-
PDF
- Split View
-
Views
-
Cite
Cite
Hironori Itoh, Akie Sasaki, Miyako Ueguchi-Tanaka, Kanako Ishiyama, Masatomo Kobayashi, Yasuko Hasegawa, Eiichi Minami, Motoyuki Ashikari, Makoto Matsuoka, Dissection of the Phosphorylation of Rice DELLA Protein, SLENDER RICE1, Plant and Cell Physiology, Volume 46, Issue 8, August 2005, Pages 1392–1399, https://doi.org/10.1093/pcp/pci152
Close - Share Icon Share
Abstract
DELLA proteins are repressors of gibberellin signaling in plants. Our previous studies have indicated that gibberellin signaling is derepressed by SCFGID2-mediated proteolysis of the DELLA protein, SLENDER RICE1 (SLR1), in rice. In addition, the gibberellin-dependent increase of phosphorylated SLR1 in the loss-of-function gid2 mutant suggests that the SCFGID2-mediated degradation of SLR1 might be initiated by gibberellin-dependent phosphorylation. To confirm the role of phosphorylation of SLR1 in its gibberellin-dependent degradation, we revealed that SLR1 is phosphorylated on an N-terminal serine residue(s) within the DELLA/TVHYNP and polyS/T/V domain. However, gibberellin-induced phosphorylation in these regions was not observed in the gid2 mutant following the constitutive expression of SLR1 under the control of the rice actin1 promoter. Treatment with gibberellin induced both the phosphorylated and non-phosphorylated forms of SLR1 with similar induction kinetics in gid2 mutant cells. Both the phosphorylated and non-phosphorylated SLR1 proteins were degraded by gibberellin treatment with a similar half-life in the rice callus cells, and both proteins interacted with recombinant glutathione S-transferase (GST)–GID2. These results demonstrate that the phosphorylation of SLR1 is independent of its degradation and is dispensable for the interaction of SLR1 with the GID2/F-box protein.
Introduction
Regulated protein degradation is essential for the growth and development of all organisms. Many short-lived proteins are degraded through the ubiquitin-mediated proteolytic pathway (Hershko and Ciechanover 1998, Deshaies 1999, Pickart 2001). In ubiquitin-mediated proteolysis, activated ubiquitin is transferred to the substrate protein by a ubiquitin ligase (E3). Then, polyubiquitinated proteins are recognized and finally degraded by the 26S proteasome.
Since the ubiquitin-mediated proteolytic system shows the strict specificity of substrate ubiquitination, the E3 ligase has been considered as a regulatory element of the ubiquitin–proteasome pathway. The SCF complex is one member of the E3 ubiquitin ligases which is structurally and functionally well characterized in mammals, yeast and plants (Patton et al. 1998). The complex is named after the first three subunits identified in yeast and mammals: Skp1, cullin1 (or Cdc53) and F-box protein. In this complex, the F-box protein functions as a receptor that recruits a protein substrate for ubiquitin-mediated degradation (Hershko and Ciechanover 1998, Deshaies 1999). According to the sequence analysis of the Arabidopsis genome, >700 genes are predicted to encode an F-box protein (del Pozo and Estelle 2000, Gagne et al. 2002, Kuroda et al. 2002). Some of these proteins have been identified as components of SCF complexes involved in auxin, jasmonate, ethylene and light signaling (Ingram et al. 1997, Ruegger et al. 1998, Xie et al. 1998, Nelson et al. 2000, Somers et al. 2000, Gray et al. 2001). These studies suggested that, like yeast and mammals, plants require the SCF-mediated proteolytic pathway for various signaling pathways.
The signaling pathway involving the phytohormone gibberellin is also controlled by the SCF-mediated proteolytic pathway (McGinnis et al. 2003, Sasaki et al. 2003). Gibberellin regulates many growth and developmental processes throughout the life cycles of plants (Davies 1995). Recent studies in certain plant species have shown that the gibberellin signaling pathway is regulated by the activity of DELLA protein(s). DELLA proteins have been shown to function as negative regulators of gibberellin signaling in Arabidopsis, rice, barley, wheat and maize (Olszewski et al. 2002). DELLA proteins belong to a plant-specific family of putative transcription factors known as GRAS [Gibberellin-INSENSITIVE (GAI), REPRESSOR OF ga1-3 (RGA) and SCR] (Pysh et al. 1999). Molecular and biochemical studies of DELLA proteins, including SLR1 (SLENDER RICE1) in rice, RGA in Arabidopsis and SLN1 (SLENDER1) in barley, have demonstrated that these proteins are localized in nuclei and are rapidly degraded in response to a gibberellin signal (Dill et al. 2001, Silverstone et al. 2001, Gubler et al. 2002, Itoh et al. 2002).
