Abstract
Selective binding between pre- and postsynaptic adhesion molecules can induce synaptic differentiation. Here we report the crystal structure of a synaptogenic trans-synaptic adhesion complex between Slit and Trk-like family member 2 (Slitrk2) and receptor protein tyrosine phosphatase (RPTP) δ. The structure and site-directed mutational analysis revealed the structural basis of splicing-dependent adhesion between Slitrks and type IIa RPTPs for inducing synaptic differentiation.
Similar content being viewed by others
Introduction
Synapse formation is initiated at contact sites between axon terminals and dendrites, where pre- and postsynaptic adhesion molecules form trans-synaptic complexes to induce synaptic differentiation. Presynaptic type IIa receptor protein tyrosine phosphatases (RPTPs) (PTPδ, PTPσ and LAR in mammals) can induce synaptic differentiation by binding to various postsynaptic adhesion molecules such as IL-1 receptor accessory protein-like 1 (IL1RAPL1), IL-1 receptor accessory protein (IL-1RAcP), netrin-G ligand-3 (NGL-3), neurotrophin receptor tyrosine kinases C (TrkC) and Slit- and Trk-like (Slitrk) proteins1,2,3,4,5,6,7,8. In mammals, the Slitrk family consists of at least six members (Slitrk1-6)9,10. Mutations in SLITRK genes have been reported to be associated with neuropsychiatric disorders11,12,13,14. All Slitrk members except Slitrk3 can induce both excitatory and inhibitory synapses7,8, whereas Slitrk3 selectively induces inhibitory synapses through binding to PTPδ7. Therefore, it has been proposed that the selective binding between Slitrk proteins and PTPδ plays a role in balancing excitatory and inhibitory synapses7,8.
Selective binding between the type IIa RPTPs and their cognate postsynaptic partners is regulated in part by alternative splicing of the type IIa RPTPs3,4,6. PTPδ expressed in the developing brain consists of three immunoglobulin-like (Ig) and four fibronectin type III (Fn) domains in the extracellular region, where two splice sites exist within Ig2 and between Ig2 and Ig3 (Fig. 1a). Peptide insertions (termed mini-exon peptides) at these two splice sites (referred hereafter to as meA and meB) generate splice variants of PTPδ. In this study, we determined the crystal structure between the first leucine-rich repeat (LRR1) of mouse Slitrk2 and the Ig1–Fn1 domains of PTPδ containing both meA and meB. The structure showed that Slitrk2 LRR1 directly recognizes meB but not meA. Further structure-based mutational analyses using surface-plasmon resonance (SPR) spectroscopy and synaptogenic co-culture assay demonstrated that binding of Slitrk2 to PTPδ depends on meB.
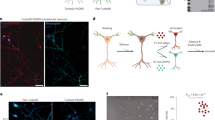
Overall structure of the complex between Slitrk2 LRR1 and PTPδ Ig1-Fn1.
(a) Domain architecture of PTPδ and Slitrk2. (b) Overall structure of the complex between Slitrk2 LRR1 and PTPδ Ig1-Fn1. Disulfide bonds and N-linked glycans are shown as sticks. PTPδ Ig1-Fn1 is colored in green, except that the meA and meB insertions are colored in pink and orange, respectively. The N-terminal cap, central LRR and C-terminal cap of Slitrk2 LRR1 are colored in blue, cyan and magenta, respectively. (c) Structure of Slitrk2 LRR1. Disulfide bonds and N-linked glycans are shown as sticks. Phenylalanine residues in the Phe spine structure are shown as orange sticks. The coloring scheme is the same as that in (b).
