes 3 and 4 show an example of fully digested DNA as judged from the disappearance of the top band. The leftmost lane (M) corresponds to a1kb plus GeneRuler ladder from Thermo Fisher.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2
Rights and permissions
About this article
Cite this article
Sanz, L.A., Chédin, F. High-resolution, strand-specific R-loop mapping via S9.6-based DNA–RNA immunoprecipitation and high-throughput sequencing. Nat Protoc 14, 1734–1755 (2019). https://doi.org/10.1038/s41596-019-0159-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0159-1
This article is cited by
-
Aberrant R-loop–mediated immune evasion, cellular communication, and metabolic reprogramming affect cancer progression: a single-cell analysis
Molecular Cancer (2024)
-
Looping out of control: R-loops in transcription-replication conflict
Chromosoma (2024)
-
Accelerated DNA replication fork speed due to loss of R-loops in myelodysplastic syndromes with SF3B1 mutation
Nature Communications (2024)
-
Integrative modeling of lncRNA-chromatin interaction maps reveals diverse mechanisms of nuclear retention
BMC Genomics (2023)
-
NKAP acts with HDAC3 to prevent R-loop associated genome instability
Cell Death & Differentiation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.