The recent discoveries of GID2 and SLY1 genes in rice and Arabidopsis, respectively, have clarified how DELLA proteins are degraded. Previous studies have shown that GID2/SLY1 genes encode F-box proteins. Furthermore, loss-of-function mutants, gid2 and sly1, are no longer sensitive to gibberellin signaling owing to the accumulation of a DELLA protein, SLR1 or RGA, respectively (McGinnis et al. 2003, Sasaki et al. 2003). More recently, biochemical studies have demonstrated that GID2 and SLY1 proteins function in vivo as components of an SCF complex and directly interact with their respective DELLA proteins, SLR1 and RGA/GAI (Dill et al. 2004, Fu et al. 2004, Gomi et al. 2004). Collectively, these results indicate that the gibberellin signal induces the proteolysis of SLR1/RGA/GAI through the SCFGID2/SLY1 ubiquitin ligase.
Biochemical studies of yeast and mammals have shown that the interaction of F-box proteins with protein substrates is dependent on modifications such as phosphorylation, glycosylation and hydroxylation (Deshaies 1999). Previous studies have demonstrated an increase in the level of phosphorylated SLR1 in response to gibberellin signaling in the gid2 mutant (Sasaki et al. 2003). Furthermore, phosphorylated SLR1 binds to recombinant glutathione S-transferase (GST)–GID2, suggesting that the gibberellin-induced phosphorylation of SLR1 induces the interaction with the SCFGID2 complex and subsequent degradation (Gomi et al. 2004).
In the present study, SLR1 was shown to be phosphorylated on serine residues in the gibberellin signal perception domain (DELLA and/or TVHYNP) and regulatory domain (polyS/T/V). However, gibberellin-induced phosphorylation was not detected following the constitutive expression of SLR1 protein. Furthermore, SLR1 was phosphorylated in the absence of gibberellin, and de-phosphorylated SLR1 still interacted with GID2 protein. Taken together, these results demonstrate that the phosphorylation of SLR1 is not essential for its interaction with GID2/F-box protein.
Results
Phosphorylated region of the SLR1 protein
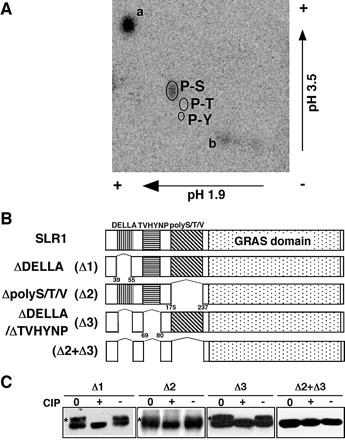
Our previous studies have shown that gibberellin treatment of the gid2 mutant resulted in the accumulation of phosphorylated SLR1 (Sasaki et al. 2003) and this phosphorylated SLR1 is capable of binding to GID2 protein (Gomi et al. 2004). To investigate further the phosphorylation of SLR1, we attempted to identify the phosphorylation sites of SLR1 in the gid2 mutant. Following the treatment of gid2 plants with 32PO4–, 32P-labeled SLR1 was purified using anti-SLR1 antibody and then subjected to acid hydrolysis. The resulting amino acids were separated using two-dimensional thin-layer electrophoresis. One radioactive spot was detected at a phosphoserine site (Fig. 1A), demonstrating that SLR1 was phosphorylated on serine residue(s). No signals corresponding to either phosphothreonine or phosphotyrosine were detected (dataa not shown).
Next, we generated transgenic rice plants that expressed four types of SLR1 deletion derivatives in the gid2 mutant (Fig. 1B). More than 20 independent transgenic plants for each construct were produced and used in the following experiments. To distinguish these derivatives from endogenous SLR1, each mutant SLR1 was tagged with hemagglutinin (HA) or green fluorescent protein (GFP). The phosphorylated SLR1 protein (form II, indicated by asterisks in Fig. 1C), which was confirmed by the disappearance of the lower mobility band following calf intestine alkaline phosphatase (CIP) treatment, migrated more slowly than the non-phosphorylated form (form I) by SDS–PAGE (see below, Sasaki et al. 2003). The lower mobility form of ΔDELLA-SLR1 protein (Δ1) disappeared following CIP treatment (panel Δ1 in Fig. 1C), suggesting that the ΔDELLA-SLR1 protein is phosphorylated. Similar results were also seen with ΔpolyS/T/V or ΔDELLA+ΔTVHYNP (panels Δ2 and Δ3 in Fig. 1C, respectively). However, the SLR1 protein with all three domains deleted was detected as a single band (panel Δ2+Δ3 in Fig. 1C), indicating no in vivo phosphorylation of this mutant protein. These results suggest that SLR1 phosphorylation occurs in at least two of the three regions, N-terminal DELLA, TVHYNP and polyS/T/V, although there is an alternative possibility that the region(s) is not phosphorylated itself but essential for phosphorylation of other sites.
