Abstract
Published transcription data from a set of 19 diploid Arabidopsis thaliana and 5 tetraploid (3 allo- and 2 auto- tetraploid) Arabidopsis accessions were re-analysed to identify reliable reference genes for normalization purposes. Five conventional and 16 novel reference genes previously derived from microarray data covering a wide range of abundance in absolute expression levels in diploid A. thaliana Col-0 were employed. Transcript abundance was well conserved for all 21 potential reference genes in the diploid A. thaliana accessions, with geNorm and NormFinder analysis indicating that AT5G46630, AT1G13320, AT4G26410, AT5G60390 and AT5G08290 were the most stable. However, conservation was less good among the tetraploid accessions, with the transcription of seven of the 21 genes being undetectable in all allotetraploids. The most stable gene was AT5G46630, while AT1G13440 was the unstable one. Hence, the choice of reference gene(s) for A. thaliana is quite wide, but with respect to the analysis of transcriptomic data derived from the tetraploids, it is probably necessary to select more than one reference gene.
Similar content being viewed by others
Introduction
The quantification of transcript abundance underpins much of current plant molecular biology1, which is usually quantified via real time PCR analysis2. The technique relies heavily on a normalization step based on the transcript abundance of a so-called “reference” gene (or genes), which is expected a priori to be stably transcribed2,3. The transcription level of an ideal reference gene is temporally and spatially constant and is unaffected by external factors3,4. These conditions are seldom fulfilled, even by the most frequently used reference sequences, such as those encoding 18S rRNA, GAPDH, EF-1α, ubiquitin, actin, α-tubulin or phosphatase 2A. Various software packages, prominently geNorm5 and NormFinder6, have been developed to identify reference genes appropriate to particular experimental systems.
The diploid angiosperm species Arabidopsis thaliana has long served as the leading genomic model for dicotyledonous plants7,8. The volume of transcriptomic data generated in this species is very large, especially given the development of microarray and next generation sequencing technologies. Such data sets have helped to identify potential replacements for the reference genes conventionally used for the analysis of transcription in A. thaliana3.
Like most plant genera, Arabidopsis includes a number of both diploid and polyploid species. As yet, the appropriateness of key reference genes, chosen on the basis of their transcription behavior in A. thaliana, especially in those with different ploidy levels, has yet to be tested. A recent comparison has been published between the transcriptomes of 19 diploid A. thaliana accessions9, chosen to maximize the range of intraspecific phenotypic variation, along with some tetraploid Arabidopsis spp. entries (A. thaliana 1001 Genomes Project)10. Here, we have exploited these data to identify an appropriate set of reference genes for normalization of transcriptomic data derived from both A. thaliana and some tetraploid Arabidopsis spp.
Results
Reference gene transcription among A. thaliana accessions
We download 38 RNA sequencing data from MAGIC (Multiparent Advanced Generation Inter-Cross) founder accessions of 19 A. thaliana diploids and all 11 available RNA sequencing data from 5 Arabidopsis tetraploids (3 allo- and 2 auto- tetraploid) from Gene Expression Omnibus (GEO) database. A set of five conventional (GAPDH, ACT2, UBQ10, UBC, EF-1α) and 16 potentially informative novel genes previously derived from microarray data in diploid A. thaliana Col-0 was assembled, the transcription of each recorded over at least 80% entries were selected (Table 1)3. The transcript abundance of these genes was measured by RPKM (reads per kilobase of exon model per million mapped reads). Among the 21 genes assessed in the 19 A. thaliana accessions, the four most highly abundant ones were EF-1α (AT5G60390, RPKM: 901.2 ± 18.4, mean ± SD, the same below), GAPDH (AT1G13440, 876.6 ± 24.2), UBC10 (AT4G05320, 242.7 ± 7.4) and ACT2 (AT3G18780, 210.3 ± 5.4). Transcript abundance of 11 out of the 21 genes was only moderate (RKPM ranging from 14.8 ± 0.4 to 97.9 ± 2.4) and that of the remaining six genes was relatively low (RKPM ranging from 0.84 ± 0.05 to 8.19 ± 0.19) (Fig. 1 and Supplementary Data S1, Online Resource). Inter-accession variation in RPKM was typically limited, with a small number of the accessions being largely responsible for the variation present.