Results and Discussion
Overall structure
For structural studies, we initially examined the PTPδ-binding region of Slitrk4 and found that mouse Slitrk4 LRR1 is sufficient for binding to the extracellular domain of PTPδ (PTPδ-ECD) (Supplementary Figure 1a). However, our co-crystallization trials using Slitrk4 LRR1 and the full-length or truncated PTPδ-ECD failed. We then screened other Slitrk members in terms of their expression level in FreeStyle293F cells and their binding activities to PTPδ-ECD and selected mouse Slitrk2 LRR1 as the next candidate for crystallization. After optimization of the length of PTPδ-ECD (Supplementary Figure 1b), we finally determined the crystal structure of a complex between Slitrk2 LRR1 and PTPδ Ig1-Fn1 at 3.35 Å resolution (Fig. 1a,b, Supplementary Figure 2a and Table 1). The asymmetric unit contains one complex, where Slitrk2 LRR1 binds to PTPδ Ig1-Fn1 at a ratio of 1:1 (Fig. 1b, Supplementary Figure 2b). Slitrk2 LRR1 interacts with PTPδ Ig2-3. The overall structure of PTPδ Ig1-Fn1 exhibits an elongated shape. PTPδ Ig1-2 forms a compact V-shaped unit, similarly to LAR or PTPσ Ig1-215. Ig3 is spatially separated from Ig1-2 by meB. The following Fn1 is linearly aligned to Ig3. Slitrk2 LRR1 comprises a central LRR with seven parallel β-strands flanked by N- and C-terminal caps, which are stabilized by disulfide bonds (Fig. 1c). The total nine β-strands in the N-terminal cap and LRR form a concave surface of Slitrk2. The interior of the convex side is rich in completely conserved phenylalanine residues, which form a “Phe spine” structure as observed in the Nogo receptor16 (Fig. 1c and Supplementary Figure 3).
Binding interface
The concave surface of Slitrk2 LRR1 surrounds two strands in PTPδ Ig2 and meB (234ELRE237) with a buried surface area of 644 Å2 (Fig. 2a). Asp167 and Glu215 of Slitrk2 hydrogen bond with Arg236 of PTPδ (the third residue of meB) (Fig. 2b). In addition, Asp142, Asn166 and Asp187 of Slitrk2 hydrogen bond with Arg233 of PTPδ (one-residue upstream of meB). Surrounding these interactions, Arg114 and Arg189 of Slitrk2 hydrogen bond with Gln209 and Glu145 of PTPδ, respectively, whereas Arg114 and His185 of Slitrk2 hydrogen bond with the main-chain O atoms of PTPδ Val232 and Leu141, respectively. Furthermore, Tyr138 of Slitrk2 is stacked with Tyr231 of PTPδ. These Slitrk2 residues involved in the interaction with PTPδ Ig2-meB are mostly conserved in all mouse Slitrk members (Supplementary Figure 3), suggesting that they can potentially recognize PTPδ Ig2-meB in the same manner as Slitrk2.
Binding interface between Slitrk2 and PTPδ.
(a) Interface between Slitrk2 LRR1 and PTPδ Ig2-meB. The coloring scheme is the same as that in Figure 1. (b) Close-up view of (a). (c) Interface between Slitrk2 LRR1 and PTPδ Ig3. The coloring scheme is the same as that in Figure 1. (d) Close-up view of (d).
Slitrk2 LRR1 also interacts with PTPδ Ig3 with a buried surface area of 450 Å2 (Fig. 2c). Phe247 and His250 of Slitrk2 hydrophobically interact with Tyr273 of PTPδ, which hydrogen bonds with the main-chain O atom of Slitrk2 Glu244 (Fig. 2d). Phe247 of Slitrk2 also hydrophobically interacts with Met289 of PTPδ and appears to play a central role in the Slitrk2 LRR1–PTPδ Ig3 interface. However, this phenylalanine residue is replaced by Thr, Ser or Pro in Slitrk1, 4 or 6, respectively (Supplementary Figure 3), which are inadequate to form hydrophobic interactions with Tyr273 and Met289 of PTPδ. In addition, His250 of Slitrk2 is not conserved in Slitrk1, 4 or 6 (Supplementary Figure 3). Therefore, the observed Slitrk2–PTPδ Ig3 interactions may be specific to Slitrk2, 3 and 5.