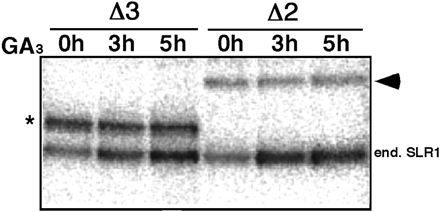
It is noteworthy that the intensity of the lower mobility band of ΔpolyS/T/V protein (panel Δ2 in Fig. 1C) was much weaker than that observed for ΔDELLA+ΔTVHYNP protein (panel Δ3 in Fig. 1C). Consistent with this, the data presented in Fig. 2 show that the intensity of the 32P-labeled ΔDELLA+ΔTVHYNP band (indicated by the asterisk) was stronger than that of the 32P-labeled ΔpolyS/T/V band (indicated by the arrowhead). These results indicate that there are multiple phosphorylation sites within these regions and that the majority of the phosphorylation is localized in the polyS/T/V region.
Phosphorylation state of SLR1 protein in the gid2 mutant
To determine which region is responsible for the gibberellin-induced phosphorylation of SLR1, the effect of GA3 on the phosphorylation of each mutated protein was examined. Under normal conditions, gid2 mutant plants accumulate high levels of bioactive gibberellin, GA1 (Sasaki et al. 2003). Therefore, the transgenic gid2 plants were grown with a gibberellin synthesis inhibitor (1 µM uniconazol) for 2 weeks, incubated with 32PO4–, and then treated with 100 µM GA3 for 0–5 h. The endogenous and mutated 32P-labeled SLR1 proteins were immunoprecipitated with anti-SLR1 antibody. Autoradiographic analysis showed that each phosphorylated SLR1 corresponded to an anti-SLR1 immunoreactive band (data not shown). Consistent with previous results (Sasaki et al. 2003), the level of the endogenous phosphorylated SLR1 was increased by GA3 treatment (indicated by ‘end. SLR1′ in Fig. 2). However, the phosphorylation of mutated SLR1 proteins expressed under the control of the actin1 promoter was not induced under the same conditions (indicated by the asterisk and arrowhead in Fig. 2).
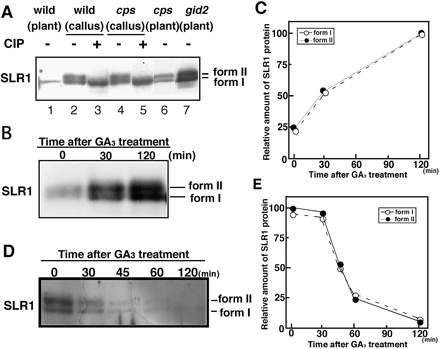
To understand why GA3 treatment did not increase the phosphorylation of the mutant proteins but did increase phosphorylation of endogenous SLR1, the pattern of phosphorylated SLR1 accumulation in gid2 mutant plants was analyzed. The gid2 callus was used for these studies because rice callus cells contained very low levels of bioactive gibberellin (Table 1), making it unnecessary to pre-treat this material with a gibberellin synthesis inhibitor. A high level of anti-SLR1 immunoreactive protein was detected in the crude extract of wild-type (WT) callus, while a low level was found in the WT seedlings (compare lanes 1 and 2 in Fig. 3A). The band intensity of the immunoreactive protein in the WT callus was almost the same as that in the callus of the cps mutant (compare lanes 2 and 4 in Fig. 3A), which is the most severe gibberellin-deficient mutant in rice (Sakamoto et al. 2004). These results confirm that the WT callus contained a very low level of bioactive gibberellin. Surprisingly, SLR1 in the WT and cps calli was detected as a doublet band by immunoblot analysis, and the lower mobility form (form II) disappeared following CIP treatment (Fig. 3A, lanes 3 and 5). This indicates that the phosphorylation of SLR1 occurred even in the absence of gibberellin. Similar migration patterns were observed for the doublet bands of these calli in the gid2 and cps seedlings, confirming that the phosphorylation of SLR1 is similar in calli and seedlings (Fig. 3A, lanes 6 and 7).
Next, we examined the change in SLR1 levels in the gid2 callus following gibberellin treatment. The treatment with GA3 resulted in an accumulation of both form I and form II SLR1 in a similar manner (Fig. 3B, C). This result suggests that the accumulation of phosphorylated SLR1 form II induced by GA3 treatment may be the result of the gibberellin-dependent accumulation of SLR1 caused by the transcriptional regulation of the endogenous SLR1 promoter (M. Ueguchi-Tanaka and M. Matsuoka, unpublished data). In this context, it is reasonable that the level of phosphorylated SLR1 was not altered by gibberellin treatment under the control of the actin1 promoter (Fig. 2).
Using the WT callus, the time course of SLR1 degradation following GA3 treatment was examined. As shown in Fig. 3D and E, both form I and form II SLR1 proteins were degraded with the same kinetics. Furthermore, a band shift from form I (non-phosphorylated) to form II (phosphorylated) was not detected. These results suggest that phosphorylated SLR1 is not selectively or preferentially degraded as a result of the gibberellin signal.