Stability was assessed based on geNorm and NormFinder software. The former associates a stability value (M) with each potential reference gene, where a low M reflects stability and a high M instability5. An M value of < 0.5 is conventionally accepted for a reference, but with high M value (≥0.5) should be avoided11. NormFinder ranks genes according to the similarity of their transcript abundances, applying a model-based approach6. The geNorm analysis showed that the M value of 19 out of the 21 candidate genes was < 0.5, suggesting that any of one (or more) of them was appropriate as a reference sequence(s). The most stable sequences were AT5G46630 (encoding a clathrin adaptor complex subunit), AT1G13320 (PP2A subunit), AT4G26410 (uncharacterised conserved protein), AT5G60390 (EF-1α) and AT5G08290 (mitosis protein YLS8) (Fig. 2a). NormFinder analysis resulted in a similar set of stable genes: AT1G13320 and AT5G46630 emerged as the most stable (Table 2). Nevertheless, even these reference genes could be unstable in two special samples, for example, the RKPM of AT1G13320 was 1.3 times of minimum between diploid WU and TSU samples, while that of AT4G34270, which was identified by both geNorm and NormFinder as being a relatively unstable sequence had the closest transcript abundance (RKPM(WU/TSU) = 14.59/14.06). The pairwise variation (V) metric generated by geNorm was informative with respect to determining the optimal number of reference genes necessary for accurate normalization. A Vn/Vn + 1 value of < 0.15 indicates that an additional reference gene is not required5,12,13. Across the 19 diploid accessions, this metric was below the threshold for the inclusion of a third reference sequence for each initial pair chosen (Fig. 2b).
Average expression stability values (M) and the pairwise variation (V) metric calculated by geNorm.
(a) M values of the 21 reference genes among 19 diploid A. thaliana accessions. A low value of M indicates greater stability. The seven columns shown in black refer to genes not transcribed in at least one tetraploid accession. (b) The pairwise variation (V) metric calculated to indicate the optimal number of reference genes required for normalization. A decrease in V indicates that the addition of the gene in question should improve normalization accuracy.
Differential stability in A. thaliana and tetraploid Arabidopsis
Although the reference genes behaved very consistently among the A. thaliana accessions, this was not the case in the tetraploids. Of the 21 reference genes, seven genes (AT1G62930, AT5G55840, AT4G38070, AT5G15710, AT3G53090, AT2G32170 and AT5G12240) of low RPKM in diploid were not detectable in all allotetraploids (Fig. 1, Supplementary Data S1), so that none of these would be suitable for normalization at the tetraploid level. A statistical analysis suggested among the remaining 14 reference genes, six genes (AT5G46630, AT4G33380, AT4G26410, AT4G05320, AT3G01150 and AT1G13320) showed a significant difference of transcript abundance between diploid and tetraploid Arabidopsis accessions according to Tukey's test and the Student's t test (P < 0.05) (Fig. 3a), meanwhile the coefficient of variation (Cv) analysis suggested that the Cv of the rest 14 genes in tetraploids was more variable than among the diploid A. thaliana accessions and the maximum Cv value of each autotetraploid (MaxA. t = 41.0%; MaxA. a = 45.3%) was less than those in the allotetraploid ones (MaxA. s = 77.2%; MaxA. t× A. a F1 = 48.3%; MaxA. t× A. a F8 = 50.9%) (Fig. 3b). Hence, it's necessary to provide a suitable list of reference gene(s) for tetraploid Arabidopsis accessions separately. According to geNorm, ten out of the 14 genes produced a M value of < 0.5 (Fig. 4), of which the most stable gene across all tetraploid accessions was AT5G46630, followed by AT4G26410, AT3G18780, AT3G01150 and AT1G13320 (Fig. 4). A similar stability ranking of 14 tested genes was observed based on analysis using NormFinder software (Table 3), i.e, the most stable gene was also AT5G46630, while AT1G13440 was the unstable one. The stability among the tetraploid accessions of the best-performing genes according to both software packages, was sufficient to allow them to be used as reference in different tetraploid Arabidopsis accessions.