Splicing-dependent interactions and synaptogenic activity
In the crystal structure of the Slitrk2–PTPδ complex, Slitrk2 LRR1 directly recognizes the meB insertion of PTPδ, whereas the meA insertion is distant from Slitrk2 LRR1 (Fig. 2a). Therefore, the present structure clearly indicates that binding of PTPδ to Slitrk2 depends on meB but not on meA. To ensure this finding, we examined the binding of Slitrk2 LRR1 to the PTPδ variants containing either or both meA and meB (meA9B-, meA-B+ and meA9B+) by surface-plasmon resonance (SPR) spectroscopy (Fig. 3a). As expected, the meA9B+ and meA-B+ variants bound to Slitrk2 LRR1 with similar affinities, whereas the meA9B- variant hardly bound to Slitrk2 LRR1 (Fig. 3a, c and Supplementary Figure 4). We next assessed the observed Slitrk2–PTPδ interactions by site-directed mutagenesis (Fig. 3 and Supplementary Figure 4). The R236E mutation of PTPδ and the D167A and E215A mutations of Slitrk2, which disrupt the meB-specific interactions, abolished the binding between Slitrk2 and PTPδ. The D187A mutation of Slitrk2, which disrupts the hydrogen bond with Arg233 of PTPδ (one-residue upstream of meB), also abolished the binding. Accordingly, the D167A, D187A or E215A mutant of Slitrk2 abolished or significantly reduced their synaptogenic activities in our co-culture assay (Fig. 4). On the other hand, the R114A mutation of Slitrk2 (PTPδ Ig2-mediated interface), the Y273A mutation of PTPδ or the F247A H250A double mutation of Slitrk2 (PTPδ Ig3-mediated interface) hardly affected the binding. Therefore, the R114A or F247A H250A mutant of Slitrk2 exhibited the synaptogenic activity comparable to wild-type Slitrk2 (Fig. 4). These results perfectly support the finding that binding of PTPδ to Slitrk2 depends on meB (Fig. 3 and Supplementary Figure 4).
Recognition of PTPδ meB by Slitrk2 LRR1.
(a) SPR equilibrium analysis of the interaction between Slitrk2 LRR1 and the splicing variants or mutants of PTPδ Ig1-Fn1. Slitrk2 LRR1 was immobilized on a CM5 sensor tip and the indicative PTPδ Ig1-Fn1 proteins were injected. Response units were plotted against concentrations of the indicative PTPδ Ig1-Fn1 proteins. (b) SPR equilibrium analysis of the interaction between Slitrk2 LRR1 mutants and PTPδ Ig1-Fn1 (meA9/meB+). (c)Dissociation constants calculated from (a) and (b).
Synaptogenic activities of Slitrk2 LRR1 mutants.
(a) Co-cultures of cortical neurons and beads conjugated with Fc or Slitrk2 LRR1 mutants fused to Fc. Slitrk2-induced presynaptic differentiation was monitored by immunostaining of the co-cultures with antibodies against Fc (red) and presynaptic active zone protein Bassoon (green). Scale bar, 5 μm. (b) Staining signal intensities for Bassoon on beads conjugated with Fc or Slitrk2 LRR1 mutants fused to Fc. All values represent as mean ± SEM. * p < 0.0001, compared to wild-type Slitrk2-LRR1-Fc. n = 24–29 beads.
Structural comparison with the Slitrk1 LRR1–PTPδ complex
Recently, the crystal structure of the complex between Slitrk1 LRR1 and PTPδ Ig1-3 has been reported17. The main-chain structure of Slitrk1 LRR1 is almost identical to that of Slitrk2 LRR1 with an rmsd of 1.3 Å (Supplementary Figure 5a). Slitrk1 LRR1 interacts with PTPδ Ig2-meB in a similar manner to Slitrk2 LRR1 (Supplementary Figure 5b and c). The hydrophilic interactions between PTPδ Ig2-meB and Slitrk LRR1 are essentially the same, consistent with the sequence conservation between Slitrk1 and 2 (Supplementary Figure 3). On the other hand, the relative position and orientation of PTPδ Ig3 to PTPδ Ig2-meB–Slitrk LRR1 are substantially different between the complexes of Slitrk1 and 2 (Supplementary Figure 5d). Concomitantly, the PTPδ Ig3-mediated interfaces are different between the Slitrk1 and 2 complexes (Supplementary Figure 5e). Physiological significance of this difference remains obscure, because the PTPδ Ig3-mediated interface appears unlikely to contribute to the affinity to Slitrk proteins as mentioned above.