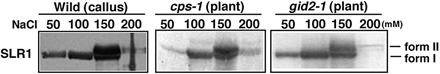
Elution profile of form I and form II SLR1 in DEAE chromatography
To determine whether the phosphorylation status of each mutant varied, form I and form II SLR1 were fractionated by DEAE ion-exchange chromatography (Gomi et al. 2004). Following extraction from gid2 (gibberellin overdose with a defect in SCFGID2 activity), form I SLR1 was eluted with 50, 100 and 150 mM NaCl. However, form II was not eluted until 150 mM NaCl and was also faintly detected in the 200 mM NaCl fraction (Fig. 4). The elution patterns of SLR1 from the WT callus (gibberellin-deficient condition) and cps seedlings (gibberellin-deficient condition) were essentially the same as that of SLR1 from the gid2 mutant, indicating that the levels of phosphorylated SLR1 are similar among these cells despite the presence or absence of the gibberellin signal.
Phosphorylation is not essential for the interaction between SLR1 and GID2 proteins
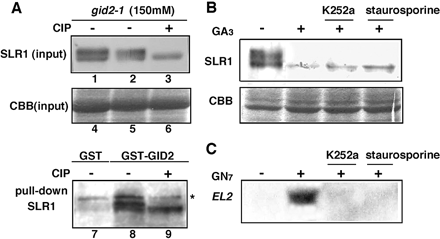
The series of results presented above suggest that phosphorylation of SLR1 is not an essential factor for gibberellin-dependent degradation. To address this question further, we examined the in vitro interaction of GID2/F-box protein with non-phosphorylated SLR1. The protein sample of the gid2 mutant eluted from the DEAE column at 150 mM NaCl was treated with CIP to dephosphorylate the SLR1 protein and was used in a subsequent pull-down assay with recombinant GST–GID2. The treatment with CIP resulted in the complete dephosphorylation of SLR1 (compare lanes 2 and 3 in Fig. 5A). Although a small amount of SLR1 was precipitated with GST only (lane 7 in Fig 5A), in the CIP-non-treated eluate, a much higher amount of both phosphorylated and non-phosphorylated SLR1 was pulled-down with GST–GID2 (lane 8 in Fig. 5A), indicating interaction of both phosphorylated and non-phosphorylated SLR1 with GST–GID2. Interaction of non-phosphorylated SLR1 with GST–GID2 was confirmed by the pull-down of dephosphorylated SLR1 in the CIP-treated eluate with GST–GID2 (lane 9 in Fig. 5A). Under this experimental condition, a band with lower mobility than that of form II phosphorylated SLR1 was always precipitated with GST protein in a non-specific manner (indicated by the asterisk in Fig. 5A). We performed three independent pull-down experiments and obtained the same results as presented above. These observations confirm that precipitation of phosphorylated and non-phosphorylated SLR1 specifically occurred due to GST–GID2 in these experiments.
The effects of kinase inhibitors on the gibberellin-dependent degradation of SLR1 were also analyzed to determine whether de novo phosphorylation of SLR1 is essential for its in vivo degradation. K252a and staurosporine are general inhibitors of Ser/Thr kinase and Tyr kinase, respectively. Gibberellin-dependent degradation of form I or form II SLR1 protein was not inhibited in the WT callus by K252a or staurosporine (Fig. 5B). For control, we examined the effects of these kinase inhibitors on the expression of an elicitor-induced gene, EL2, in the rice callus, and found that these inhibitors completely inhibited the enhancement of EL2 expression by the elicitor, GN7, under the same condition (Fig. 5C). These experiments with kinase inhibitors support that the phosphorylation of SLR1 is not necessary for its gibberellin-dependent degradation.
Discussion
In contrast to previous reports that suggested the gibberellin-dependent degradation of DELLA proteins is dependent on protein phosphorylation (Fu et al. 2002, Gomi et al. 2004), our current studies have demonstrated that the phosphorylation of SLR1 occurs even in the absence of the gibberellin signal (Fig. 3A). Furthermore, dephosphorylated SLR1 was also shown to bind to GST–GID2 (Fig. 5). The phosphorylation-dependent substrate degradation in SCF-mediated proteolysis is achieved by the direct interaction between F-box proteins and the phosphorylated region (phosphodegron) of the substrate (Winston et al. 1999, Nash et al. 2001, Yeh et al. 2001). Therefore, dephosphorylation should diminish the interaction between the F-box protein and its substrate. In this context, the interaction between dephosphorylated SLR1 and GID2/F-box protein strongly suggests that the gibberellin-dependent degradation of SLR1 does not depend on the phosphodegron-dependent machinery.
We have shown previously that form II SLR1, which is eluted at 250 mM NaCl from a DEAE column, bound to GID2 protein, but that form I SLR1, which is eluted at 100 mM NaCl, did not (Gomi et al. 2004). Moreover, form II SLR1 (phosphorylated) was enriched in fractions that eluted at higher concentrations of NaCl, while form I SLR1 (non-phosphorylated) was eluted at lower NaCl concentrations (Gomi et al. 2004). Based on these observations, we previously proposed that the phosphorylated SLR1 specifically interacted with GID2, resulting in its degradation (Gomi et al. 2004). However, our current studies demonstrate that these two phenomena are irrelevant. Although still we cannot exclude the possibility that the results with the gid2 mutant may not reflect the situation in the WT, we predict that there are at least two different types of modification for SLR1 protein, i.e. phosphorylation/dephosphorylation and an as yet unidentified gibberellin signal received/non-received. We did not use the SLR1 protein prepared from rice callus cells for the in vitro binding assay, since gibberellin signal non-received SLR1 cannot be recognized by GID2 protein even though the SLR1 is highly accumulated in such a gibberellin-deficient background (data not shown).