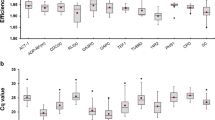
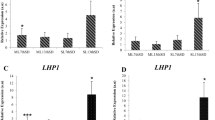
The comparison of transcript abundances and the coefficient of variation (Cv) of each reference gene between A. thaliana and tetraploid Arabidopsis accessions.
(a) The comparison of transcript abundances of 14 reference genes. Boxes indicate the 25th/75th percentiles, the line represents the median, squares represent the means and whiskers (plus/minus values) indicate the ranges of total samples. (b) The coefficient of variation (Cv). Cv values close to 0% indicate little deviation from the mean RPKM.
Discussion
The feasibility of reference genes identification from RNA-seq
Reliable reference gene(s) are an important resource for the analysis of transcriptomic data. Microarray-based transcriptomic data are derived from the hybridization between cDNA and the probe set attached to the chip, which are subjected to a degree of noise due to cross-hybridization and uneven hybridization efficiency14,15. In contrast, high throughput sequencing platforms such as RNA-seq generate a direct count of the number of transcript copies16 and this parameter is free of the hybridization artifacts which compromise the quality of microarray-based data17,18.
Reference gene stability in diploid A. thaliana
Based on microarray-acquired transcriptomic data, a set of 25 stably transcribed genes has been described for diploid A. thaliana Col-03,19. Of these, four were not well represented in the transcription profiles of the 19 diploid A. thaliana accessions used here due to their low abundance, this was consistent with the results identified by cDNA microarray3,19. Although the 19 diploid accessions represent a phenotypically diverse set, where nearly 50% of genes showed a measure of differential transcription, in the present study, the 21 genes assayed showed only a low degree of accession-to-accession variation in transcript abundance; thus each of them appears to be useful for normalization purposes9.
In spite of this, there was a little difference in gene function among the 21 genes. For example, AT5G46630, encoded proteins involved in adaptor complex subunits of claturin which mainly involved the protein transport and cell division process3,19, was most stable than others. PP2A subunit (AT1G13320) and AT5G60390 (EF-1α) also shown to be highly stable than others. They were all highly conserved and played ubiquitous roles in protein translation and degradation, binding cytoskeletal proteins and several different signal transduction pathways in the cell3,19. In addition, AT1G62930 and AT5G55840 both belonged to PPR gene were the most unstable genes, which were RNA-binding proteins that were particularly prevalent in terrestrial plants20. These proteins had a range of essential functions in posttranscriptional processes, including RNA editing, RNA splicing, RNA cleavage and translation within mitochondria and chloroplasts20. AT4G38070 was also unstable, as a kind of bHLH transcription factor, it was functionally characterized in Arabidopsis and its roles included regulation of fruit dehiscence, phytochrome signaling, flavonoid biosynthesis, hormone signaling and stress responses21. Interestingly, most of the stable genes were related to the structure and normal physiological function of cells while the unstable genes were not. Perhaps these stable reference genes played conservative roles in cellular structure, divided and physiological state in the normal growth of the diploid accessions.
We provide the ranking of reference genes which might be suitable for accurate normalization and quantification of gene expression studies in different diploid Arabidopsis accessions. However, there is no single reference gene that is the most stable in all samples tested (like AT1G13320). It has been suggested that at least four reference genes should be used for a broad range of tissues or conditions3,19, so although the ranking of the various reference genes could provide a guide for the selection of reference genes, care needs to be taken when analyzing transcription throughout a variety of tissues or conditions and the use of more than one gene is often advisable.