Trans-synaptic clustering of the Slitrk1–PTPδ complex has been proposed on the basis of the lateral interaction between neighboring complexes in the crystal17. This lateral interaction involves Arg72, Phe74 and Arg143 of Slitrk1 and Glu237 and Glu286 of PTPδ. On the other hand, no higher-order clustering was observed in the crystal of the Slitrk2–PTPδ complex (Supplementary Figure 2b). Arg143 of Slitrk1 corresponds to Ser147 of Slitrk2, which is exposed to the solvent (Supplementary Figure 5f). Arg72 and Phe74 of Slitrk1 correspond to Arg76 and Tyr78 of Slitrk2, respectively, which interact with Asp378 of the neighboring PTPδ Fn1 but appear unable to form higher-order clustering (Supplementary Figure 5f).
In conclusion, we revealed that binding of Slitrk2 to PTPδ depends on meB but not on meA, based on the crystal structure and mutational studies at the molecular and cellular levels. Our previous analysis of cDNA from the developing mouse brain showed that PTPσ and LAR also have the splicing variants containing a four-residue insert as meB with the conserved third arginine residue4, suggesting that the meB-containing variants of all type IIa RPTPs can potentially bind to all Slitrk proteins. In fact, a recent study showed that the meB-containing variant of LAR can bind to Slitrk1 (ref. 17). Other example of splicing-dependent regulation of trans-synaptic adhesion through the type IIa RPTPs is the adhesion between PTPδ and IL1RAPL1 or IL-1RAcP3,4. Further studies for these complexes, together with the present study, will lead to complete understanding of splicing-dependent regulation of trans-synaptic adhesions for synaptic differentiation.
Methods
Protein expression and purification
The gene encoding mouse PTPδ Ig1-Fn1 (residues 28–418) was amplified from cDNA (accession No. NM_011211.3) by PCR and cloned into pEBMulti-Neo vector (Wako Pure Chemical Industries) with the N-terminal signal sequence derived from pHLsec vector18 and a C-terminal hexahistidine tag. The gene encoding mouse Slitrk2 LRR1 was cloned into pEBMulti-Neo vector with a C-terminal hexahistidine tag. The proteins were transiently expressed in Freestyle293F cells and purified by Ni-affinity chromatography. Slitrk2 LRR1 was further purified by size-exclusion chromatography with Superdex200 increase (GE healthcare).
Crystallization and structure determination
We tested the interaction between the various truncation mutants of mouse Slitrk proteins and PTPδ by pull down experiments using Fc-fused PTPδ. Consequently, we found that first LRR domain (LRR1) of Slitrk proteins and PTPδ extracellular regions containing Ig1-Fn1 are sufficient for their binding. Slitrk2 LRR1 and PTPδ Ig1-Fn1 were mixed at a molar ratio of 1:1. Prior to the crystallization, the sample was treated with neuraminidase for trimming N-linked glycans. Crystallization was carried out with the sitting drop vapor diffusion method at 293 K by mixing the equal volumes (1 μL) of the protein solution and the mother liquor containing 15% PEG4000, 0.1 M sodium acetate, 0.1 M MES (pH 6.0). The crystals were flash-frozen in liquid nitrogen after soaking in the mother liquor containing 25% ethylene glycol for cryoprotection.
Diffraction data were collected at 100 K at BL41XU in SPring-8 and processed with HKL200019. The structure of Slitrk2 LRR1 and PTPδ Ig1-Fn1 was solved by the molecular replacement method using the program MolRep20. For PTPδ Ig1-Fn1, Ig1-2 of PTPδ (PDB ID:2YD6), Ig3 of PTPσ (PDB ID:2YD9) and Fn2 of LAR (PDB ID:2DJU) were used as the search model. For Slitrk2 LRR1, the Nogo receptor LRR domain (PDB ID:1OZN) was used as the search model. Model building was carried out using the program Coot21. The model was refined using the program Phenix22 to the final Rwork/Rfree factors of 23.46/28.65%. The stereochemistry of the final model was assessed by Procheck23. Data collection and refinement statistics were summarized in Table 1. The buried surface area was calculated using the program PISA24. All structural figures are prepared using the program PyMol (Schrödinger, LLC).