In this study, we found a trend similar to the previous result that the SLR1 eluted at a higher concentration of NaCl exhibited a higher affinity for GID2 than did the SLR1 eluted at a lower NaCl concentration. This suggests that the gibberellin signal received form SLR1 may be more negatively charged than the non-received form. Enrichment of the phosphorylated form (form II) and the gibberellin signal received form in the eluate of the high NaCl fraction led us to an incorrect speculation that phosphorylated SLR1 corresponds to the received form gibberellin signal.
Recently, Dill et al. (2004) reported that the N-terminal truncated GAI protein lacking the DELLA, TVHYNP and poly S/T/V regions can still interact with an SLY1 gain-of-function mutant, sly1-d. Based on this observation, they concluded that the C-terminal conserved GRAS region in RGA/GAI is responsible for the direct interaction with SLY1. In this study, we suggested that SLR1 phosphorylation occurs in the DELLA/TVHYNP domain and in the polyS/T/V region, but not in the C-terminal GRAS region (Fig. 1). Collectively, these results suggest that phosphorylation is not necessary for the interaction between DELLA proteins and the F-box proteins GID2 and SLY1. Recent studies by Fu et al. (2004) have demonstrated that gai is phosphorylated in vivo and that phosphorylated gai has a higher in vitro binding affinity for SLY1 than does dephosphorylated gai (Fu et al. 2004). This observation is seemingly contradictory to the result shown in Fig. 5A. However, there is one difference between the experimental conditions of the two studies. Fu et al. (2004) used the DELLA-truncated version of GAI (gai-1) for their experiment, while we used full-length SLR1 protein. As the gai-1 protein is stable in vivo even under the gibberellin-treated condition, its molecular features would be different from those of full-length DELLA protein, which is rapidly degraded by the gibberellin signal. Actually, the binding affinity between gai-1 and SLY1 was much higher than that between full-length GAI and SLY1 in yeast cells (Dill et al. 2004). Therefore, it is possible that phosphorylation may exhibit different effects on the binding of full-length and DELLA-truncated DELLA proteins to F-box proteins. In this context, it is curious that gai-1 and rga-Δ17 showed higher binding affinity for SLY1/F-box protein than did either full-length mutant, although these truncated proteins are resistant to gibberellin-dependent degradation. These results indicate that the interaction of DELLA proteins with SLY1 or GID2 in yeast does not correspond to the degradation of DELLA proteins in plants. Similarly, in vitro binding of SLR1 to GID2 may not be indicative of the molecular feature of SLR1 in vivo. Further studies are needed to clarify the effect of gibberellin-dependent modifications of SLR1 and its degradation by the SCFGID2 pathway.
Materials and Methods
Plant materials
The gid2 and cps mutants, and their original cultivar Oryza sativa L. cv. Taichung 65, were used in this study. Rice seeds were immersed in water for 2 d and grown in a greenhouse. Suspension-cultured cells of O. sativa L. cv. Nipponbare were maintained in modified N-6 medium and were subcultured weekly as previously described (Kuchitu et al. 1993).
Analysis of gibberellin content
Quantitative analyses of the endogenous gibberellins were performed by gas chromatography-selected ion monitoring (GC/SIM) (Kobayashi et al. 1995).
Plant transformation and growth conditions
Binary vectors were introduced into Agrobacterium tumefaciens strain EHA101 (Hood et al. 1986) by electroporation. Rice transformation was performed as described by Hiei et al. (1994). Transgenic plants were selected on medium containing 50 mg l–1 hygromycin. Hygromycin-resistant plants were transplanted to soil and grown at 30°C in a 16 h light/8 h dark cycle.
Plasmid construction
The binary plasmids, HA-ΔDELLA-GFP and HA-ΔpolyS/T/V-GFP, were constructed as previously described (Itoh et al. 2002). The truncated constructs of ΔDELLA+ΔTVHYNP and ΔDELLA+ΔTVHYNP+ΔpolyS/T/V were encoded as three copies of the HA epitope (Gomi et al. 2004) fusion at their N-termini. These SLR1 derivative constructs were ligated into the binary vector pBI-101 at the position between the actin1 promoter and a NOS terminator.