Variation in transcription of reference genes among tetraploid Arabidopsis accessions
It has been recognized that transcription profiles among tetraploid Arabidopsis accessions are more variable than within diploid A. thaliana10,22,23,24. This diversity was reflected in the set of 21 tested reference genes. For seven of the genes, there was no transcription in all allotetraploid accessions. However, it must be pointed that the newly formed Arabidopsis interspecific hybrid of autotetraploid A. arenosa x A. thaliana F1 are self-pollinated to generate A. arenosa x A. thaliana F8, which are A. suecica-like and could be comparable with the natural allotetraploids of A. suecica10,22 and these seven reference genes are all low expression abundance genes in chosen subset with a low RPKM in diploid (Fig. 1). Therefore these genes may enable better normalization and quantification of these genes of low transcript levels in diploid Arabidopsis, but should be neglected in the selection of reference genes for A. suecica or A. suecica-like allopolyploid combinations. We also supposed that high abundance stably expressed reference genes (eg. ACT225,26,27,28,29,30) are more suitable for normalization and quantification of these genes with high expression abundance in tetraploid than the low abundance stably expressed one(s), considering the impact of PCR efficiency, specificity and/or yield for RT-PCR and noise and/or hybridization efficiency of the probe for microarray2,3.
The statistical analysis also suggested that six out of 14 transcribed reference genes showed a significant difference in transcript abundance between diploid and tetraploid Arabidopsis accessions (Fig. 3a), meanwhile the Cv of the 14 genes in tetraploids was more variable than among the diploid A. thaliana accessions (Fig. 3b). Therefore, some mechanisms differentially regulated their expression from diploid to polyploidy maybe exists3,30,31 and determining the mode of regulation of these genes in tetraploids will provide further information regarding the effect of polyploidization on gene expression.
Followed polyploids formation in the plant kingdom, it was observed that the gene expression patterns varied with changes in a ploidy series, however, differences in autotetraploids were usually less pronounced than in allotetraploids32,33,34,35. In the present study, maximum Cv value all appeared in allotetraploids. Autopolyploidy occurred as a consequence of homologous genomes duplication from a single species, whereas allopolyploidy described the union of diverged genomes from different species31,32. Both allo- and auto- tetraploids could be affected by gene dosage generated by the interactions between homoeologous genes, however, the allotetraploid would be affected by not only the gene dosage but also the gene divergence under interspecific hybridization which played a major role in speciation and evolution32,33,36,37,38. It would be likely that these phenomena underlie the poorer stability of the reference genes in tetraploids in comparison to their highly reliable behavior in diploids32,38,39,40.
Interestingly, the most stable gene among the remaining 14 reference genes detectable in both allo- and auto- tetraploids was also AT5G46630, which is the same with the ranking in diploid accessions. AT4G26410 (the 2nd stable in geNorm analysis) encoded uncharacterized conserved protein was related to the cell structure41, while ACT2 (AT3G18780, the 3th stable in geNorm analysis) was structurally constituent of cytoskeleton. These stable reference genes mainly involved in cell structure, cytoskeleton, the protein transport and cell division process in the allo- and auto- tetraploids. Notably, ACT2 was the most commonly used reference gene in polyploid studies26,27,28,29,30 for not only its steady but also the similar expression level between diploids (Fig. 3a). Nevertheless, between different individuals of tetraploid, both phenotypic and genotypic variation and photosynthetic efficiency diversity (like heterosis) may lead to associated genes expression unstable (like GAPDH, the least stable). Hence, studies on the effect of (both allo- and auto-) polyploidy on gene expression are possible; however the genetic basis of ploidy changes in analyzed materials cannot be ignored.