SPR analysis
SPR experiments were carried out by using Biacore T200 (GE healthcare) at 25°C in 10 mM HEPES-Na buffer (pH 7.9) containing 150 mM NaCl and 0.05% Tween-20. The wild-type or mutant Slitrk2 LRR1 (D167A, D187A, E215A, R114A or F247A H250A) was immobilized on a CM5 sensor tip by the amine-coupling method. The splicing variant (meA9B+, meA9B- or meA-B+) or mutant (R236E or Y273A) of PTPδ Ig1-Fn1 was injected at concentrations ranging from 15.625 to 4,000 nM. 10 mM NaOH was used as a regeneration buffer.
Synaptogenic assay
Primary cortical cultures were prepared from mice at E18 essentially as described previously4. Expression vectors for the mutated forms of Slitrk2-LRR1-Fc were generated by PCR-based mutagenesis using pEB6-Slitrk2-LRR1-Fc as a template. Fc and the mutated forms of Slitrk2-LRR1-Fc in FreeStyle293F cell culture medium were bound to Protein A-conjugated magnetic particles (smooth surface, 4.0–4.5 μm diameter; Spherotech). Beads coupled with Fc or the Fc fusion proteins were added to cortical neurons at days in vitro 16. After 24 hours, cultures were fixed for immunostaining with mouse anti-Bassoon antibody (Stressgen, 1:400) and rabbit anti-human Fc (Rockland, 1:1000).
Image acquisition and quantification
Images of bead-neuron co-cultures were collected from at least two separate experiments. Image acquisition and quantification for the co-culture assays were performed essentially as previously described4. Briefly, the intensities of immunostaining signals for Bassoon were measured as the optical mean density within a circle of 7-μm diameter enclosing coated-bead. Statistical significance was evaluated by one-way ANOVA followed by post hoc Tukey’s test.
Additional information
How to cite this article: Yamagata, A. et al. Structure of Slitrk2–PTPδ complex reveals mechanisms for splicing-dependent trans-synaptic adhesion. Sci. Rep. 5, 9686; DOI:10.1038/srep09686 (2015).
References
Takahashi, H. & Craig, A. M. Protein tyrosine phosphatases PTPδ, PTPσ and LAR: presynaptic hubs for synapse organization. Trends Neurosci. 36, 522–534 (2013).
Um, J. W. & Ko, J. LAR-RPTPs: synaptic adhesion molecules that shape synapse development. Trends Cell Biol. 23, 465–475 (2013).
Yoshida, T. et al. Interleukin-1 receptor accessory protein organizes neuronal synaptogenesis as a cell adhesion molecule. J. Neurosci. 32, 2588–2600 (2012).
Yoshida, T. et al. IL-1 receptor accessory protein-like 1 associated with mental retardation and autism mediates synapse formation by trans-synaptic interaction with protein tyrosine phosphatase δ. J. Neurosci. 31, 13485–13499 (2011).
Woo, J. et al. Trans-synaptic adhesion between NGL-3 and LAR regulates the formation of excitatory synapses. Nat. Neurosci. 12, 428–437 (2009).
Takahashi, H. et al. Postsynaptic TrkC and presynaptic PTPσ function as a bidirectional excitatory synaptic organizing complex. Neuron 69, 287–303 (2011).
Takahashi, H. et al. Selective control of inhibitory synapse development by Slitrk3-PTPdelta trans-synaptic interaction. Nat. Neurosci. 15, 389–398 (2012).
Yim, Y. S. et al. Slitrks control excitatory and inhibitory synapse formation with LAR receptor protein tyrosine phosphatases. Proc. Natl. Acad. Sci. U. S. A. 110, 4057–4062 (2013).
Aruga, J. & Mikoshiba, K. Identification and characterization of Slitrk, a novel neuronal transmembrane protein family controlling neurite outgrowth. Mol. Cell. Neurosci. 24, 117–129 (2003).
Proenca, C. C., Gao, K. P., Shmelkov, S. V., Rafii, S. & Lee, F. S. Slitrks as emerging candidate genes involved in neuropsychiatric disorders. Trends Neurosci. 34, 143–153 (2011).