Preparation of protein extracts and immunoblot analysis
Total proteins were extracted by grinding seedlings or calli with liquid nitrogen and then resuspending the ground tissue in extraction buffer [20 mM Tris–HCl, pH 7.5, 150 mM NaCl, 0.5% Tween-20, 1 mM EDTA, 1 mM dithiothreitol (DTT)] containing the Complete protease inhibitor cocktail (Roche, Mannheim, Germany). For CIP treatment, crude protein extracts were incubated with 100 U of CIP at 37°C for 15 min as previously described (Sasaki et al. 2003). After the addition of an equal volume of 2× sample buffer [1× sample buffer: 67.5 mM Tris–HCl (pH 6.8), 2% (w/v) SDS, 10% (w/v) glycerol, 0.01% (w/v) bromophenol blue and 0.1 M DTT], the samples were boiled for 5 min, separated by 7.5% SDS–PAGE (Sambrook and Russell 2000) and transferred to a Hybond ECL membrane (Amersham Pharmacia Biotech, Little Chalfont, UK) by semi-dry blotting. The blots were incubated with rabbit anti-SLR1 antiserum (Sasaki et al. 2003) and then with goat anti-rabbit IgG horseradish peroxidase-conjugated secondary antibody. Detection of the peroxidase activity was performed according to the manufacturer’s instructions (Pierce, Rockford, IL, USA).
In vivo labeling of SLR1 protein and phosphoamino acid analysis
The preparation of 32P-labeled SLR1 protein and immunoprecipitation with anti-SLR1 antibody were described in Sasaki et al. (2003). Radioactive SLR1 was separated by SDS–PAGE and then blotted onto a polyvinylidene difluoride membrane (Millipore Co., Billerica, MA, USA). The band corresponding to the radioactive SLR1 was excised from the membrane and hydrolyzed in 6 M HCl for 1 h at 110°C (Kamps and Sefton 1989). The hydrolyzed amino acids were separated using two-dimensional thin-layer electrophoresis at pH 1.9 in the first dimension and pH 3.5 in the second dimension (King et al. 1983). Phosphoserine, phosphothreonine and phosphotyrosine standards were separated and detected with ninhydrin (ninhydrin spray; Wako, Tokyo, Japan). The gel was placed on an imaging plate for 1 week to detect the radiographic image as described above.
Preparation of SLR1 protein by DEAE column chromatography and the in vitro binding assay with GST–GID2 protein
The proteins were extracted from gid2, cps and WT calli (approximately 2 g) in liquid nitrogen and by resuspending in the extraction buffer as described above. The crude extracts were desalted on a Sephadex G-25 column (175 ml bed volume, Amersham Pharmacia Biotech) equilibrated and eluted with buffer A [20 mM Tris–HCl, pH 7.5, 0.5% Tween-20, 1 mM DTT, 1 mM EDTA, 1 mM phenylmethylsulfonyl fluoride (PMSF), 0.5× Complete). The fractions containing total proteins were pooled together and loaded onto a DEAE Sepharose column (4 ml bed volume, Amersham Pharmacia Biotech) equilibrated with buffer A. The column was washed with 10 column volumes of buffer A, and the bound proteins were eluted with 10 column volumes of buffer A containing 50, 100, 150 and 200 mM NaCl in a stepwise manner. Protein-enriched fractions (about 10 of 40 ml) were concentrated to 1 ml with a BioMax™ filter (Millipore Co., Billerica, MA, USA).
For the pull-down assay with recombinant GST–GID2, the proteins in the 150 mM NaCl elution fraction were desalted on a PD-10 column (Amersham Pharmacia Biotech) in CIP buffer and incubated with CIP at 37°C for 15 min. The CIP-treated and non-treated proteins (100 µl) were then incubated with GST or GST–GID2 (3 µg), which had been expressed in Escherichia coli (Gomi et al. 2004), in buffer B [900 µl: 50 mM Tris–HCl, pH 7.5, 150 mM NaCl, 5 mM MgCl2, 1 mM EDTA, 10% glycerol, 0.01% (w/v) Tween-20, 1 mM DTT] for 2 h at 4°C with gentle agitation. After incubation, the bound proteins were washed three times with 1 ml of buffer B and eluted with SDS sample buffer.
Acknowledgments
We thank Dr. H. Mori (Nagoya University) for assistance with the phosphoamino acid analysis, M. Hattori and I. Aichi for excellent technical assistance, and Dr. N. Shibuya for providing N-acetylchitoheptaose. This work was supported by a Grant-in-Aid for the Center of Excellence, a Grant-in-Aid from the Program for the Promotion of Basic Research Activities for Innovation Bioscience (M.M.) and a research fellowship from the Japan Society for the Promotion of Science (H.I. and A.S.).
These authors contributed equally to this work.