Methods
Transcriptomic data
All data set was obtained from the Gene Expression Omnibus (GEO) (http://www.ncbi.nlm.nih.gov/geo/) database. Diploid transcriptomic sequences of the 19 diploid A. thaliana accessions were acquired from a MAGIC population (GEO accession: GSE30720; Dataset name: GSM762074-GSM762111; a total of 38 transcriptomic data; Plant materials include: seedling stage of Col, Bur, Can, Ct, Edi, Hi, Kn, Ler, Mt, No, Oy, Po, Rsch, Sf, Tsu, Wil, Ws, Wu and Zu; RNA was collected over a three-week period; All plants were grown under a 16/8-h light/dark cycle). Analyzed tetraploids dataset includes 5 tetraploids (GEO accession: GSE29687; Dataset name: GSM736442-GSM736446, a total of 11 transcriptomic data): autotetraploid A. thaliana (A. t; ABRC, CS3900) and A. arenosa (A. a; ABRC, CS3901), two hybrids (A. t × A. a F1 and A. t × A. a F8) between autotetraploid A. thaliana and A. arenosa and the natural allotetraploid A.suecica (A. s; ABRC, CS22508) (Total RNA from 3–4 week A. thaliana; 6–7-week A. arenosa or allotetraploids; All plants were grown under a 16/8-h light/dark cycle).
Selection of candidate reference genes
A set of five conventional (GAPDH, ACT2, UBQ10, UBC, EF-1α) and 16 potentially informative novel genes was assembled and the transcription of each was recorded over at least 80% entries, the selected genes covered a wide range of abundance in absolute expression levels (Table 1)3. Notably, these novel reference genes were chosen from hundreds of Arabidopsis genes which have been previously proven to outperform traditional reference genes in terms of expression stability throughout different developmental stages, organs, tissues, genotypes and under a range of environmental conditions in diploid Col-03. Meanwhile, these novel genes were all excellent for designing gene-specific PCR primers using a standard set of design criteria (e.g., from the 3′-untranslated region, primer Tm = 60 ± 1°C, length 18 to 25 bases, GC content between 40 and 60%, generate a unique, short PCR product between 60 to 150 bp of the expected length), which enabled better normalization and quantification of transcript levels in Arabidopsis2.
Stability was assessed based on geNorm and NormFinder software. The former associates a stability value (M) with each potential reference gene, where a low M reflects stability and a high M instability5. An M value of < 0.5 is conventionally accepted for a reference, but with high M value (≥0.5) should be avoided11. NormFinder ranks genes according to the similarity of their transcript abundances, applying a model-based approach6. The results represent the means ± SD. The statistical differences between diploid and tetraploid Arabidopsis accessions were analyzed according to Tukey's test and the Student's t test (P < 0.05). A coefficient of variation (Cv) was calculated according to the following formula:

A Cv close to 0 is produced when the RPKM (reads per kilobase of exon model per million mapped reads) value of a given gene deviates non-significantly from the mean RPKM19.
References
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29, e45–e45 (2001).
Udvardi, M. K., Czechowski, T. & Scheible, W.-R. Eleven golden rules of quantitative RT-PCR. Plant Cell 20, 1736–1737 (2008).
Czechowski, T., Stitt, M., Altmann, T., Udvardi, M. K. & Scheible, W. R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol 139, 5–17 (2005).
Czechowski, T., Bari, R. P., Stitt, M., Scheible, W. R. & Udvardi, M. K. Real-time RT-PCR profiling of over 1400 Arabidopsis transcription factors: unprecedented sensitivity reveals novel root-and shoot-specific genes. Plant J 38, 366–379 (2004).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3, research0034 (2002).
Andersen, C. L., Jensen, J. L. & Ørntoft, T. F. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64, 5245–5250 (2004).
Mandakova, T., Joly, S., Krzywinski, M., Mummenhoff, K. & Lysak, M. A. Fast diploidization in close mesopolyploid relatives of Arabidopsis. Plant cell 22, 2277–2290 (2010).