Abelson, J. F. et al. Sequence variants in SLITRK1 are associated with Tourette's syndrome. Science 310, 317–320 (2005).
Zuchner, S. et al. SLITRK1 mutations in trichotillomania. Mol. Psychiatry 11, 887–889 (2006).
Piton, A. et al. Systematic resequencing of X-chromosome synaptic genes in autism spectrum disorder and schizophrenia. Mol. Psychiatry 16, 867–880 (2011).
Smith, E. N. et al. Genome-wide association of bipolar disorder suggests an enrichment of replicable associations in regions near genes. PLoS Genet. 7, e1002134 (2011).
Coles, C. H. et al. Proteoglycan-specific molecular switch for RPTPσ clustering and neuronal extension. Science 332, 484–488 (2011).
He, X. L. et al. Structure of the Nogo receptor ectodomain: a recognition module implicated in myelin inhibition. Neuron 38, 177–185 (2003).
Um, J. M. et al. Structural basis for LAR-RPTP/Slitrk complex-mediated synaptic adhesion. Nat. Commun. 5, 5423 (2014).
Aricescu, A.R., Lu, W.. & Jones, E.Y. A time- and cost-efficient system for high-level protein production in mammalian cells.. Acta. Crystallogr. D 62, 1243–50 (2006).
Otwinowski, Z. & Minor, W. Processing of X-ray Diffraction Data Collected in Oscillation Mode. Methods Enzymol. 276, 20 (1997).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 4 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. The Phenix software for automated determination of macromolecular structures. Methods 55, 94–106 (2011).
Laskowski, R. A., Macarthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallog. 26, 9 (1993).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Acknowledgements
We thank the beam-line staffs at BL41XU of SPring8 (Hyogo, Japan) for technical help during data collection. This work was supported by Grant-in-Aid for Scientific Research on Innovative Areas 22121003 (S.F.), 25117711 (Y.S.) and 25112505 (Y.S.), Grant-in-Aid for Scientific Research (A) 24247014 (S.F.), Grant-in-Aid for Scientific Research (B) 25293057 (T.Y.), Grant-in-Aid for challenging Exploratory Research 26640038 (T.Y.) and 25650018 (S.G.-I.), Grant-in-Aid for Scientific Research (C) 24570126 (A.Y.), Grant-in-Aid for Young Scientists (A) 24687012 (Y.S.), PRESTO, JST (T.Y.) and CREST, JST (S.F. and T.U.).
Author information
Authors and Affiliations
Contributions
A.Y., A.M., T.S., T.U. and T.Y. performed gene cloning, protein purification and crystallization. A.Y., S.F., Y.S. and S.G.-I. collected the diffraction data sets. A.Y. and S.F. analyzed the collected data and determined the structures. T.S. performed SPR measurement. A.Y. and S.F. wrote the paper with editing by T.U. and T.Y., S.F. designed and supervised the study.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Accession codes: Coordinates and structure factors of the Slitrk2 LRR1–PTPδ Ig1-Fn1 complex have been deposited in the Protein Data Bank under accession code 4Y61.
Electronic supplementary material
Supplementary Information
Supplementary Figures 1-5
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Yamagata, A., Sato, Y., Goto-Ito, S. et al. Structure of Slitrk2–PTPδ complex reveals mechanisms for splicing-dependent trans-synaptic adhesion. Sci Rep 5, 9686 (2015). https://doi.org/10.1038/srep09686
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep09686
This article is cited by
-
MicroExonator enables systematic discovery and quantification of microexons across mouse embryonic development
Genome Biology (2021)
-
Canonical versus non-canonical transsynaptic signaling of neuroligin 3 tunes development of sociality in mice
Nature Communications (2021)
-
Genome-wide association study of pediatric obsessive-compulsive traits: shared genetic risk between traits and disorder
Translational Psychiatry (2021)
-
Structural insights into selective interaction between type IIa receptor protein tyrosine phosphatases and Liprin-α
Nature Communications (2020)
-
Structural basis of SALM3 dimerization and synaptic adhesion complex formation with PTPσ
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.