Fig. 1 Identification of the SLR1 phosphorylation site. (A) Radiographic image of the phosphoamino acids of SLR1 protein separated by two-dimensional thin-layer electrophoresis. The circles indicate the locations of standard phosphoamino acid residues detected by ninhydrin. P-S, phosphoserine; P-T, phosphothreonine; P-Y, phosphotyrosine. a, free Pi; b, non- or partially hydrolyzed 32P-labeled peptide fragments. (B) Diagram of the truncated constructs. Each of the four domains, DELLA, TVHYNP, Ser/Thr/Val-rich domain (polyS/T/V) and C-terminal GRAS, is indicated by different shading. Constructs Δ1 and Δ2 were N-terminally fused to HA and C-terminally fused to GFP as described in Itoh et al. (2002). Constructs Δ3 and Δ2+Δ3 were N-terminally fused with three copies of HA. The deletion points in each SLR1 mutant are shown below each box. (C) The effect of CIP treatment on each SLR1 mutant by immunoblot analysis. A crude extract of each transgenic plant was incubated at 37°C for 15 min with (+) or without (–) CIP. 0 indicates a sample without incubation. Asterisks indicate the phosphorylated forms of mutated SLR1 protein. A 10 µg aliquot of crude extracts was loaded per lane and probed with anti-SLR1 antibody.
Fig. 2 In vivo phosphorylation of endogenous SLR1 and SLR1 derivatives. Transgenic plants expressing each mutant SLR1 protein under the control of the actin1 promoter were incubated with 32PO4– and then treated with 100 µM GA3 for the indicated period. The 32P-labeled SLR1 protein was immunoprecipitated with anti-SLR1 antibody from 1.5 mg of total protein extract and then separated by SDS–PAGE. Finally, labeled SLR1 was detected by autoradiography. The asterisk and arrowhead indicate the phosphorylated forms of 3× HA-ΔDELLA+ΔTVHYNP (Δ3) and HA-ΔpolyS/T/V-GFP (Δ2), respectively. The phosphorylated form of the endogenous SLR1 was indicated by ‘end. SLR1′.
Fig. 3 Accumulation and degradation of SLR1 in gid2 and WT calli. (A) High level of accumulation of SLR1 protein in the callus. A crude extract of each sample was separated by SDS–PAGE and then probed with anti-SLR1 antibody. Each extract was incubated with (+) or without (–) CIP. Aliquots of 10 (lanes 2–5) or 15 (lanes 1, 6 and 7) µg of crude extracts were loaded per lane. Form I and form II indicate the non-phosphorylated and phosphorylated SLR1, respectively. (B) Gibberellin-dependent accumulation of SLR1 protein in the gid2 callus. The gid2 mutant callus was treated with 1 µM GA3 for the indicated time. A 10 µg aliquot of protein extracts was loaded per lane and probed with anti-SLR1 antibody. (C) Relative amount of phosphorylated (filled circles) and non-phosphorylated (open circles) SLR1 protein after GA3 treatment. Protein levels were quantified with NIH image software version 1.63. The level of non-phosphorylated SLR1 protein at 120 min after GA3 treatment was set as 100. (D) Gibberellin-dependent degradation of SLR1 protein in the WT callus. The WT callus was incubated with 1 µM GA3 for the indicated time. A 10 µg aliquot of protein extract was loaded per lane and probed with anti-SLR1 antibody. (E) Relative amounts of phosphorylated (filled circles) and non-phosphorylated (open circles) SLR1 protein after GA3 treatment. Protein levels were quantified as in (C). The level of phosphorylated SLR1 protein before GA3 treatment (time 0) was set as 100.
Fig. 4 Comparison of the phosphorylation status in the WT callus and in the cps and gid2 mutant plants. Crude extracts were separated by DEAE column chromatography as described in Materials and Methods. The bound proteins were eluted in a stepwise manner with buffer containing increasing concentrations of NaCl. The eluates from the DEAE columns were subjected to immunoblot analysis with anti-SLR1 antibody. A 5 µl aliquot of each elution fraction was loaded per lane.
Fig. 5 The phosphorylation of SLR1 protein is not required for its interaction with GID2. (A) The interaction between SLR1 and GID2 in vitro. Upper panel: immunoblot analysis of SLR1 protein in the 150 mM NaCl eluate of the gid2-1 extract. CIP treatment (+) dephosphorylated the phosphorylated SLR1 with increased mobility (lane 3). Twenty times the amounts of extracts were used for the pull-down experiment. Middle panel: Coomassie brilliant blue staining of the large subunit of RuBPC was shown as a loading control. Lower panel: protein pre-treated with (+) CIP or without (–) CIP was incubated with glutathione beads bound to either GST or GST–GID2. The interaction of SLR1 with GST–GID2 was detected by immunoblot analysis with anti-SLR1 antibody. The asterisk indicates the non-specific signal of the E. coli proteins co-purified with GST fusion protein under our experimental conditions. (B) The effects of kinase inhibitors on the gibberellin-dependent degradation of SLR1. WT calli were treated with K252a or staurosporine followed by treatment with 1 µM GA3. A 10 µg aliquot of each crude extract was loaded per lane and probed with anti-SLR1 antibody. Coomassie brilliant blue staining was shown as a loading control. (C) The effect of kinase inhibitors on the induction of EL2 mRNA in response to N-acetylchitooligosaccharide elicitor, GN7. WT calli were treated with K252a or staurosporine followed by the treatment with GN7 for 30min. A 10 µg aliquot of total RNA was analyzed by RNA gel blot analysis.