Chen, Z. J. et al. The development of an Arabidopsis model system for genome-wide analysis of polyploidy effects. Biol J Linn Soc 82, 689–700 (2004).
Gan, X. et al. Multiple reference genomes and transcriptomes for Arabidopsis thaliana. Nature 477, 419–423 (2011).
Shi, X. et al. Cis-and trans-regulatory divergence between progenitor species determines gene-expression novelty in Arabidopsis allopolyploids. Nature Commun 3, 950 (2012).
Hellemans, J., Mortier, G., De Paepe, A., Speleman, F. & Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol 8, R19 (2007).
Fernandez, P. et al. Comparison of predictive methods and biological validation for qPCR reference genes in sunflower leaf senescence transcript analysis. Plant Cell Rep 30, 63–74 (2011).
Maroufi, A., Van Bockstaele, E. & De Loose, M. Validation of reference genes for gene expression analysis in chicory (Cichorium intybus) using quantitative real-time PCR. BMC Mol Biol 11, 15 (2010).
Hassibi, A., Vikalo, H., Riechmann, J. L. & Hassibi, B. Real-time DNA microarray analysis. Nucleic Acids Res 37, e132 (2009).
Schena, M., Shalon, D., Davis, R. W. & Brown, P. O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Fu, X. et al. Estimating accuracy of RNA-Seq and microarrays with proteomics. BMC genomics 10, 161 (2009).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nature Rev Genet 10, 57–63 (2009).
Fu, X. et al. Estimating accuracy of RNA-Seq and microarrays with proteomics. BMC genomics 10, 161 (2009).
Hong, S. M., Bahn, S. C., Lyu, A., Jung, H. S. & Ahn, J. H. Identification and testing of superior reference genes for a starting pool of transcript normalization in Arabidopsis. Plant & cell physiology 51, 1694–1706 (2010).
Schmitz-Linneweber, C. & Small, I. Pentatricopeptide repeat proteins: a socket set for organelle gene expression. Trends in plant science 13, 663–670 (2008).
Feller, A., Machemer, K., Braun, E. L. & Grotewold, E. Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. The Plant Journal 66, 94–116 (2011).
Ha, M. et al. Small RNAs serve as a genetic buffer against genomic shock in Arabidopsis interspecific hybrids and allopolyploids. Proceedings of the National Academy of Sciences 106, 17835–17840 (2009).
Comai, L., Madlung, A., Josefsson, C. & Tyagi, A. Do the different parental ‘heteromes’ cause genomic shock in newly formed allopolyploids? Philosophical Transactions of the Royal Society of London. Series B: Biological Sciences 358, 1149–1155 (2003).
Madlung, A. et al. Genomic changes in synthetic Arabidopsis polyploids. The Plant Journal 41, 221–230 (2005).
Chen, M., Ha, M., Lackey, E., Wang, J. & Chen, Z. J. RNAi of met1 reduces DNA methylation and induces genome-specific changes in gene expression and centromeric small RNA accumulation in Arabidopsis allopolyploids. Genetics 178, 1845–1858 (2008).
Miller, M., Zhang, C. & Chen, Z. J. Ploidy and Hybridity Effects on Growth Vigor and Gene Expression in Arabidopsis thaliana Hybrids and Their Parents. G3 (Bethesda) 2, 505–513 (2012).
Ni, Z. et al. Altered circadian rhythms regulate growth vigour in hybrids and allopolyploids. Nature 457, 327–331 (2009).
Wang, J., Tian, L., Lee, H. S. & Chen, Z. J. Nonadditive regulation of FRI and FLC loci mediates flowering-time variation in Arabidopsis allopolyploids. Genetics 173, 965–974 (2006).
Yu, Z. et al. Impact of natural genetic variation on the transcriptome of autotetraploid Arabidopsis thaliana. Proceedings of the National Academy of Sciences 107, 17809–17814 (2010).
Paun, O., Fay, M. F., Soltis, D. E. & Chase, M. W. Genetic and epigenetic alterations after hybridization and genome doubling. Taxon 56, 649–656 (2007).
Osborn, T. C. et al. Understanding mechanisms of novel gene expression in polyploids. Trends in genetics: TIG 19, 141–147 (2003).
Chen, Z. J. Genetic and epigenetic mechanisms for gene expression and phenotypic variation in plant polyploids. Annual review of plant biology 58, 377–406 (2007).
Wang, H. et al. Rapid genomic and transcriptomic alterations induced by wide hybridization: Chrysanthemum nankingense x Tanacetum vulgare and C. crassum x Crossostephium chinense (Asteraceae). BMC genomics 14, 902 (2013).
Wang, H. et al. Rapid genetic and epigenetic alterations under intergeneric genomic shock in newly synthesized Chrysanthemum morifolium× Leucanthemum paludosum hybrids (Asteraceae). Genome biology and evolution 6, 247–259 (2014).
McClintock, B. The significance of responses of the genome to challenge. SCIENCES, 180 (1983).
Wang, J. et al. Genomewide nonadditive gene regulation in Arabidopsis allotetraploids. Genetics 172, 507–517 (2006).
Chen, Z. J. Molecular mechanisms of polyploidy and hybrid vigor. Trends in plant science 15, 57–71 (2010).
Jackson, S. & Chen, Z. J. Genomic and expression plasticity of polyploidy. Current opinion in plant biology 13, 153–159 (2010).
Lee, H. S. & Chen, Z. J. Protein-coding genes are epigenetically regulated in Arabidopsis polyploids. Proceedings of the National Academy of Sciences of the United States of America 98, 6753–6758 (2001).
Chen, Z. J. & Ni, Z. Mechanisms of genomic rearrangements and gene expression changes in plant polyploids. Bioessays 28, 240–252 (2006).
Schmid, M. et al. A gene expression map of Arabidopsis thaliana development. Nature genetics 37, 501–506 (2005).
Acknowledgements
We thank Dr Z. Jeffrey Chen (D. J. Sibley Centennial Professor of Plant Molecular Genetics, The University of Texas at Austin) for helpful discussions. The study was supported by the National Natural Science Foundation of China (31372100, 31272203, 31272202, 31272196, 31171987), the Program for New Century Excellent Talents in University of the Chinese Ministry of Education (NCET-12-0890), the Natural Science Fund of Jiangsu Province (BK2012773, BK2011641), the Fundamental Research Funds for the Central Universities” (KYZ201147, KYZ201112), a scholarship granted to Dr. Sumei Chen (File No. 201306855010, CSC, Ministry of Education, China).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: H.W., J.W., J.J., S.C. Performed the experiments: H.W., F.C., J.J., Z.G. Analyzed the data: H.W. Contributed reagents/materials/analysis tools: Y.L. Wrote the paper: H.W., J.J., S.C. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Data S1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Wang, H., Wang, J., Jiang, J. et al. Reference genes for normalizing transcription in diploid and tetraploid Arabidopsis. Sci Rep 4, 6781 (2014). https://doi.org/10.1038/srep06781
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06781
This article is cited by
-
Overexpression of the elongation factor MtEF1A1 promotes salt stress tolerance in Arabidopsis thaliana and Medicago truncatula
BMC Plant Biology (2023)
-
α-Aminoadipic acid metabolism is controlled by the glutathione-dependent redox environment in Arabidopsis
Journal of Plant Biochemistry and Biotechnology (2023)
-
Plastid ribosome protein L5 is essential for post-globular embryo development in Arabidopsis thaliana
Plant Reproduction (2022)
-
Identification of reliable reference genes for quantitative real-time PCR normalization in pitaya
Plant Methods (2019)
-
Reference gene selection for cross-species and cross-ploidy level comparisons in Chrysanthemum spp.
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.