Amounts of endogenous GA20 and GA1 in calli and seedlings of WT, gid2-1 and cps (ng g–1 FW)
| Calli | Seedling | |||||
| WT | gid2-1 | cps | WT | gid2-1 | ||
| GA20 | 0.07 | ND | ND | 0.4 | 2.4 | |
| GA1 | 0.05 | 0.03 | ND | 0.3 | 51 | |
| Calli | Seedling | |||||
| WT | gid2-1 | cps | WT | gid2-1 | ||
| GA20 | 0.07 | ND | ND | 0.4 | 2.4 | |
| GA1 | 0.05 | 0.03 | ND | 0.3 | 51 | |
ND, not detected.
Amounts of endogenous GA20 and GA1 in calli and seedlings of WT, gid2-1 and cps (ng g–1 FW)
| Calli | Seedling | |||||
| WT | gid2-1 | cps | WT | gid2-1 | ||
| GA20 | 0.07 | ND | ND | 0.4 | 2.4 | |
| GA1 | 0.05 | 0.03 | ND | 0.3 | 51 | |
| Calli | Seedling | |||||
| WT | gid2-1 | cps | WT | gid2-1 | ||
| GA20 | 0.07 | ND | ND | 0.4 | 2.4 | |
| GA1 | 0.05 | 0.03 | ND | 0.3 | 51 | |
ND, not detected.
Abbreviations
- CIP
calf intestine alkaline phosphatase
- cps
CDP synthase
- GFP
green fluorescent protein
- gid2
gibberellin insensitive dwarf 2
- GST
glutathione S-transferase
- HA
hemagglutinin
- WT
wild type
References
Davies, P.J. (
del Pozo, J.C. and Estelle, M. (
Deshaies, R.J. (
Dill, A., Jung, H.S. and Sun, T.P. (
Dill, A., Thomas, S.G., Hu, J., Steber, C.M. and Sun, T.P. (
Fu, X., Richards, D.E., Ait-Ali, T., Hynes, L.W., Ougham, H., Peng, J. and Harberd, N.P. (
Fu, X., Richards, D.E., Fleck, B., Xie, D., Burton, N. and Harberd, N.P. (
Gagne, J.M., Downes, B.P., Shiu, S.H., Durski, A.M. and Vierstra, R.D. (
Gomi, K., Sasaki, A., Itoh, H., Ueguchi-Tanaka, M., Ashikari, M., Kitano, H. and Matsuoka, M. (
Gray, W.M., Kepinski, S., Rouse, D., Leyser, O. and Estelle, M. (
Gubler, F., Chandler, P.M., White, R.G., Llewellyn, D.J. and Jacobsen, J.V. (
Hiei, Y., Ohta, S., Komari, T. and Kumashiro, T. (
Hood, E.E., Helmer, G.L., Fraley, R.T. and Chilton, M.D. (
Ingram, G.C., Doyle, S., Carpenter, R., Schultz, E.A., Simon, R. and Coen, E.S. (
Itoh, H., Ueguchi-Tanaka, M., Sato, Y., Ashikari, M. and Matsuoka, M. (
Kamps, M.P. and Sefton, B.M. (
King, M.M., Fitzgerald, T.J. and Carlson, G.M. (
Kobayashi, M., Gomi, M., Agematsu, J., Asami, T., Yoshida, S. and Sakurai, A. (
Kuchitu, K., Kikuyama, M. and Shibuya, N. (
Kuroda, H., Takahashi, N., Shimada, H., Seki, M., Shinozaki, K. and Matsui, M. (
McGinnis, K.M., Thomas, S.G., Soule, J.D., Strader, L.C., Zale, J.M., Sun, T.P. and Steber, C.M. (
Nash, P., Tang, X., Orlicky, S., Chen, Q., Gertler, F.B., Mendenhall, M.D., Sicheri, F., Pawson, T. and Tyers, M. (
Nelson, D.C., Lasswell, J., Rogg, L.E., Cohen, M.A. and Bartel, B. (
Olszewski, N., Sun, T.P. and Gubler, F. (
Patton, E.E., Willems, A.R. and Tyers, M. (
Pysh, L.D., Wysocka-Diller, J.W., Camilleri, C., Bouchez, D. and Benfey, P.N. (
Ruegger, M., Dewey, E., Gray, W.M., Hobbie, L., Turner, J. and Estelle, M. (
Sambrook, J. and Russell, D.W. (
Sakamoto, T., Miura, K., Itoh, H., Tatsumi, T., Ueguchi-Tanaka, M., et al. (
Sasaki, A., Itoh, H., Gomi, K., Ueguchi-Tanaka, M., Ishiyama, K., et al. (
Silverstone, A.L., Jung, H.S., Dill, A., Kawaide, H., Kamiya, Y. and Sun, T.P. (
Somers, D.E., Schultz, T.F., Milnamow, M. and Kay, S.A. (
Winston, J.T., Strack, P., Beer-Romero, P., Chu, C.Y., Elledge, S.J. and Harper, J.W. (
Xie, D.X., Feys, B.F., James, S., Nieto-Rostro, M. and Turner, J.G. (